Abstract
Imaging across different scales is essential for understanding healthy organ morphology and pathophysiological changes. The macro- and microscale three-dimensional morphology of large samples, including intact human organs, is possible with X-ray microtomography (using laboratory or synchrotron sources). Preparation of large samples for high-resolution imaging, however, is challenging due to limitations such as sample shrinkage, insufficient contrast, movement of the sample and bubble formation during mounting or scanning. Here, we describe the preparation, stabilization, dehydration and mounting of large soft-tissue samples for X-ray microtomography. We detail the protocol applied to whole human organs and hierarchical phase-contrast tomography at the European Synchrotron Radiation Facility, yet it is applicable to a range of biological samples, including complete organisms. The protocol enhances the contrast when using X-ray imaging, while preventing sample motion during the scan, even with different sample orientations. Bubbles trapped during mounting and those formed during scanning (in the case of synchrotron X-ray imaging) are mitigated by multiple degassing steps. The sample preparation is also compatible with magnetic resonance imaging, computed tomography and histological observation. The sample preparation and mounting require 24–36 d for a large organ such as a whole human brain or heart. The preparation time varies depending on the composition, size and fragility of the tissue. Use of the protocol enables scanning of intact organs with a diameter of 150 mm with a local voxel size of 1 μm. The protocol requires users with expertise in handling human or animal organs, laboratory operation and X-ray imaging.
Similar content being viewed by others
Introduction
The quantification of human organ morphology, in both health and disease, is a complex task that can be tackled by multimodal spatial imaging modalities, capable of spanning across dimensional scales. Complete tissue morphological characterization requires the detection of interactions between and across scales; however, most imaging techniques are limited by either resolution or field of view, making it difficult to bridge macroscopic with microscopic observations and data. Conventional histology1,2,3 or electron microscopy4,5,6 approaches permit the visualization of the tissue’s microstructural organization and composition through serial sections, and the data can be suitably quantified; however, these approaches normally require sampling and sectioning of the tissue and are extremely labor intensive and time consuming. Optical clearing combined with light sheet microscopy can provide a large field of view with high resolution; however, tissue clearing also requires large timescales and is often expensive; in addition, the depth of imaging for a light sheet microscope is limited by the objective lens working distance7,8. Even where whole adult human organs8 or whole animals9 have been cleared over a period of several months, imaging them remains challenging. Similar drawbacks apply to optical coherence tomography10,11, multiphoton microscopy12 or confocal microscopy13,14, which can capture the local three-dimensional (3D) microstructure of the tissue at the cellular scale, but have limited tissue penetration, hindering deep tissue imaging15. Recently, high-resolution magnetic resonance imaging (MRI) achieved an isotropic voxel size of 100 µm in a whole ex vivo human brain16. Although MRI is nondestructive and has a large field of view17, the resolution is still not sufficient to examine tissue microstructure. Hierarchical imaging techniques are capable of overcoming the trade-off between resolution and field of view. In a hierarchical approach, multiple images of the same sample are acquired at different resolutions to bridge the different scales. Microcomputed tomography (µCT) has been used to image entire lungs with a resolution of 150 μm voxels, followed by subsequent extraction of biopsy cores in the lungs; these small cores were then scanned with µCT to achieve 10 μm voxels18.
Considerable progress has been made in the field of X-ray imaging over the past decade, especially for the visualization of soft tissue19. Synchrotron X-ray computed tomography (sCT) has proved to be one of the most powerful X-ray-based imaging techniques due to its high brightness20, enabling the observation of soft tissue at high resolution, with enough contrast to detect microstructural components21, such as individual neurons22 or elastin fibers23. In particular, phase-contrast-based sCT24, combined with optimized sample preparation and mounting procedures has provided extensive information of heart fiber orientation25,26 and, separately, of brain cellular maps27. Phase-contrast imaging refers to the detection of phase shifts of an X-ray beam imparted by a sample28. This technique enables the visualization of soft tissues that would be undetectable with conventional X-ray tomography in the absence of a contrast agent29. Nevertheless, the studies using sCT are mostly limited to fetal organs30, small subsamples of adult human organs31 or organs from small animal models, e.g., mouse32 and rabbit33, due to the restricted field of view. Some studies have demonstrated the imaging of larger samples, such as coelacanths with a diameter reaching 10 cm and a height of 30 cm34,35; however, the voxel size was limited to 30 µm and the structure of interest were hard tissues or cartilages.
Fourth-generation synchrotron sources, such as the European Synchrotron Radiation Facility (ESRF)’s Extremely Brilliant Source, provide enough beam spatial coherence and flux to visualize intact human organs from the macro- to the microscale. Using the ESRF’s Extremely Brilliant Source, we have recently developed a technique termed hierarchical phase-contrast tomography36 (HiP-CT) that allows the scanning of large intact human organs with ~20 µm isotropic voxels, with subsequent zooming (without sectioning), achieving up to 1 μm isotropic voxels locally. Although often overlooked, sample preparation and mounting are crucial to achieve the highest resolutions with this and other techniques, especially when visualizing large soft tissue samples, such as human organs, where the ratio of voxel size to organ diameter is 1:150,000 (1 µm in 150 mm). The maximum imageable diameter is limited by the equipment and setup parameters of the beamline, such as the width of the X-ray beam, the size of the detector, the computing power available to reconstruct the data and the size of the data. Currently, the maximum organ diameter we have imaged is 150 mm, but the present setup would be compatible up to 250 mm diameter by 500 mm vertically.
