Abstract
Arabidopsis NODULIN HOMEOBOX (NDX) is a plant-specific transcriptional regulator whose role in small RNA biogenesis and heterochromatin homeostasis has recently been described. Here we extend our previous transcriptomic analysis to the flowering stage of development. We performed mRNA-seq and small RNA-seq measurements on inflorescence samples of wild-type and ndx1-4 mutant (WiscDsLox344A04) Arabidopsis plants. We identified specific groups of differentially expressed genes and noncoding heterochromatic siRNA (hetsiRNA) loci/regions whose transcriptional activity was significantly changed in the absence of NDX. In addition, data obtained from inflorescence were compared with seedling transcriptomics data, which revealed development-specific changes in gene expression profiles. Overall, we provide a comprehensive data source on the coding and noncoding transcriptomes of NDX-deficient Arabidopsis flowers to serve as a basis for further research on NDX function.
Similar content being viewed by others
Background & Summary
NODULIN HOMEOBOX (NDX) is a specific member of the homeobox family of transcription factors in flowering plants1,2, particularly in the Brassicaceae family, which includes the genetic model Arabidopsis thaliana. Besides its homeobox domain, NDX has two atypical domains called NDX-A and NDX-B, whose molecular functions are less understood. Recent molecular data indicate that NDX regulates the activity of some euchromatic genes in Arabidopsis thaliana plants. NDX was shown to interact with the E3 ubiquitin ligase module of the Polycomb Repressive Complex 1 (PRC1), RING1A and RING1B, establishing a functional link between histone H2A ubiquitination (H2Aub) and abscisic acid (ABA)-mediated repressive chromatin signalling3. Through the ABA pathway, NDX regulates seed germination and root growth. NDX was also shown to interact with the transcriptional repressor VIVIPAROUS1/ABI3-LIKE (VAL1) (through binding RING1A and RING1B), which regulates the activity of FLOWERING LOCUS C (FLC), a central integrator of the flowering transition in Arabidopsis4.
In addition to the above functions, NDX exhibits significant binding affinity to different nucleic acid substrates3,5. NDX was shown to bind to an R-loop structure at the 3′-end of FLC, which serves as a promoter for antisense (COOLAIR) transcription. COOLAIR is a set of antisense long noncoding RNAs that act as a repressor of FLC. The association of NDX and COOLAIR suggested a model in which COOLAIR transcription was repressed by NDX-mediated R-loop stabilization, which in turn altered FLC expression and flowering time.
Recent genomic data challenged the idea that NDX acts as a general R-loop regulator that controls the formation of chromosomal R-loops throughout the Arabidopsis genome. Instead, NDX appears to be linked to heterochromatin function by regulating the activity of heterochromatic siRNAs (het-siRNAs) and non-CG DNA methylation pathways at pericentromeric regions6. This observation is consistent with earlier data showing that NDX coincides with a heterochromatic patch of H3K9 dimethylation and Pol IV-dependent siRNA transcripts within the FLC terminator/COOLAIR promoter region5,7, linking NDX to heterochromatin function.
The molecular phenotype supporting the role of NDX in heterochromatin homeostasis was identified in genomic data of 10-day-old seedlings, however, similar functional studies were not performed in other stages of plant development. In particular, there is a lack of transcriptome data in Arabidopsis inflorescence, which awaits to be generated to understand the effect of NDX on the expression of flowering-related coding and noncoding RNAs. Herein, we extend our transcriptomic measurements to the flowering stage and present novel mRNA-seq and small RNA (sRNA)-seq data on inflorescence tissues of wild-type and ndx1-4 T-DNA insertion mutant (WiscDsLox344A04) plants (Fig. 1). Our analysis identified specific sets of genes differentially expressed in ndx1-4, as well as noncoding heterochromatic siRNA (het-siRNA) loci whose activity was significantly changed in the absence of NDX. Transcriptomic data obtained from inflorescence were also compared with previously published seedling data6, highlighting development-specific changes in gene expression profiles.
General workflow of mRNA-seq and sRNA-seq experiments performed in Arabidopsis thaliana. Two different tissue types were analysed: flowers and seedlings. The seedling data were published in ref. 6, which were used for comparison with the flower transcriptomic data generated in this study.
