Abstract
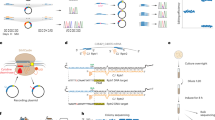
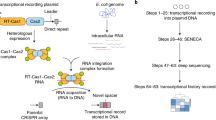
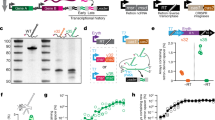
Biological signals occur over time in living cells. Yet most current approaches to interrogate biology, particularly gene expression, use destructive techniques that quantify signals only at a single point in time. A recent technological advance, termed the Retro-Cascorder, overcomes this limitation by molecularly logging a record of gene expression events in a temporally organized genomic ledger. The Retro-Cascorder works by converting a transcriptional event into a DNA barcode using a retron reverse transcriptase and then storing that event in a unidirectionally expanding clustered regularly interspaced short palindromic repeats (CRISPR) array via acquisition by CRISPR–Cas integrases. This CRISPR array-based ledger of gene expression can be retrieved at a later point in time by sequencing. Here we describe an implementation of the Retro-Cascorder in which the relative timing of transcriptional events from multiple promoters of interest is recorded chronologically in Escherichia coli populations over multiple days. We detail the molecular components required for this technology, provide a step-by-step guide to generate the recording and retrieve the data by Illumina sequencing, and give instructions for how to use custom software to infer the relative transcriptional timing from the sequencing data. The example recording is generated in 2 d, preparation of sequencing libraries and sequencing can be accomplished in 2–3 d, and analysis of data takes up to several hours. This protocol can be implemented by someone familiar with basic bacterial culture, molecular biology and bioinformatics. Analysis can be minimally run on a personal computer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Sequencing data associated with this study are available in the NCBI SRA (PRJNA838025).
Code availability
The latest version of the analysis code can be accessed through our lab GitHub (https://github.com/Shipman-Lab/Spacer-Seq_Nat-Protocols). The release at time of manuscript publishing is available at https://doi.org/10.5281/zenodo.7549326.
References
Siuti, P., Yazbek, J. & Lu, T. K. Synthetic circuits integrating logic and memory in living cells. Nat. Biotechnol. 31, 448–452 (2013).
Bonnet, J., Yin, P., Ortiz, M. E., Subsoontorn, P. & Endy, D. Amplifying genetic logic gates. Science 340, 599–603 (2013).
Yang, L. et al. Permanent genetic memory with >1-byte capacity. Nat. Methods 11, 1261–1266 (2014).
Farzadfard, F. & Lu, T. K. Genomically encoded analog memory with precise in vivo DNA writing in living cell populations. Science 346, 1256272 (2014).
Courbet, A., Endy, D., Renard, E., Molina, F. & Bonnet, J. Detection of pathological biomarkers in human clinical samples via amplifying genetic switches and logic gates. Sci. Transl. Med. 7, 289ra83–289ra83 (2015).
Roquet, N., Soleimany, A. P., Ferris, A. C., Aaronson, S. & Lu, T. K. Synthetic recombinase-based state machines in living cells. Science 353, aad8559 (2016).
Hsiao, V., Hori, Y., Rothermund, P. W. & Murray, M. M. A population-based temporal logic gate for timing and recording chemical events. Mol. Syst. Biol. 12, 869 (2016).
Weinberg, B. H. et al. Large-scale design of robust genetic circuits with multiple inputs and outputs for mammalian cells. Nat. Biotechnol. 35, 453–462 (2017).
Perli, S. D., Cui, C. H. & Lu, T. K. Continuous genetic recording with self-targeting CRISPR-Cas in human cells. Science 353, aag0511 (2016).
Tang, W. & Liu, D. R. Rewritable multi-event analog recording in bacterial and mammalian cells. Science 360, eaap8992 (2018).
Kempton, H. R., Love, K. S., Guo, L. Y. & Qi, L. S. Scalable biological signal recording in mammalian cells using Cas12a base editors. Nat. Chem. Biol. 1–9 (2022)
Chen, W. et al. Multiplex genomic recording of enhancer and signal transduction activity in mammalian cells. Preprint at bioRxiv https://doi.org/10.1101/2021.11.05.467434 (2021).
Loveless, T. B. et al. Molecular recording of sequential cellular events into DNA. Preprint at bioRxiv https://doi.org/10.1101/2021.11.05.467507 (2021).
Choi, J. et al. A time-resolved, multi-symbol molecular recorder via sequential genome editing. Nature 608, 98–107 (2022).
Shipman, S. L., Nivala, J., Macklis, J. D. & Church, G. M. Molecular recordings by directed CRISPR spacer acquisition. Science 353, aaf1175 (2016).
Sheth, R. U., Yim, S. S., Wu, F. L. & Wang, H. H. Multiplex recording of cellular events over time on CRISPR biological tape. Science 358, 1457–1461 (2017).
Schmidt, F., Cherepkova, M. Y. & Platt, R. J. Transcriptional recording by CRISPR spacer acquisition from RNA. Nature 562, 380–385 (2018).
Yim, S. S. et al. Robust direct digital-to-biological data storage in living cells. Nat. Chem. Biol. 17, 246–253 (2021).
Sheth, R. U. & Wang, H. H. DNA-based memory devices for recording cellular events. Nat. Rev. Genet. 19, 718–732 (2018).
Lear, S. K. & Shipman, S. L. Molecular recording: transcriptional data collection into the genome. Curr. Opin. Biotechnol. 79, 102855 (2023).
Bhattarai-Kline, S. et al. Recording gene expression order in DNA by CRISPR addition of retron barcodes. Nature https://doi.org/10.1038/s41586-022-04994-6 (2022).
Yosef, I., Goren, M. G. & Qimron, U. Proteins and DNA elements essential for the CRISPR adaptation process in Escherichia coli. Nucleic Acids Res. 40, 5569–5576 (2012).