The success of all these experimental techniques depends on the careful preparation of the sample to avoid imaging artifacts. Soft tissue imaging using µCT or sCT presents a number of challenges compared with hard tissues. One of these issues is the lack of contrast when imaged with X-rays due to the similar densities of the sample components. Furthermore, the back-projection algorithms typically used to reconstruct the 3D volume from X-ray projections assume no movement of the sample during scanning. Most sample preparation methods are designed to prevent drifting or deformation of the sample, to avoid movement artifacts that would reduce the quality of the images37. Some reconstruction algorithms, such as motion compensation38,39 or machine learning based40,41, have been developed to overcome this issue42 but they remain complex and computationally costly to operate19.
Although the protocol we describe is applicable to a wide range of biological samples of varying sizes and using a range of imaging modalities, here we focus on samples to be imaged using synchrotron phase contrast imaging. The method is also applicable to other imaging modalities; however, the procedure may need optimizing according to sample size and modality chosen (see protocol steps for possible optimization recommendations).
Development and overview of the protocol
The sample preparation and mounting protocol described herein was developed from the need to image large biological volumes, such as intact human lung, brain, heart, kidney and spleen. The technique enables scanning of whole organs with 25 µm isotropic voxel size. Areas are then selected for further high-resolution scanning without requiring biopsies. To image such large structures, a high-energy X-ray beam is required to penetrate the samples. We used a polychromatic beam with energy ranging from 64 to 120 keV for organ imaging, but higher energies (up to 140 keV) would be optimal, if available.
Preparation and stabilization of the organ for this technique is essential as any density inhomogeneity, gas bubbles or movement during scanning would greatly reduce the image quality. During the development of the method, several challenges arose, as described in the Extended Data Fig. 1. Bubble entrapment (during mounting) and bubble formation (during scanning), both create motion blurring and phase contrast artifacts, and reduce the scan quality. This was resolved by limiting the dose absorbed by the sample and including multiple degassing steps during the organ preparation and mounting procedure. Sample movement during scanning is a further challenge, particularly as the samples are large, and hence often require a long time (>5 h) to image. This challenge was solved by carefully packing a mixture of crushed agar gel and liquid (in our case ethanol 70%) as a mounting media around the organ. In addition, the dehydration of the organ with ethanol increased the contrast of the images43 and diminished the bubble formation.
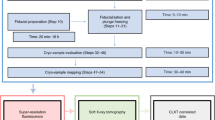
Here, we present a procedure to prepare whole human organs for imaging with sCT, µCT, medical CT and MRI that is compatible with a final stage of classical paraffin embedded histology. In this protocol, we mainly describe the sample preparation and mounting with ethanol-agar; the X-ray imaging protocol using HiP-CT and its application to a medical application (quantifying the damage coronavirus disease 2019 (COVID-19) does to lung vasculature)44,45. In brief, after fixation of the body (Step 1A(i)), the organ(s) are extracted (Step 1A(ii–iii)), immersion fixed (Step 2), dehydrated and degassed with vacuum (Step 4A) or thermal cycles (Step 4B) depending on its fragility, mounted with crushed agar gel mixture (Steps 5–33) and imaged using sCT (Steps 34–54), µCT (Step 55), clinical CT (Step 55) and MRI (Step 55), and finally, histological analysis is performed (Step 56). For an overview of the procedure, see Fig. 1.
The protocol contains three major steps (organ acquisition and preparation, organ mounting, and imaging) indicated in color-coded boxes. The organ can be retrieved either from a biobank, a surgery, a dismissed transplantation or it can be extracted from a donated body. We provide two protocols for degassing depending on the fragility of the organ (vacuum degassing and thermal cycles degassing). Once the sample is mounted with the agar gel, different imaging techniques can be performed (µCT, sCT, clinical CT and MRI). After imaging, histology can be carried out on the sample.
Advantages and limitations
This protocol was developed and optimized for the preparation of whole human organs to be imaged at high-resolution with HiP-CT, as shown in Fig. 2, and has been applied to various organs including brain, heart, lung, kidney and spleen. Nevertheless, the organ preparation procedure is flexible and can be used with soft tissues of animal origin or even with intact small animals. A list of human organs and biological samples imaged with this method and the preparation time for each step is provided in Extended Data Fig. 3. Most common sample preparation methods for X-ray imaging include fixation, alcoholic dehydration, paraffin embedding or critical-point drying. One of the main drawback of wet embedding such as alcoholic immersion is sample drift, which creates movement artifacts37. Paraffin embedding and critical-point drying modify the specimen structure to increase its rigidity, enabling the specimen to be immobile for a long period of time.
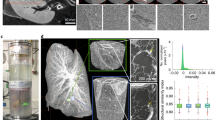
a, 3D view of the human heart imaged using the beamline BM05 at ESRF. b–d, Cross-sections of the heart with an isotropic voxel size of 25 µm (b), 6.5 µm (c) and 2.5 µm (d). e, Magnification of the 2.5 voxel image with annotation on the principal structures observed (ca, coronary artery; mc, myocyte cells; ad, adipose tissue). All the basic information on the patient from whom this organ originates is provided in Extended Data Fig. 2. All experiments followed the relevant governmental and institutional ethics regulations for human experiments.
Prevention of sample movement and shrinkage
In this protocol, sample movement is prevented by the use of small blocks of agar gel and crushed agar gel in density equilibrium with the mounting liquid, holding the sample in place. Scans taken several months apart on the same sample prepared with our organ-stabilization protocol can simply be registered using manual rigid transformation. This demonstrates the stability of this method over long time periods. An issue common to all these sample preparation methods is tissue shrinkage. This effect can hinder morphological quantification and lead to erroneous analyses. However, compared with paraffin embedding46,47 and critical point drying37,48, the sample shrinkage in this protocol can be mitigated using multiple ethanol baths in ascending ethanol concentrations43 up to 70%, thus preserving the morphology of the tissue. Furthermore, the dehydration of the sample with ethanol ensures a higher contrast with µCT43,49 or sCT50. This specimen preparation protocol is compatible with many MRI, clinical CT and histology (after 3D imaging and dismounting) techniques. Finally, whilst the specific imaging equipment may be specialized, the preparation protocol can be implemented in any biological laboratory with an adequate fume hood. The equipment and materials are readily available from standard scientific suppliers, with the most specialized equipment being a vacuum pump and desiccator chamber.