Methods
Arabidopsis thaliana Col-0 wild-type accession and ndx1-4 mutant (WiscDsLox344A04)5 plants were directly sown on soil and grown side-by-side at 21 °C on long day conditions (LD, 16 h light, 8 h dark photoperiod) until flowering stage. After five weeks, when the central flowering stem of plants reached approximately 20–25 cm and side branches were also present, inflorescence tissues were collected. Total RNA isolation was performed by the standard phenol-chloroform extraction method as described previously8. Briefly, approximately 30 mg plant material per sample was homogenized and resuspended in 600 μl of extraction buffer (0.1 M glycine-NaOH, pH 9.0, 100 mM NaCl, 10 mM EDTA, 2% SDS). The sample was mixed with an equal volume of phenol (pH 4.3, Sigma-Aldrich, P3803). The aqueous phase was treated with 600 μl of phenol-chloroform and chloroform, precipitated with ethanol. RNA samples were eluted in nuclease-free water and used in subsequent steps. To remove genomic contaminations, Dnase I treatments were performed based on manufacturer’s instructions (Ambion AM2222, www.thermofisher.com); RNA quality was assessed by testing RNA degradation based on agarose gel electrophoresis and presence of contaminants by Nanodrop spectrophotometer measurements (260/230 and 260/280 ratios). RNA quality was further assayed by capillary electrophoresis (Agilent 2100).
For RNA-seq, cDNA libraries were prepared from two independent biological replicates according to Illumina’s TruSeq RNA Library Preparation Kit v2 protocol. Equal amounts of total RNA extracts from 5 inflorescence tissues of different plants were pooled (to create one pool) for each biological replicate (see Tables 1, 2 for description of samples). The mRNA-seq libraries were sequenced using an Illumina NextSeq 500 instrument with 1 × 75 bp reads, generating 17.3–22.4 million reads per sample (Fig. 2a). Raw sequence data quality was assessed using FastQC and summarized reports were generated with MultiQC9. Duplication detection indicated >50% of reads as unique (Fig. 2b). Mean Phred quality scores for each read position in each mRNA sample were higher than 28 (indicative of ‘very good quality’) (Fig. 2c). Per sequence mean Phred quality scores were higher than 28 (indicative of ‘very good quality’) for at least 95% of reads in each mRNA sample (Fig. 2d).
MultiQC analysis of mRNA-seq data from flowers. (a) General NGS statistics of mRNA-seq samples. (b) Sequence Counts plot showing the total number of reads, classified as unique and duplicate. Duplication detection requires an exact sequence match over the whole length of the sequence. Any reads over 75 bp in length were truncated to 50 bp for this analysis. More about duplicate calculation: https://multiqc.info/. (c) Mean quality scores. The higher the score, the better the base call. Background color of the graph divides the y axis into very good quality calls (green zone), calls of reasonable quality (orange zone), and calls of poor quality (red zone). (d) Per sequence quality scores. The graph shows if a subset of sequences have universally low-quality values. Background color: of the graph very good quality calls (green), calls of reasonable quality (orange), and calls of poor quality (red).
Next, Salmon10 was used to map mRNA-seq reads and get transcript quantities for protein coding genes, noncoding genes and transposable elements. Reads were mapped to the Arabidopsis thaliana reference transcriptome (Araport11, coding and noncoding genes) to quantify gene expression levels as well as to transposable elements (Araport11) to quantify transposable element expression (see Table 3 for detailed description of Arabidopsis thaliana reference data used in the analyses). Transcript quantities of protein coding genes, noncoding genes, and transposable elements were corrected for potential fragment level GC bias using ‘salmon quant–gcBias’ that was shown to reduce isoform quantification errors10,11. Duplicate reads were filtered out from alignment to protein coding and noncoding genes (default behaviour of ‘salmon quant’) while duplicates were kept for transposable element alignments (‘salmon quant’ was used with ‘–keepDuplicates’ option set).
DESeq2 was employed to identify differentially expressed genes (DEGs) and transposable elements (ndx1-4 versus Col-0 wild type)12. The significance level was established based on the adjusted p-values with independent hypothesis weighting13, where p(adjusted) <0.01 was used to define DEGs and p(adjusted) <0.05 for differentially expressed transposable elements. For the purpose of data visualization, RNA-seq reads were aligned to the TAIR10 reference genome using HISAT214, which allowed for reporting spliced alignments. To create RPKM (Reads Per Kilobase per Million) normalized coverage files, we utilized deepTools bamCoverage15.