Nuñez, J. K. et al. Cas1–Cas2 complex formation mediates spacer acquisition during CRISPR–Cas adaptive immunity. Nat. Struct. Mol. Biol. 21, 528–534 (2014).
Shipman, S. L., Nivala, J., Macklis, J. D. & Church, G. M. CRISPR–Cas encoding of a digital movie into the genomes of a population of living bacteria. Nature 547, 345–349 (2017).
Silas, S. et al. Direct CRISPR spacer acquisition from RNA by a natural reverse transcriptase–Cas1 fusion protein. Science 351, aad4234 (2016).
Tanna, T., Schmidt, F., Cherepkova, M. Y., Okoniewski, M. & Platt, R. J. Recording transcriptional histories using Record-seq. Nat. Protoc. 15, 513–539 (2020).
Schmidt, F. et al. Noninvasive assessment of gut function using transcriptional recording sentinel cells. Science 376, eabm6038 (2022).
Yehl, K. & Lu, T. Scaling computation and memory in living cells. Curr. Opin. Biomed. Eng. 4, 143–151 (2017).
Nuñez, J. K., Bai, L., Harrington, L. B., Hinder, T. L. & Doudna, J. A. CRISPR immunological memory requires a host factor for specificity. Mol. Cell 62, 824–833 (2016).
Yoganand, K. N. R., Sivathanu, R., Nimkar, S. & Anand, B. Asymmetric positioning of Cas1–2 complex and Integration Host Factor induced DNA bending guide the unidirectional homing of protospacer in CRISPR-Cas type I-E system. Nucleic Acids Res. 45, 367–381 (2017).
Sharon, E. et al. Functional genetic variants revealed by massively parallel precise genome editing. Cell 175, 544–557.e16 (2018).
Kong, X. et al. Precise genome editing without exogenous donor DNA via retron editing system in human cells. Protein Cell 12, 899–902 (2021).
Lopez, S. C., Crawford, K. D., Lear, S. K., Bhattarai-Kline, S. & Shipman, S. L. Precise genome editing across kingdoms of life using retron-derived DNA. Nat. Chem. Biol. 18, 199–206 (2022).
Zhao, B., Chen, S.-A. A., Lee, J. & Fraser, H. B. Bacterial retrons enable precise gene editing in human cells. CRISPR J. 5, 31–39 (2022).
Palka, C., Fishman, C. B., Bhattarai-Kline, S., Myers, S. A. & Shipman, S. L. Retron reverse transcriptase termination and phage defense are dependent on host RNase H1. Nucleic Acids Res. 50, 3490–3504 (2022).
Munck, C., Sheth, R. U., Freedberg, D. E. & Wang, H. H. Recording mobile DNA in the gut microbiota using an Escherichia coli CRISPR–Cas spacer acquisition platform. Nat. Commun. 11, 95 (2020).
Lee, P. Y., Costumbrado, J., Hsu, C. Y. & Kim, Y. H. Agarose gel electrophoresis for the separation of DNA fragments. J. Vis. Exp. 62, 3923 (2012).
Kluyver, T., et al. Jupyter Notebooks—a publishing format for reproducible computational workflows. In: Loizides, F. & Schmidt, B. (eds.) Positioning and Power in Academic Publishing: Players, Agents and Agendas, 87–90 (IOS Press, 2016).
Joshi, N.A. & Fass, J.N. Sickle: a sliding-window, adaptive, quality-based trimming tool for FastQ files (Version 1.33) Available at: https://github.com/najoshi/sickle. (2011).
JoVE Science Education Database. Microbiology. Serial Dilutions and Plating: Microbial Enumeration (JoVE, 2022).
Acknowledgements
This work was supported by funding from the National Science Foundation (2137692), the NIH/NIGMS (1DP2GM140917-01) and the Pew Biomedical Scholars Program. S.L.S. is a Chan Zuckerberg Biohub investigator and acknowledges additional funding support from the L.K. Whittier Foundation. S.K.L. was supported by an NSF Graduate Research Fellowship (2034836). S.C.L. was supported by a Berkeley Fellowship for Graduate Study.
Author information
Authors and Affiliations
Contributions
S.K.L. and S.C.L. wrote the protocol, with S.K.L. focusing on the experimental components and S.C.L. focusing on the computational components. The protocol was revised on the basis of input from A.G.-D., S.B.-K. and S.L.S. Additional simulation data was generated and analyzed by S.K.L. and S.C.L. using the Spacer-Seq code adapted into a JupyterLab notebook written by S.C.L. Figures were contributed by S.K.L., S.C.L. and A.G.D.
Corresponding author
Ethics declarations
Competing interests
S.L.S. is a named inventor on a patent application assigned to Harvard College, ‘Method of recording multiplexed biological information into a CRISPR array using a retron’ (US20200115706A1). The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks Michelle Chan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key reference using this protocol
Bhattarai-Kline, S. et al. Nature 608, 217–225 (2022): https://doi.org/10.1038/s41586-022-04994-6
Supplementary information
Supplementary Information
Supplementary Method and Supplementary Fig. 1.
Supplementary Table 1
Indexing primer sequences for multiplexing samples during library preparation.
Supplementary Table 2
KAPA Library Quantification Data Analysis Template as described for the KAPA Library Quantification Kit. Worksheet provides a readme page, an analysis page for data input, and a summary page for data output.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lear, S.K., Lopez, S.C., González-Delgado, A. et al. Temporally resolved transcriptional recording in E. coli DNA using a Retro-Cascorder. Nat Protoc 18, 1866–1892 (2023). https://doi.org/10.1038/s41596-023-00819-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-023-00819-6
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.