Although this procedure presents several advantages compared with other sample preparation protocols, some limitations should be noted.
Fixation and dehydration of the sample
The timing of the different steps of the protocol involving fixation and degassing are heavily dependent on the composition and size of the tissue. In the present procedure, we give examples of timing for different human organs; whereas timing for other types of tissues would need to be tested and verified. Some guidelines for optimization of timings are provided in this protocol.
Whilst fixation of the organ is critical for long-term tissue preservation, it alters tissue mechanical51,52 and diffusion properties, which could confound other measurements. Despite increasing the image contrast, ethanol dehydration also affects the mechanical properties of the tissue43,53. This limits the use of this preparation method for in situ testing; however, by not fixing the tissue and replacing ethanol with water, it is possible to use in situ imaging13,54. Thus, the protocol could be expanded in the future to cover dynamic experiments and quantification of mechanical properties over a short timeframe (as biological degradation would occur). If the contrast provided by ethanol is not high enough, or not adapted to the experimental needs, various contrast agents, such as iodine-based55 or tungsten-based contrast agents56 could be used to increase the overall contrast of the tissue, or resolve specific components of interest13,56. If agar gel was not desirable for a particular application, it could be replaced with other solid or elastic media that would be in equilibrium with the mounting liquid and relatively amorphous in its structure. For instance, in case of mounting with 96% ethanol, transparent candle crystal gel can be used instead of agar gel. In case of mounting with water, gelatin blocks or polyacrylamide blocks can also be used.
X-ray dose limit
One of the main drawbacks of wet embedding methods is the formation of bubbles due to dose rate or dose accumulation in the case of sCT57. These bubbles can move or damage the sample, in addition they can create strong artifacts, dramatically reducing the image quality58. Although bubbling is an issue with other standard preparation methods, e.g., paraffin embedding, it is not present with critical-point drying. In this protocol, the issue is mitigated by multiple degassing steps during the procedure, delaying the nucleation of bubbles during the scan and further mitigated by the use of high-energy X-rays with strong phase contrast. However, only a few beamlines are equipped to image soft tissue at high energy with sufficient coherence properties and propagation capabilities. Spatial coherence must be high enough that the propagation distance can be set to distinguish the density variation of the sample without geometric blurring. For low-energy X-ray tomography, the degassing steps must be performed conscientiously, and the dose rate controlled even more carefully because of the photodissociation of the water molecule.
Experimental design
Sample collection and fixation
Biological tissue can be collected by various methods. If the sample is collected directly from a biobank, surgery or dismissed transplantation (Step 1B), it can be taken directly to the sample fixation stage (Step 2). If the organs come from a donated body, organ extraction must be performed (Step 1A(ii)). Embalming of the body is carried out shortly after death, before organ extraction (by a licenced practitioner). The body is fixed by injecting formalin diluted in a solution containing lanolin into the right carotid artery (Step 1A(i)). After evisceration, complete fixation of the organ is ensured by immersing it in 4% neutral buffered formaldehyde (Step 2). The duration of fixation is defined by the size of the organ, e.g., 4 d for a human brain. When eviscerating the organ, ensure as much surrounding tissue as possible is removed to decrease the time of penetration of the fixative into the organ. The volume of fixative is also important, we recommend a volume at least four times the tissue volume.
Some organs, such as the lung, may require inflation. This can be partially accomplished at the fixation stage by using the instillation of formalin in the lungs under controlled pressure59. The lung is perfused with 4% formalin through the trachea using a 30 cm water column, the trachea can then be ligated to maintain the inflated configuration over a period of 2 d. The lung is subsequently immersed in a 4% formalin solution after extraction. Once fixed, the replacement of formalin by the successive baths of ethanol with vacuum pumping is coupled with the injection of ethanol in the bronchia to maintain a consistent 3D shape. Strictly controlled pressure with ethanol degassing is currently not possible with the proposed protocol, yet should become possible using the circulation of degassed ethanol instilled at a controlled pressure. Our results show that, even without strictly controlled inflation at the ethanol dehydration stage, the data still have high scientific utility.
Organ dehydration and degassing
The organ is dehydrated with multiple pre-degassed ethanol baths. The transition to 70% ethanol must be gradual to avoid shrinkage43. An ethanol concentration of 70% was chosen as the best compromise between controlled shrinkage and sufficient contrast to observe structures of interest using phase contrast; however, a different final ethanol concentration can be used depending on the application. During this step, degassing has to be performed to remove free and dissolved gas present in the tissue to avoid bubble formation or volume increase during imaging. For human organs, we present two methods, depending on the fragility of the organ for dehydration and degassing. The vacuum degassing (Step 4A(i–v)) can be used for most organs (heart, lung, liver, kidney, spleen). The organ is immersed at room temperature (20 °C) in four successive pre-degassed ethanol baths of concentrations of 50%, 60% and 70%, and then a second 70% bath to ensure equilibrium is reached. A degassing step is performed between baths. Equilibrium is generally reached in 4 d for each concentration for a human organ such as a lung or a heart. Degassing is performed with a vacuum pump and a desiccator by successive cycling down to an absolute pressure between 15 and 10 mbar. The degassing is considered suitable when no strong bubbling can be observed at 10 mbar. Alternatively, thermal cycling (between room temperature and 4 °C) (Step 4B(i)) was developed for fragile organs, such as human brain, as some damage was observed after using the vacuum degassing method36 if the bubbles were not able to find a way out of the brain. In this method, four thermal cycles are performed by immersing the organ in four successive baths of 50%, 60% and 70%, and a second 70% pre-degassed ethanol. Each thermal cycle consists of immersing the organ in the highly degassed ethanol bath at room temperature. The container has to be closed with care to avoid entrapping bubbles of air. It is then kept in a refrigerator for 4–5 d at 4 °C. During this period, the dissolved gas will diffuse into the surrounding ethanol, and the bubbles will progressively dissolve. After this time, the solutions and organ must be brought back to room temperature, and a new cycle can be started using a new strongly pre-degassed ethanol bath. For both methods, the minimum number of ethanol baths is four to reach a final concentration of 70% without having substantial shrinkage. The immersion times have to be adapted and optimized to the type and fragility of the organ. The result of the degassing can be tested by making a radiograph of the organ in its jar without the mounting media described hereafter. If some remaining bubbles are still visible, more thermal cycles can be performed with ethanol at 70%. Organs with adipose tissues, such as the brain, require a longer time to equilibrate with ethanol. At each stage, dehydration can be checked by disturbing the container with the organ inside and looking for streaks of different density (different transparency) forming in the surrounding ethanol solution, this indicates the presence of water in the ethanol.