Expression changes of 31987 protein coding and noncoding genes and 34856 transposable elements were analysed. DEGs were visualized by hierarchical clustering in three different relations: (1) in inflorescence, we identified 269 (0.84%) up- and 145 (0.45%) downregulated genes in the ndx1-4 mutant vs. Col-0 control samples (Fig. 3a). Representative genes are shown in Fig. 3b–d. (2) The transcriptomes of inflorescence and seedlings6,16 were compared in Col-0 (control) samples (Fig. 4a), identifying 5555 (17.37%) up- and 5618 (17.56%) downregulated genes in flowers relative to seedlings. Representative genes are shown in Fig. 4b–d. 3) The transcriptomes of inflorescence and seedlings6,16 were compared in ndx1-4 mutant samples (Fig. 5a), identifying 5405 (16.9%) up- and 5645 (17.65%) downregulated genes in flowers vs. seedlings. In addition, 200 DEGs identified at the flowering stage between ndx1-4 and Col-0 overlapped with DEGs identified in seedlings. Representative genes are shown in Fig. 5b–d. Collectively, the above analysis identified a great number of ndx1-4-specific and development-specific changes in gene expression profiles.
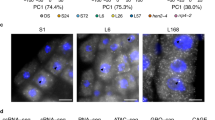
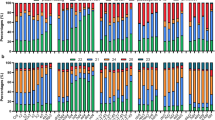
Differentially expressed genes in Col-0 and ndx1-4 mutant Arabidopsis inflorescence. (a) Hierarchical clustering of log2 fold change values (ndx1-4 vs. Col-0) calculated by DESeq2 analysis. Orange shows upregulated genes, blue shows downregulated genes. Two independent biological replicates are shown. (b) Representative screenshots of upregulated genes in ndx1-4 flowers. (c) Representative screenshots of downregulated genes in ndx1-4 flowers. (d) Representative screenshots of genes showing no differential expression in ndx1-4 flowers.
Differentially expressed genes in Col-0 seedlings and flowers. (a) Hierarchical clustering of log2 fold change values (seedling, Col-0 vs. flower, Col-0) calculated by DESeq2 analysis. Orange shows upregulated genes, blue shows downregulated genes. Two independent biological replicates are shown. (b) Representative screenshots of downregulated genes in flowers. (c) Representative screenshots of upregulated genes in flowers. (d) Representative gene showing no differential expression in Col-0 seedlings and flowers.
Differentially expressed genes in ndx1-4 mutant seedlings and flowers. (a) Hierarchical clustering of log2 fold change values (seedling, ndx1-4 vs. flower, ndx1-4) calculated by DESeq2 analysis. Orange shows upregulated genes, blue shows downregulated genes. Two independent biological replicates are shown. (b) Representative screenshots of downregulated genes in ndx1-4 flowers. (c) Representative screenshots of upregulated genes in ndx1-4 flowers. (d) Representative gene showing no differential expression in ndx1-4 seedlings and flowers.
Differential expression of 34856 transposable elements were analysed in inflorescence samples. Only a small number of transposable elements were differentially expressed: we identified 33 (0.09%) upregulated and 12 (0.03%) downregulated transposable elements in the ndx1-4 mutant vs. Col-0 control samples.
Comparing DEGs with TEs, 42 upregulated and 19 downregulated DEGs overlap with transposable elements, 7 upregulated and 3 downregulated DEGs overlap with differentially expressed transposable elements.
Description and accessibility of the generated datasets can be found in Tables 1, 2.
For the analysis of the noncoding transcriptome, sRNA libraries were prepared from 3-3 biological replicates according to Illumina’s NEBNext® Multiplex sRNA Library Prep protocol. For sRNA library preparations, equal amounts of total RNA extracts from 5 flowering tissues were pooled (to create one pool) for each biological replicate (see Tables 1, 2 for description of samples). Total RNA extraction was done as described above for mRNA-seq. The same total RNA prep was used for sRNA and mRNA sequencing. The sRNA-seq libraries were sequenced using an Illumina NextSeq500 instrument with 1 × 50 bp reads, generating 13.5–17.5 million reads per sample (Fig. 6a). Raw sequence data quality was assessed using FastQC and summarized reports were generated with MultiQC9. Duplication detection indicated 5–6 million unique reads per sample (Fig. 6b). Mean Phred quality scores for each read position in each sRNA sample were higher than 28 (indicative of ‘very good quality’) (Fig. 6c). Per sequence mean Phred quality scores were higher than 28 (indicative of ‘very good quality’) for at least 95% of reads in each sRNA sample (Fig. 6d).
