Abstract
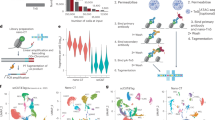
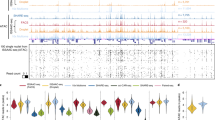
The ability to comprehensively analyze the chromatin state with single-cell resolution is crucial for understanding gene regulatory principles in heterogenous tissues or during development. Recently, we developed a nanobody-based single-cell CUT&Tag (nano-CT) protocol to simultaneously profile three epigenetic modalities—two histone marks and open chromatin state—from the same single cell. Nano-CT implements a new set of secondary nanobody-Tn5 fusion proteins to direct barcoded tagmentation by Tn5 transposase to genomic targets labeled by primary antibodies raised in different species. Such nanobody-Tn5 fusion proteins are currently not commercially available, and their in-house production and purification can be completed in 3–4 d by following our detailed protocol. The single-cell indexing in nano-CT is performed on a commercially available platform, making it widely accessible to the community. In comparison to other multimodal methods, nano-CT stands out in data complexity, low sample requirements and the flexibility to choose two of the three modalities. In addition, nano-CT works efficiently with fresh brain samples, generating multimodal epigenomic profiles for thousands of brain cells at single-cell resolution. The nano-CT protocol can be completed in just 3 d by users with basic skills in standard molecular biology and bioinformatics, although previous experience with single-cell assay for transposase-accessible chromatin using sequencing (scATAC-seq) is beneficial for more in-depth data analysis. As a multimodal assay, nano-CT holds immense potential to reveal interactions of various chromatin modalities, to explore epigenetic heterogeneity and to increase our understanding of the role and interplay that chromatin dynamics has in cellular development.
Key points
-
This protocol involves profiling gene regulatory dynamics in complex tissues at the single-cell level. This is achieved by generating two nanobody-Tn5 fusion proteins to recognize primary antibodies against widespread histone modifications.
-
The nanobody-Tn5 fusions are combined with ATAC-seq to simultaneously profile three epigenetic modalities from the same single cell, thousands of cells at the same time. A bioinformatic pipeline, Nanoscope, for seamless analysis of nano-CT datasets is also described.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
Raw nano-CT data were deposited in Gene Expression Omnibus under accession GSE198467. Source data are provided with this paper.
Code availability
All code, scripts, analysis pipeline and instructions for reproducibility can be found on Github at https://github.com/bartosovic-lab/nanoscope, and the analysis vignette can be found at https://fansalon.github.io/vignette_single-cell-nano-CT.html.
References
Spitz, F. & Furlong, E. E. M.Transcription factors: from enhancer binding to developmental control. Nat. Rev. Genet. 13, 613–626 (2012).
Moris, N., Pina, C. & Arias, A. M. Transition states and cell fate decisions in epigenetic landscapes. Nat. Rev. Genet. 17, 693–703 (2016).
Valencia, A. M. & Pașca, S. P. Chromatin dynamics in human brain development and disease. Trends Cell Biol. 32, 98–101 (2022).
Roadmap Epigenomics Consortium. et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
The ENCODE Project Consortium. et al. Expanded encyclopaedias of DNA elements in the human and mouse genomes. Nature 583, 699–710 (2020).
Zwart, W. et al. A carrier-assisted ChIP-seq method for estrogen receptor-chromatin interactions from breast cancer core needle biopsy samples. BMC Genomics 14, 232 (2013).
Schmidl, C., Rendeiro, A. F., Sheffield, N. C. & Bock, C. ChIPmentation: fast, robust, low-input ChIP-seq for histones and transcription factors. Nat. Methods 12, 963–965 (2015).
Skene, P. J. & Henikoff, S. An efficient targeted nuclease strategy for high-resolution mapping of DNA binding sites. eLife 6, e21856 (2017).
Kaya-Okur, H. S. et al. CUT&Tag for efficient epigenomic profiling of small samples and single cells. Nat. Commun. 10, 1930 (2019).
Wang, Q. et al. CoBATCH for high-throughput single-cell epigenomic profiling. Mol. Cell 76, 206–216.e7 (2019).