Ethanol dehydration is sufficient to observe the structures of interest with µCT and sCT when using phase contrast; however, it should be possible to combine this protocol with the use of a contrast agent to resolve specific components of the sample or when using less sensitive imaging techniques56. The contrast agent must be miscible with 70% ethanol, or should be applied to the fixed organ before the ethanol dehydration. Contrast agents would increase the absorption of the sample, which may enhance bubble formation during imaging if using intense synchrotron X-ray beam.
In cases where the characterization technique is not be compatible with ethanol (for instance, MRI for diffusion), the same protocol for degassing and then for mounting can be applied using water with formalin at the desired concentration. For in situ applications requiring close-to-biological conditions, this protocol can also be used with water only, but the samples can be used for a only few hours as biological degradation would occur. Specific safety aspects (working under an adapted fume hood in a well-ventilated laboratory, wearing gloves, a laboratory coat, closed shoes and safety glasses) have to be taken as solutions of formalin or ethanol produces dangerous vapors.
Bubble formation
One of the main problems in wet embedding is the presence of bubbles. These can come from the mounting protocol (bubbles in the organs or trapped in the mounting media) and/or from very high X-ray doses resulting in evaporation of the ethanol (in case of sCT). In both cases they can create substantial artifacts (Fig. 3), and in case of bubbling from the X-ray dose, the movements of the bubbles during the scanning often renders the scans unusable60,61. Preliminary tests showed that degassing the sample before imaging removes trapped bubbles and delays the nucleation and growth of new bubbles. Hence, several degassing steps were incorporated in the protocol to mitigate the bubble formation.
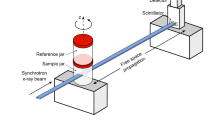
a–d, Artifacts due to the radiation dose (a,b) or to a stable bubble entrapped during mounting (c,d). a, A cross-section of a human liver with numerous bubbles that have developed due to too high a dose uptake due to a scan crashed with the beam on for several hours, causing artifacts in the images. ba, bubble artifact. b, A cross-section of the liver imaged after removal of bubbles by degassing the sample with vacuum degassing. c, A cross-section of a human brain with a bubble entrapped during mounting. d, A zoom on the bubble trapped inside the brain. The images were obtained at BM05 beamline at ESRF. All experiments followed the relevant governmental and institutional ethics regulations for human experiments.
Organ mounting
The purpose of this step is to maintain the organ in position during the scan to avoid motion artifacts and to ensure that more scans can be performed at a later stage with good 3D rigid registration between subsequent scans/experiments. The agar gel cannot be prepared with 70% ethanol directly, but must be done by slowly adding powder of agar agar in agitated hot demineralized water (~85 °C) up to 20 g/L. Once fully dissolved, the agar agar solution is poured into a suitable container and let to cool down for several hours. Once gelled, the mixture can be cut into small cubes (Step 8) and added to 96% ethanol (6 L:2 L ethanol:agar agar) (Step 9), ensuring a final ethanol concentration of 70%. After degassing (Step 12) and crushing of a part of the agar cubes (Step 14), the mixture is ready for mounting. If mounting with formalin is preferred to ethanol (e.g., to improve contrast with MRI), the 96% ethanol bath can be replaced with a 4% formalin bath. The agar gel can be prepared in advance and stored in an airtight container to prevent the reintroduction of gas into the solution after its degassing. The amount prepared depends on the size of the organ and container used for final mounting. An agar gel prepared with 70% ethanol ensures a drastic reduction of bubble formation during imaging. It is important to use a crushed agar gel and not a blended agar gel, as the blended gel does not hold the sample as firmly as the crushed agar gel and could lead to movement during imaging. Initially, only agar cubes were used to hold the sample in position; however, the cubes were found to be too rigid, creating deformations where they contacted the surface of soft organs such as lungs. Thus, the cubes should only be used at the bottom and top of the container. A few centimeters are used to create a solid base and avoid rotation of the sample (Step 16). Crushed agar is used in the remainder of the container to maintain the sample in position. The mixture of crushed agar in the mounting liquid (70% ethanol or 4% formalin) must be added to the container gently with a ladle to avoid gas bubble entrapping during the process (Fig. 4c). Rapid vacuum degassing should be performed at least three times when adding the agar crushed gel to remove entrapped bubbles. These vacuum cycles should be performed ideally down to 15 mbars. The dimensions of the container should be as close as possible to the specimen to minimize the amount of material that the X-ray beam has to pass through; however, the organ should not touch the side of the container to avoid artifacts in the images that would compromise the accuracy of the reconstruction (i.e., a minimum of 5 mm of crushed agar gel should surround the organ to avoid direct contact with the container). For sCT, the container used for the mounting must be made of a material resistant to X-rays and not too dense, such as polyethylene terephthalate. Glass should be avoided as its high density compared with the samples would result in strong absorption contrast artifacts. Once the agar gel has been compacted around the sample (Step 22) and properly degassed, the container can be sealed with a liquid-tight lid (Step 28). The mounting should be assessed to ensure that no movement of the sample in the container is possible and no bubbles can be seen inside. If some bubbles are entrapped, rapid vacuum degassing can be used to remove them. The same approach can be used to remove bubbles in an organ in case of a bubbling event due to high dose during sCT scanning (do not degas the sealed container, remove the lid, put a flat rigid sieve on top of the agar gel to prevent movement of the organ and then degas, complement the ethanol level if it decreases and eventually add a small amount of crushed agar gel, then close again). An example of insufficiently compacted agar is shown in the online Supplementary Video 1. With the organ fixed and placed in 70% ethanol, the sample can be stored for years (Fig. 4d).