MultiQC analysis of sRNA-seq data from flowers. (a) General NGS statistics of sRNA-seq samples. (b-d) Same as in Fig. 2.
The sRNA-seq data underwent additional processing using the sRNAnalyzer pipeline17. The Illumina adapters were initially removed by Cutadapt18, and the reads were then size-selected to range between 19–25 nt. These size-selected reads were subsequently aligned to a published comprehensive sRNA locus database19 (public data: www.ebi.ac.uk/ena/browser/view/PRJEB22276, additional material: https://github.com/seb-mueller/Arabidopsis_smallRNA_loci) using sRNAnalyzer. To determine differential expression of sRNA loci between Col-0 and ndx1-4 samples, DESeq2 was employed (p(adjusted) <0.05), and the hits were further filtered by absolute fold change (abs(FC) >1.5). For data visualization, reads were aligned to the Arabidopsis thaliana reference sequence (TAIR10) with bowtie220. Finally, we utilized deepTools bamCoverage to generate RPKM (Reads per Kilobase per Million) normalized coverage files. Differential expression of 16517 sRNA loci were analysed. Our analysis identified 642 (3.89%) up- and 991 (6%) downregulated sRNA loci in the ndx1-4 mutant relative to Col-0 control samples. At absolute fold change (abs(FC) >1.5) cutoff, we identified 375 (2.27%) up- and 179 (1.08%) downregulated sRNA loci that were visualized by hierarchical clustering (Fig. 7a). Representative sRNA loci are shown in Fig. 7b. Description and accessibility of the generated datasets can be found in Tables 1, 2.
Differentially expressed sRNA loci in Col-0 and ndx1-4 mutant Arabidopsis inflorescence. (a) Hierarchical clustering of log2 fold change values (flower, Col-0 vs. ndx1-4) calculated by DESeq2 analysis. Orange shows upregulated sRNA loci, blue shows downregulated sRNA loci. Three independent biological replicates are shown. (b) Representative screenshots of upregulated, downregulated sRNA loci in ndx1-4 flowers. An sRNA locus with no transcriptional change is also shown (bottom panel).
Data Records
The mRNA-seq and sRNA-seq datasets generated from Arabidopsis inflorescence tissues were deposited in Gene Expression Omnibus (GEO) under the accession number GSE22359121.
Genome browser (JBrowse) tracks containing relevant transcriptomic data are available at:
https://geneart.med.unideb.hu/pub/2023_ndx (login: ndx; password: athal23)
The mRNA and sRNA datasets generated from Arabidopsis seedlings can be accessed at GEO under the accession number GSE20184116 and in Supplementary Data 1–16 accompanying the related paper6.
Tables 1, 2 summarizes all transcriptomic experiments, NGS data, result tables, and identifiers to access relevant datasets. Table 3 lists all reference data used for analyses.
Technical Validation
The genotypes of Col-0 and ndx1-4 mutant Arabidopsis lines were confirmed by PCR using different combinations of genotyping primers designed for the NDX locus and the T-DNA (WiscDsLox344A04) insertion site (Fig. 8). The integrity, purity, and yield of extracted total RNA were determined by agarose gel electrophoresis and nanodrop spectrophotometry (Fig. 9) as well as Agilent bioanalyzer measurements (resulting in RIN values >9). Lack of mRNA expression from the mutagenized ndx1-4 locus was confirmed by RT-qPCR analysis (Fig. 10). RNA samples were extracted from the 10-day-old seedling and inflorescence tissues (Col-0 and ndx1-4 genotypes). To remove genomic contaminations, Dnase I treatments were performed based on manufacturer’s instructions (Ambion AM2222, www.thermofisher.com). These RNA samples were reverse transcribed with random hexamers using SuperScript IV reverse transcriptase (Thermo Fisher Scientific) as previously described8. For qPCR, we employed the ΔΔCt method and internal primers for normalization between samples. Primer sequences used for qPCR were as follows: NDX F AGCTGTAAAGTCAACTAACTGAGA; NDX R TCTAGATCCCATCTAACAAGAAACA; GAPDH1 F AGGAGCAAGGCAGTTAGTGGT; GAPDH1 R AGATGCGCCCATGTTCGTT. Real-time qPCR was subsequently conducted with a LightCycler 480 SYBR Green I Master mix (Roche) utilizing a QuantStudio 12 K Flex Real-Time PCR System (Thermo Fisher Scientific). NDX mRNA expression levels were normalized to GAPDH1 gene expression.