Bartosovic, M., Kabbe, M. & Castelo-Branco, G. Single-cell CUT&Tag profiles histone modifications and transcription factors in complex tissues. Nat. Biotechnol. 39, 825–835 (2021).
Wu, S. J. et al. Single-cell CUT&Tag analysis of chromatin modifications in differentiation and tumor progression. Nat. Biotechnol. 39, 819–824 (2021).
Adey, A. C. Tagmentation-based single-cell genomics. Genome Res. 31, 1693–1705 (2021).
Ma, S. et al. Chromatin potential identified by shared single-cell profiling of RNA and Chromatin. Cell 183, 1103–1116.e20 (2020).
Xu, W. et al. ISSAAC-seq enables sensitive and flexible multimodal profiling of chromatin accessibility and gene expression in single cells. Nat. Methods 19, 1243–1249 (2022).
Trevino, A. E. et al. Chromatin and gene-regulatory dynamics of the developing human cerebral cortex at single-cell resolution. Cell 184, 5053–5069.e23 (2021).
Gopalan, S., Wang, Y., Harper, N. W., Garber, M. & Fazzio, T. G. Simultaneous profiling of multiple chromatin proteins in the same cells. Mol. Cell 81, 4736–4746.e5 (2021).
Meers, M. P., Llagas, G., Janssens, D. H., Codomo, C. A. & Henikoff, S. Multifactorial profiling of epigenetic landscapes at single-cell resolution using MulTI-Tag. Nat. Biotechnol. 41, 708–716 (2023).
Stuart, T. et al. Nanobody-tethered transposition enables multifactorial chromatin profiling at single-cell resolution. Nat. Biotechnol. 41, 806–812 (2023).
Janssens, D. H. et al. CUT&Tag2for1: a modified method for simultaneous profiling of the accessible and silenced regulome in single cells. Genome Biol. 23, 81 (2022).
Yeung, J. et al. scChIX-seq infers dynamic relationships between histone modifications in single cells. Nat. Biotechnol. 41, 813–823 (2023).
Bartosovic, M. & Castelo-Branco, G. Multimodal chromatin profiling using nanobody-based single-cell CUT&Tag. Nat. Biotechnol. 41, 794–805 (2023).
Pleiner, T., Bates, M. & Görlich, D. A toolbox of anti–mouse and anti–rabbit IgG secondary nanobodies. J. Cell Biol. 217, 1143–1154 (2018).
Tedesco, M. et al. Chromatin Velocity reveals epigenetic dynamics by single-cell profiling of heterochromatin and euchromatin. Nat. Biotechnol. 40, 235–244 (2022).
Corces, M. R. et al. An improved ATAC-seq protocol reduces background and enables interrogation of frozen tissues. Nat. Methods 14, 959–962 (2017).
Amini, S. et al. Haplotype-resolved whole-genome sequencing by contiguity-preserving transposition and combinatorial indexing. Nat. Genet. 46, 1343–1349 (2014).
Plessy, C. et al. Linking promoters to functional transcripts in small samples with nanoCAGE and CAGEscan. Nat. Methods 7, 528–534 (2010).
Rykalina, V., Shadrin, A., Lehrach, H. & Borodina, T. qPCR-based characterization of DNA fragmentation efficiency of Tn5 transposomes. Biol. Methods Protoc. 2, bpx001 (2017).
Datlinger, P. et al. Ultra-high-throughput single-cell RNA sequencing and perturbation screening with combinatorial fluidic indexing. Nat. Methods 18, 635–642 (2021).
Stuart, T., Srivastava, A., Madad, S., Lareau, C. A. & Satija, R. Single-cell chromatin state analysis with Signac. Nat. Methods 18, 1333–1341 (2021).
Granja, J. M. et al. ArchR is a scalable software package for integrative single-cell chromatin accessibility analysis. Nat. Genet. 53, 403–411 (2021).
O’Neill, L. Immunoprecipitation of native chromatin: NChIP. Methods 31, 76–82 (2003).
Kasinathan, S., Orsi, G. A., Zentner, G. E., Ahmad, K. & Henikoff, S. High-resolution mapping of transcription factor binding sites on native chromatin. Nat. Methods 11, 203–209 (2014).
Mulqueen, R. M. et al. High-content single-cell combinatorial indexing. Nat. Biotechnol. 39, 1574–1580 (2021).