a, Degassing of the organ in its container using a desiccator and a vacuum pump. b,c, After compaction of the agar gel present in the container, some of the ethanol is removed using a syringe and a sieve (b), then agar gel is added (c). This process is continued until the agar gel is sufficiently compressed around the sample to hold it in position. d, Mounted sample stabilized and degassed in the agar gel, ready to be scanned. e, Schematic of a mounted brain, like in d, fixed with the crushed agar gel and agar cubes, in a sealed container. All experiments followed the relevant governmental and institutional ethics regulations for human experiments. Panel e image of brain from Smart Servier Medical Art (https://smart.servier.com/) licensed under a CC BY 3.0 license.
sCT imaging and reconstruction
The X-ray imaging and 3D reconstruction are not described in detail in this protocol; however, a detailed description is available in the literature36 or by contacting the corresponding authors. This method was developed for large-organ imaging with HiP-CT but many aspects would be beneficial for more classical sCT and μCT (laboratory source) imaging on smaller samples, including the immobilization and contrast enhancement linked to ethanol preparation. Once the sample is mounted, scanning can be performed at any time, with typical sCT imaging results of an intact human heart shown in Fig. 2. The main limitation of this technique is the nucleation and growth of bubbles following the application of high-dose X-ray to the organ in the case of sCT. If only a few bubbles appear during imaging, the sample can be left in a refrigerator at 4 °C to dissolve them again. If this is not successful, the sample can be re-degassed with vacuum pumping without having to dismount, but there will inevitably be some small movements of the sample during the process, which complicates the multiresolution scanning and registration procedure. Several solutions exist to avoid bubble formation during sCT, for the optimization of detection, the increase of relative contrast to work with lower dose, a better control of the dose, better data processing or having resting period for the samples between successive scans of the same area. Bubbling seems to be strongly linked to the thermal effect of dose on the evaporation rate of the ethanol. Developing sample environmental chambers to work at a lower temperature and ensuring a better thermal equilibrium may be an efficient way to alleviate the risk of bubbling linked to the use of 70% ethanol, especially for multiple submicron resolution scans on the same organ.
MRI imaging
The sample preparation method presented in this protocol is compatible with MRI (Fig. 5c), making it highly amenable to multimodal studies. However, a number of considerations should be taken into account before deciding on the sample preparation method and scanning order. Dehydration of the sample with ethanol increases the contrast for X-ray imaging, but may affect the MR images62. The majority of MR techniques are highly dependent on water content. Thus, by dehydrating the sample, the contrast between tissues that retain water well (e.g., adipose tissue) and those that dehydrate efficiently (e.g., muscle tissue) is enhanced. However, by decreasing the total water content, the signal is reduced and some techniques such as diffusion-weighted MR are no longer possible. One method of overcoming this problem is to not dehydrate the sample but to prepare the sample directly in a formalin mounting for performing MRI first. The contrast for sCT images is then reduced, but the signal-to-noise ratio of MRI images is increased. Although formalin- or paraformaldehyde-fixed tissue is the most used sample preparation method for ex vivo MRI, this technique also affects the quality of the images63. The fixation of tissue with formalin or paraformaldehyde reduces T1, T2 and diffusivity of tissue, hence reducing the contrast‐to‐noise in images. One approach is to wash the brain to remove fixative before MRI imaging, which can restore the pre-fixation T2 values though not the T1 (ref. 64). Washing will also compromise the preceding degassing steps, so the relative importance of each imaging modality must be carefully weighed against the overall imaging pipeline and multimodal registration requirements.
a, 3D view of the human kidney. b, Cross-section of the kidney imaged with medical CT, with a voxel size of 625 µm. c, Cross-section of the kidney imaged with a 3T medical MRI, with a voxel size of 220 × 200 × 200 µm3. d, Cross-section of the kidney imaged with sCT, with a voxel size of 25 µm on the BM5 ESRF beamline. e, Comparison of sCT and a histopathological section stained with hematoxylin and eosin (H&E). Histology was performed after all other imaging modalities. Histological techniques used are standard and well documented in ref. 72. The left column shows light micrography images of H&E-stained histopathological sections, the right column shows 2D sCT images taken at BM05 beamline at ESRF with a voxel size of 1.4 μm. Images with a star on the top left have been pseudocolored (from sCT) or converted to gray levels before being inverted in contrast (from histology). All the basic information on the patient from whom this organ originates is provided in Extended Data Fig. 2. All experiments followed the relevant governmental and institutional ethics regulations for human experiments. Panel e adapted with permission from ref. 36, Springer Nature America, Inc.
Expertise needed to implement the protocol
The protocol outlined here requires prior experience in handling human or animal organs. If human organs are used, a pathologist should be part of the study to handle body embalming and organ dissection. The degassing of the organ, the preparation of the agar solution and the mounting of the organ require the expertise of a laboratory technician, as these steps involve the handling of hazardous materials such as formalin and ethanol solution. This protocol was developed and optimized for imaging large organs, a scientist who has experience in CT or sCT is required as imaging such large structures is not trivial.