Genotyping of the ndx1-4 mutant plant line (Col-0 background). (a) Schematic representation of T-DNA insertion into the NDX locus along with the genotyping PCR primers ndx1-4_gLP AAAGCTCGTGTTGGCTAAGTG; ndx1-4_gRP AGGTTTCTGCAAACACCAGTG; WISC-TDNA AACGTCCGCAATGTGTTATTAAGTTGTC. Expected sizes of PCR amplicons in Col-0 and ndx1-4 mutant backgrounds are shown below. (b) Representative genotyping PCR in ndx1-4 mutant and Col-0 plant lines. (−): no template control.
Total RNA isolated from Col-0 and ndx1-4 mutant Arabidopsis seedlings and inflorescence samples. Upper panel: RNA integrity analysis performed by agarose gel electrophoresis. Dominant bands represent intact rRNAs. Lower panel: Purity and yield of total RNA samples measured by Nanodrop spectrophotometry and Agilent Bioanalyzer. OD 260/280 values > 1.8 and 260/230 values > 2.2 indicate the high purity of the samples. RIN: RNA integrity number.
RT-qPCR validation of NDX mRNA expression in Col-0 and ndx1-4 plants. (a) Genome browser screenshot showing mRNA-seq expression profiles over the NDX locus. Two independent biological replicates are shown. (b) RT-qPCR validation performed in Col-0 and ndx1-4 flowers and seedlings. NDX mRNA expression levels were normalized to GAPDH1 expression, as described in the related paper6. NDX mRNA is barely detected in the ndx1-4 mutant.
Usage Notes
We have created a comprehensive database of coding and noncoding transcriptomes of Arabidopsis inflorescence and seedling tissues that can be used for downstream genomic analysis. The NGS tracks provided can be displayed directly in freely accessible genome browsers. The list of differentially expressed genes can be directly used for molecular pathway analysis, GO term analysis or gene set enrichment analysis. The identified differentially regulated sRNA loci can be integrated with histone modification, DNA methylation, and transcriptomic maps for in-depth epigenomic analysis. Altogether, the data source we generated in Col-0 and ndx1-4 mutant plants is expected to lead us to a deeper understanding of the molecular function of NDX.
Code availability
No custom code was generated or applied for analysis of the genomic data presented. All software tools were referenced and used with default settings unless otherwise noted. Non-default parameters were as follows:
FastQC (v0.11.9), multiqc (1.14): Quality check of mRNA-seq and sRNA-seq reads.
Cutadapt (1.9.1): Adapter trimming of sRNA-seq reads (first pass parameters: -a AGATCGGAAGAGCACACGTCT -n 1 -e 0.2 -O 5 -m 1 --match-read-wildcards; second pass parameters: -g GTTCAGAGTTCTACAGTCCGACGATC -n 1 -e 0.125 -O 8 -m 1 --match-read-wildcards). After adapter trimming, reads were size-selected (using standard linux command line tools) to keep 18–25 nt reads.
sRNAnalyzer pipeline: Alignment of sRNA reads (kit: NEB; min-length: 8; alignment type: multiple) to the A. thaliana sRNA locus database19.
Salmon (1.7.0): Quantification of mRNA-seq reads has been performed with fragment GC bias correction (--gcBias). Duplicate reads were filtered out (default behaviour) for protein coding and noncoding genes. Duplicate reads were kept for transposable elements (--keepDuplicates).
DESeq2 (1.24.0): Adjusted p-value < 0.05 for differential gene expression analysis, adjusted p-value < 0.05 for differential transposable element expression analysis, adjusted p-value < 0.05 for differential sRNA expression analysis.
HISAT2 (2.1.0): mRNA-seq data alignment (reference: TAIR10) for visualization.
bowtie2 (2.3.5): sRNA-seq data alignment (reference: TAIR10) for visualization.
deepTools (3.3.0) bamCoverage: RPKM normalized coverage files of mRNA-seq (--binSize 1 --normalizeUsing RPKM) and sRNA-seq (-bs 15–smoothLength 45 --normalizeUsing RPKM) data for visualization in JBrowse.
References
Jørgensen, J. E. et al. A new class of plant homeobox genes is expressed in specific regions of determinate symbiotic root nodules. Plant Mol. Biol. 40, 65–77 (1999).