Picelli, S. et al. Tn5 transposase and tagmentation procedures for massively scaled sequencing projects. Genome Res. 24, 2033–2040 (2014).
Kaya-Okur, H. S., Janssens, D. H., Henikoff, J. G., Ahmad, K. & Henikoff, S. Efficient low-cost chromatin profiling with CUT&Tag. Nat. Protoc. 15, 3264–3283 (2020).
Heumos, L. et al. Best practices for single-cell analysis across modalities. Nat. Rev. Genet. 24, 550–572 (2023).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Luecken, M. D. et al. Benchmarking atlas-level data integration in single-cell genomics. Nat. Methods 19, 41–50 (2022).
Mannens, C. C. A. et al. Dynamics of chromatin accessibility during human first-trimester neurodevelopment. Preprint at bioRxiv https://doi.org/10.1101/2023.08.18.553878 (2023).
Zhu, C. et al. Joint profiling of histone modifications and transcriptome in single cells from mouse brain. Nat. Methods 18, 283–292 (2021).
Xiong, H., Luo, Y., Wang, Q., Yu, X. & He, A. Single-cell joint detection of chromatin occupancy and transcriptome enables higher-dimensional epigenomic reconstructions. Nat. Methods 18, 652–660 (2021).
Xie, Y. et al. Droplet-based single-cell joint profiling of histone modifications and transcriptomes. Nat. Struct. Mol. Biol. 30, 1428–1433 (2023).
Matsuoka, T. et al. Neural crest origins of the neck and shoulder. Nature 436, 347–355 (2005).
Sousa, V. H., Miyoshi, G., Hjerling-Leffler, J., Karayannis, T. & Fishell, G. Characterization of Nkx6-2-derived neocortical interneuron lineages. Cereb. Cortex 19, i1–i10 (2009).
Meyerhoff, J. et al. Microdissection of mouse brain into functionally and anatomically different regions. J. Vis. Exp. 2021, e61941 (2021).
Grandi, F. C., Modi, H., Kampman, L. & Corces, M. R. Chromatin accessibility profiling by ATAC-seq. Nat. Protoc. 17, 1518–1552 (2022).
Köster, J. & Rahmann, S. Snakemake—a scalable bioinformatics workflow engine. Bioinformatics 28, 2520–2522 (2012).
Zhang, Y. et al. Model-based Analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Meers, M. P., Tenenbaum, D. & Henikoff, S. Peak calling by Sparse Enrichment Analysis for CUT&RUN chromatin profiling. Epigenetics Chromatin 12, 42 (2019).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Zhang, B. et al. Characterizing cellular heterogeneity in chromatin state with scCUT&Tag-pro. Nat. Biotechnol. 40, 1220–1230 (2022).
Acknowledgements
We thank T. Jimenez-Beristain for writing laboratory animal ethics permit 1995_2019 and assistance with animal experiments; L. Kirby, M. Meijer, C. Zheng and P. Kukanja for assistance with animal experiments; and the staff at Comparative Medicine-Biomedicum. We thank M. Leboeuf and E. Hedlund´s group for valuable suggestions with figures. We acknowledge support from the National Genomics Infrastructure in Stockholm funded by the Science for Life Laboratory, the Knut and Alice Wallenberg Foundation and the Swedish Research Council. Computation/data handling was enabled by resources provided by the Swedish National Infrastructure for Computing (SNIC) at the Uppsala Multidisciplinary Center for Advanced Computational Science, partially funded by the Swedish Research Council through grant agreement number 2022-06725. We thank Abcam for providing a set of histone modification antibodies for CUT&Tag testing. M.B. was funded by the Vinnova Seal of Excellence Marie-Sklodowska Curie Actions grant RNA-centric view on Oligodendrocyte lineage development (RODent). Work in the research group of G.C.-B. was supported by the Swedish Research Council (grant no. 2019-01360), the European Union (Horizon 2020 Research and Innovation Programme/European Research Council Consolidator Grant EPIScOPE, grant agreement number 681893), the Swedish Cancer Society (Cancerfonden; 190394 Pj), the Knut and Alice Wallenberg Foundation (grants 2019-0107 and 2019-0089), the Swedish Society for Medical Research (SSMF, grant JUB2019), the Göran Gustafsson Foundation for Research in Natural Sciences and Medicine, the Ming Wai Lau Centre for Reparative Medicine and the Karolinska Institutet. Work in M.B.’s laboratory is supported by the Swedish Research Council (grant no. 2021-01476), Carl Tryggers Stiftelse (CTS22:2079), Jaensson’s Stiftelse, Ole Engkvist Stiftelse – Swedish foundation’s starting grant and Cancerfonden Junior Investigator Award (23 0758 JIA). F.A. was supported by the European Committee for Treatment and Research in Multiple Sclerosis (ECTRIMS) in the scope of the Postdoctoral Research Fellowship. All experimental procedures on animals were performed by following the European directive 2010/63/EU, local Swedish directive L150/SJVFS/2019:9, Saknr L150 and Karolinska Institutet complementary guidelines for the procurement and use of laboratory animals, Dnr. 1937/03–640. The procedures described were approved by the local committee for ethical experiments on laboratory animals in Sweden (Stockholms Norra Djurförsöksetiska nämnd), license number 144/16, 1995_2019 and 7029/2020.