Regulatory approvals
All the experiments herein were authorized by the ESRF, the Laboratoire d’Anatomie des Alpes Françaises and the Hannover Institute of Pathology at Medizinische Hochschule, Hannover (ethics vote no. 9022_BO_K_2020). Transport and imaging protocols were approved by the Health Research Authority and Integrated Research Application System (200429) and the French Health Ministry. All organ dissections respected the memory of the deceased. The post-mortem study was conducted according to the Quality Appraisal for Cadaveric Studies scale recommendations65.
Institutional and governmental ethics regulations concerning use of human tissue in research must be sought and followed before undertaking any procedures described in this protocol. The specific requirements will be set by the relevant authorities. Typically, after identifying a suitable sample, either from a body donation or from a biobank, an ethics committee should review the project. Once the project is accepted, contracts and/or Material Transfer Agreements (MTAs) must be established to transfer the biospecimens between institutions. The time taken to obtain the ethical agreement and the MTA can be considerable (several weeks to years), depending on the countries involved and the specificities of the contract.
Materials
Biological material
-
Human organs obtained from a donated body
Caution
Institutional and governmental ethics regulations concerning the use of human for scientific research must be followed.
Reagents
-
4% neutral buffered formalin, (50% Hygeco, cat. no. 07040 + 50% Hygeco, cat. no. 07020; diluted to 4%)
Caution
Toxic, carcinogenic, mutagenic, reprotoxic and flammable. Handle with care, in a fume hood. Avoid inhalation and exposure to the skin or eyes.
-
Ethanol 96% (Cheminol France, cat. no. 20× 4-2)
-
Agar Agar (Nature et Aliments, cat. no. 510190)
Critical
The gelling properties of agar differ depending on the producer. We recommend using the supplier listed to reproduce the results presented here. If using others, some preliminary tests would be required.
-
Demineralized water (Sigma Aldrich, cat. no. 38796)
Equipment
-
Fume hood (Iberis laboratoire, cat. no. SPR18)
-
Gloves Nitril (Showa, cat. no. 7500PF)
-
Paper towels (Paredes, cat. no. 404857)
-
Dissecting instruments (Fisher Scientific, cat. no. 11738551)
-
Vacuum pump (Marshall Scientific, cat. no. VME8)
-
Vacuum dessicator (SP Bel-Art, cat. no. F42400-4031)
-
Refrigerator (Fisher Scientific, cat. no. 22651264)
-
Large leak-proof container in polyethylene terephthalate (Medline Scientific, cat. no. 129-0592, or Lock & Lock, cat. no. INL-403 or Lock & Lock, cat. no. INL-203, shown in Extended Data Fig. 4)
-
Magnetic vortex system equipped with a heating plate (Fisherbrand, cat. no. 15349654)
-
Electric grater (Seb, cat. no. DO201141)
-
Ladle (Tefal, cat. no. K1180214)
-
Long knife (Ikea, cat. no. 402.947.22)
-
Syringe 50 ml (PentaFerte, cat. no. 002022960)
-
Sieve (Profilstore, cat. no. toilinox_PGRI1ME213)
-
General purpose vinyl tape (3 M, cat. no. 764)
-
Drill (DeWALT, cat. no. DCD791P2-QW)
-
Container holder (custom-made, described in Extended Data Fig. 5 and in ref. 36)
-
Synchrotron microtomography setup. We used the BM05 and BM18 beamlines of the ESRF. The beamline parameters of all the organ scans presented in this paper are detailed in Extended Data Fig. 6.
-
Data reconstruction: Dell EMC PowerEdge R7525 Rack Server (operating system: Ubuntu 20.04.3 LTS; central processing unit: AMD EPYC 75F3 2.95 GHz, 64 cores; memory: DDR4-3200 32 GB (32×); graphics processing unit: NVIDIA Ampere A40 ×2; hard drive: 2.4 TB 10 K RPM SAS 12 Gbps)
-
Data visualization: Dell Precision 7920 Tower (operating system: Windows Server 2016; central processing unit: Intel Xeon Platinum 8160 M processor, 2.10 GHz, 24 cores; memory: 1.5 TB; graphics processing unit: NVIDIA Quadro P6000; hard drive: AVAGO MR9460-16i SCSI 4.67 TB)
-
Reconstruction code written in house using MATLAB 2017 available on GitHub (https://github.com/HiPCTProject/Tomo_Recon)
-
PyHST2 (3D tomographic reconstruction code, http://ftp.esrf.fr/scisoft/PYHST2, open source) developed by the ESRF
-
ImageJ/Fiji (data pre- and postprocessing, https://imagej.nih.gov/ij/, http://fiji.sc/, open source)
-
VGSTUDIO MAX version 3.5 (volume segmentation and visualization, https://www.volumegraphics.com/en/products/vgsm.html)
Procedure
Organ procurement
Caution
Any experiments involving the use of human organs must be ethically approved by the relevant institutional or governmental committees.
-
1
Collect the organ of interest either by extracting it from a donated body (option A) or through a biobank, a surgery or a dismissed transplantation (option B).
-
(A)
Embalming of the body and organ extraction
Timing ~3 d
-
(i)
(Optional) Embalm the body after death by injecting sequentially 4,500 ml formalin diluted to 1.15% in a solution containing lanolin and 4,500 ml formalin diluted to 1.44% into the right carotid artery (performed by a licenced medical practitioner), ideally with a jugular draining.
Caution
Formalin is volatile and highly toxic. It is an eye, respiratory and skin irritant and a probable carcinogen. Work inside a chemical fume hood in a well-ventilated area and wear appropriate personal protective equipment (gloves, eye protection and laboratory coat).