Mukherjee, K., Brocchieri, L. & Bürglin, T. R. A comprehensive classification and evolutionary analysis of plant homeobox genes. Mol. Biol. Evol. 26, 2775–2794 (2009).
Zhu, Y. et al. The Arabidopsis Nodulin Homeobox Factor AtNDX Interacts with AtRING1A/B and Negatively Regulates Abscisic Acid Signaling. Plant Cell 32, 703–721 (2020).
Mikulski, P., Benfey, P. N., Lu, T., Zhu, D. & Dean, C. VAL1 as an assembly platform co-ordinating co-transcriptional repression and chromatin regulation at Arabidopsis FLC. BioRxiv https://doi.org/10.1101/2021.07.21.453204.
Sun, Q., Csorba, T., Skourti-Stathaki, K., Proudfoot, N. J. & Dean, C. R-loop stabilization represses antisense transcription at the Arabidopsis FLC locus. Science 340, 619–21 (2013).
Karányi, Z. et al. NODULIN HOMEOBOX is required for heterochromatin homeostasis in Arabidopsis. Nat. Commun. 13, 5058 (2022).
Swiezewski, S. et al. Small RNA-mediated chromatin silencing directed to the 3′ region of the Arabidopsis gene encoding the developmental regulator, FLC. Proc. Natl. Acad. Sci. 104, 3633–3638 (2007).
Szaker, H. M. et al. miR824/AGAMOUS-LIKE16 Module Integrates Recurring Environmental Heat Stress Changes to Fine-Tune Poststress Development. Front. Plant Sci. 10 (2019).
Ewels, P., Magnusson, M., Lundin, S. & Käller, M. MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinforma. Oxf. Engl. 32, 3047–3048 (2016).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Love, M. I., Hogenesch, J. B. & Irizarry, R. A. Modeling of RNA-seq fragment sequence bias reduces systematic errors in transcript abundance estimation. Nat. Biotechnol. 34, 1287–1291 (2016).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Ignatiadis, N., Klaus, B., Zaugg, J. B. & Huber, W. Data-driven hypothesis weighting increases detection power in genome-scale multiple testing. Nat. Methods 13, 577–580 (2016).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Karányi, Z. et al. GEO. https://identifiers.org/geo/GSE201841 (2022).
Wu, X. et al. SRNAnalyzer-A flexible and customizable small RNA sequencing data analysis pipeline. Nucleic Acids Res. 45, 12140–12151 (2017).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 10–12 (2011).
Hardcastle, T. J., Müller, S. Y. & Baulcombe, D. C. Towards annotating the plant epigenome: The Arabidopsis thaliana small RNA locus map. Sci. Rep. 8, 1–15 (2018).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Feró, O. et al. GEO. https://identifiers.org/geo/GSE223591 (2023).
Acknowledgements
This study was supported by NKFIH-K-142137, NKFIH-NNE-130913, the Thematic Excellence Programme (TKP2021-EGA-18) of the Ministry for Innovation and Technology in Hungary. L.Sz. was supported by the Bolyai Janos fellowship of the Hungarian Academy of Sciences, and the UNKP-21-5-DE-11 new national excellence program of the Ministry for Innovation and Technology from the source of the National Research, Development and Innovation Fund. T.Cs received support from NKFIH-K129283, K137722, and K136513. We thank Caroline Dean (John Innes Centre, UK) for providing all the plant lines utilized in this study. We are grateful for the Genomic Medicine and Bioinformatics Core Facility (University of Debrecen) for the NGS service.
Funding
Open access funding provided by University of Debrecen.
Author information
Authors and Affiliations
Contributions
Á.M., Sz.H. and É.N. performed the experiments research, O.F., Zs.K., É.N., T.Cs. and L.Sz. analysed the data, L.Sz., and T.Cs. secured funding, L.Sz. supervised the work, O.F. and L.Sz. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Feró, O., Karányi, Z., Nagy, É. et al. Coding and noncoding transcriptomes of NODULIN HOMEOBOX (NDX)-deficient Arabidopsis inflorescence. Sci Data 10, 364 (2023). https://doi.org/10.1038/s41597-023-02279-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-023-02279-9
This article is cited by
-
Chromosomal R-loops: who R they?
Biologia Futura (2024)
-
DNA methylome, R-loop and clinical exome profiling of patients with sporadic amyotrophic lateral sclerosis
Scientific Data (2024)