Author information
Authors and Affiliations
Contributions
M.B. and G.C.-B. designed and developed the nano-CT protocol. M.B. performed the experiments. M.B., F.A. and B.H. performed bioinformatic data analysis and developed the Nanoscope pipeline and vignette. E.S. and T.N. performed the purification of nanobody-Tn5 recombinant protein and wrote the part of the manuscript describing recombinant protein expression and purification. J.R.B.-W. and M.B. wrote the manuscript and prepared the figures.
Corresponding author
Ethics declarations
Competing interests
G.C.-B. and M.B. have filed a patent application on the basis of this work (European patent application number EP22160860.7).
Peer review
Peer review information
Nature Protocols thanks Leif Ludwig and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key reference using this protocol
Bartosovic, M. & Castelo-Branco, G. Nat. Biotechnol. 41, 794–805 (2023): https://doi.org/10.1038/s41587-022-01535-4
Extended data
Extended Data Fig. 1 Workflows for activity assessment of in-house-produced Tn5.
a, Supplementary Method for assessing Tn5 activity on the basis of inverse qPCR detection. Step numbers corresponding to the Supplementary Method A numeration are shown to the right. b, Titration of in-house-produced Tn5 (ME-B) from different batches by qPCR. By assessing the activity of Tn5 assembly batches with respect to fragmenting a known template, we can obtain information about the volume to be used for the tagmentation of LA products from nano-CT library preparation (Step 108 from the Procedure). c, Supplementary Method for unimodal nano-CT in bulk (without single-cell barcoding) to perform titration with Tn5 (ME-B) and use this value for single-cell nano-CT. Step numbers corresponding to the Supplementary Method B numeration are shown to the right. d, Bioanalyzer traces performing nano-CT in bulk, showing different ratios of Tn5 (ME-B) and different size distributions of the final nano-CT libraries. Ct, cycle threshold.
Extended Data Fig. 2 Nano-CT library structure.
a and b, Full amplification sequence with Illumina P5 (a) and P7 (b) sequences for the nano-CT library. Note that both a and b correspond to the same nano-CT library molecule and are divided by genomic insert sequences. The custom primers required are for Read 1 and for Index 2 (shown in purple). Importantly, the ‘Modality barcode (8 nt)’ will contain the sequence from oligo adapters used during transposome complex loading (barcodes A, B and C in Steps 28–32). Library sequences will be demultiplexed by modalities by the Nanoscope pipeline. c, Schematic representation of the nano-CT library structure.
Supplementary information
Source data
Source Data Fig. 3
Unprocessed Coomassie-stained SDS-PAGE gels loaded with 10 µl of each selected fraction from psfaMsTn5 (c) and psfaRbTn5 (d) purifications.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bárcenas-Walls, J.R., Ansaloni, F., Hervé, B. et al. Nano-CUT&Tag for multimodal chromatin profiling at single-cell resolution. Nat Protoc 19, 791–830 (2024). https://doi.org/10.1038/s41596-023-00932-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-023-00932-6
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.