Pause point
The body can be stored up to 2 years if properly fixed and refrigerated.
-
(ii)
Extract the organs of interest from the body.
-
(iii)
Remove surrounding fat and connective tissue.
-
(i)
-
(B)
Organ procurement from a biobank
Timing depends on the administrative and delivery delays
-
(i)
Retrieve the organ from a biobank, a surgery or a dismissed transplantation. The time between retrieval of the organ and formalin fixation should be minimal. In the meantime, the organ should be kept at 4 °C to avoid degradation of the tissue. Donor data should be anonymized in accordance with governmental regulations. The organ should be transported in a sealed container.
-
(i)
-
(A)
Organ fixation
Timing 4 d
-
2
(Optional) Fix completely the organ by immersing it in 4% neutral buffered formalin at room temperature. We used 4 d for fixation by formalin to comply with safety rules linked to autopsies in case of pathogenic agent (such as COVID-19). We recommend the formalin volume to be at least four times greater than the volume of the organ. The volume is estimated by the pathologist, for whom this is standard practice.
Preparation of the organ before mounting
Caution
Degassing with a vacuum pump should be carried out under a fume hood or in a room equipped with a ventilation system of sufficient capacity to avoid inhalation of formalin or ethanol gases.
Critical
Room temperature was set typically at 20 °C.
-
3
(Optional) Wash the sample with tap water for 2 min to remove the fixative.
-
4
Prepare the organ for mounting by dehydration using multiple baths of ethanol and degassing it under vacuum (option A). With the vacuum degassing protocol, damage could be seen in some fragile organs (typically the brain). Thus, an alternative protocol to prepare the sample without vacuum degassing was developed using thermal cycles (option B). The ethanol can be replaced with 4% formalin solution to avoid the dehydration of the sample where this is undesirable e.g., MR imaging or in case of X-ray imaging with contrast close to biological conditions. Modifications to this procedure may be necessary as timing and number of cycles depend heavily on the composition and size of the organ. For samples of different origin, type and size, some optimization of this step should be considered.
-
(A)
Dehydration and vacuum degassing
Timing 16 d
-
(i)
Immerse the organ in a bath of pre-degassed 50% ethanol. The solution must be at least four times the volume of the organ.
-
(ii)
Place the container with the organ inside a desiccator
-
(iii)
Remove the free and dissolved gas in the tissue by diminishing the pressure by successive cycles using a diaphragm vacuum pump until strong bubbling occurs.
-
(iv)
The first cycles are typically 2–3 min long and reaches 15–30 min after three to four cycles for a human organ, such as a heart or a kidney.
Critical step
these times are strongly dependent on the user’s vacuum configuration.
-
(v)
Continue these cycles until reaching 15–10 mbar without strong bubbling.
Critical step
The degassing should be performed in cycles of increased duration. For each cycle, the vacuum pumping is stopped when bubbling becomes suddenly stronger in intensity. Each time a new cycle starts, the time before first bubble formation should increase. The degassing time has to be adapted to each organ depending on its composition and size. There is no typical time, it has to be chosen empirically by looking at the bubbling regime. This will strongly depend on the vacuum pump and desiccator quality.
-
(vi)
Wait until equilibrium (our tests show that typically 4 d for a human organ such as lung, heart or brain is sufficient for each ethanol concentration; with 2 d only, we saw incomplete equilibrium visible as density gradients in the scans).
-
(vii)
Repeat Steps (i–vi) with three successive baths of pre-degassed ethanol at 60%, 70% and finally 70% with a degassing between each bath.
-
(viii)
Perform a final degassing of the organ for few minutes just before mounting it with the agar-agar crushed gel in the mounting jar.
-
(i)
-
(B)
Dehydration and thermal cycles degassing (alternative protocol for fragile organs)
Timing 27 d
-
(i)
At least four thermal cycles must be performed each in successively higher concentrations of 50%, 60%, 70% and 70% ethanol.
-
(ii)
For each successive concentration, use the vacuum pump to degas a volume of ethanol of at least four times the volume of the organ, at room temperature.
-
(iii)
Add the ethanol to the container with the sample and close the container with care to avoid entrapping bubbles.
-
(iv)
Store at 4 °C for 5 d. The dissolved gas will diffuse into the surrounding ethanol, and the bubbles will progressively dissolve. The immersion times have to be adapted and optimized to the type and fragility of the organ.
-
(v)
For each cycle, remove the container from the refrigerator to bring it back to room temperature (~12 h). A thermometer can be used to measure the temperature of the solution.
Critical step
The result of the degassing can be tested by making a radiograph of the organ in its jar without the mounting media described hereafter. If some remaining bubbles are still visible, more thermal cycles can be performed with ethanol at 70%. Organs with adipose tissues, such as the brain, require a longer time to equilibrate with ethanol.
Pause point
Once at 70% ethanol, the organ can be stored at room temperature for months to years before continuing with the mounting steps of the protocol.
-
(i)
-
(A)
Preparation of the organ mounting gel
Timing 12 h to 2 d
Critical
The following produces 5 L of agar gel:
-
5
Boil 5 L of demineralized water in a container using a hot plate equipped with a magnetic vortex system.
Caution
Boiling water or steam can cause severe burns. Never carry the full container by hand. Instead, roll it on a cart or dolly. Use safety equipment (gloves, a laboratory coat, closed shoes and safety glasses) and work in a well-ventilated room.
-
6
Once above 80°, slowly pour 100 g of agar agar (20 g/L) powder in the water and keep the vortex until good dissolution.
-
7
Once dissolved, stop the agitation and remove the stirrer with a ladle.
-
8
Pour the liquid into a large container. Typically, we use a container of 40 × 30 × 12 cm.
Caution
Boiling water or steam can cause severe burns. Use safety equipment (gloves, a laboratory coat, closed shoes and safety glasses) and work in a well-ventilated room.
-
9
Once gelation has been achieved (~12 h depending on the temperature), cut the agar gel with a long knife in cubes of ~2 cm3.
Caution
Use knives carefully and always concentrate on what you are doing.
-
10
Immerse the cubes in 11.7 L of 96% ethanol in a 20 L container. The volume ratio is chosen to ensure a final ethanol concentration of 70%. If the organ was prepared with 4% formalin solution instead of ethanol to avoid the dehydration of the sample, replace the 96% ethanol with 4% formalin.
Caution
Ethanol is flammable, keep away from heat sources and always have a fire extinguisher close at hand. Handling of ethanol or formalin should always be carried under a fume hood. Avoid inhalation and exposure to the skin or eyes. Use safety equipment (gloves, a laboratory coat, closed shoes and safety glasses).Wait until the density of the blocks is close to that of ethanol (~24 h).
-
11
Check the equilibrium of the solution by agitating the solution to put the gel cubes in suspension and ensure that they sink slowly to the bottom of the container (over several dozen of seconds).
-
12
Place the container with the agar cubes and ethanol in the desiccator. To close the desiccator, place the lid and close slowly by applying a mild force. Carefully twist the lid in both directions to ensure an airtight seal. Make sure the desiccator is connected to the vacuum pump.
-
13
Degas the solution using the vacuum pump for two cycles of ~1 h to avoid cracking the agar cubes. The cubes should be kept submerged during the process to avoid dehydration. Avoid exposing the cubes to air for more than 10 min. Store a third of the degassed cubes with ethanol 70%, in an air-tight container.
-
14
Crush the remaining cubes using an electric grater.
Caution
Use of electric equipment with 70% ethanol is a fire risk, the electrical device in question should have a motor with spark protection and appropriate fire regulations followed.
-
15
Store in an airtight container for future use with enough 70% ethanol solution to ensure that no agar is out of the solution,
-
16
Degas the solution again just before use.
Organ mounting and degassing
Timing ~3 h
-
17
Fill the bottom of the leak-proof cylindrical container with agarose cubes a few centimetres in length/width. The container is shown in Extended Data Fig. 4.
-
18
Fill half the container with crushed gel.
Critical step
Use a ladle with careful slow movements when manipulating the agar solution to avoid increasing the dissolved gas in the solution or entrapment of bubbles.
-
19
Carefully immerse the organ in the gel and place it in the desired position for imaging. Cover the organ with crushed agar.
-
20
Degas the whole container to remove entrapped bubbles.
Critical step
This degassing step only aims at removing entrapped bubbles. As all the components were degassed before, this should limit the amount of dissolved gas. This vacuum degassing should be done only with short pumping times (2–3 min) for several cycles to help remove the visible bubbles.
Critical step
Gentle tapping on the desiccator can help bubbles travel to the top
-
21
Add more of the agar–ethanol solution.
-
22
Use a sieve to press on the agar gel from the top to compact it around the organ. Use a syringe to remove the excess ethanol from on top of the sieve, then add a volume of agar and ethanol equivalent to one-quarter of the container.
Critical step
Be careful as once the agar is compact, the bubbles cannot easily rise to the top, and therefore degassing can no longer be carried out on this part of the container. All movements have to be done slowly and carefully to ensure not to entrap bubbles when compacting the agar.
-
23
Manually apply vertical pressure around the sample to compact the agar around the sample and maintain the sample in position until it cannot move anymore in the jar.
Critical step
At least 5 mm of crushed agar gel must be left between the sample and the container wall to avoid border effects during X-ray imaging.
-
24
Degas the whole container using a vacuum pump to remove the gas added during the last three steps and to avoid trapping bubble in the agar gel. Use cycles to facilitate the escape of bubbles. After a few cycles, if no bubbles are visible on inspection, proceed to the next step.
-
25
Repeat the last four steps until the agar is compact enough below, around and above the sample to avoid any movement of the specimen. An example of insufficiently compacted agar is shown in the online Supplementary Video 1.
-
26
Fill the leak-proof container to the top with the agar–ethanol solution and finish filling with some agar cubes with solution to ensure good sample blocking.
-
27
Drill a small hole of a few millimetres in diameter in the center of the lid.
-
28
Screw the lid on the container.
Critical step
If the container is not directly leak tight, use sealing material such as parafilm latex or silicon rubber on the container thread to ensure a good seal.
-
29
After sealing, check that all air was removed from the container by applying a slight pressure on the lid and ensuring that ethanol comes out of the hole.
Critical step
Agar can block the hole and prevent ethanol from coming out, in which case use a needle to clear the hole
-
30
Apply a piece of tape onto the hole on the lid to avoid gas exchange between the inside and outside of the container, and to act as a safety exhaust in case of bubbling event during scanning.
-
31
Apply tape around the lid to protect it and prevent it from being accidentally opened.
-
32
Assess that no bubbles remain in the container. Once the agar is compact, the bubbles cannot travel easily to the top.
Pause point
The mounted organ can be stored at room temperature for months to years before imaging, and can be imaged many times without any intervention as long as there is no bubbling event due to too high an X-ray dose.
sCT 3D imaging of the mounted sample
Timing variable depending on the imaging setup
Caution
-
33
In case of refrigerated, mounted organs, take the organ out of the refrigerator at least 12 h before the experiment and leave at room temperature to avoid any change in shape during imaging due to temperature change.
-
34
Place the organ in its sealed container on the container holder. The custom-made container holder we are using is described in Extended Data Fig. 5.
-
35
Place a second equivalent container (called the reference container) above the sample container, filled only with the appropriate mounting media, such as ethanol–agar gel.
-
36
Setup the detector with a pixel size that enables the whole organ to be imaged. Typically for a brain of 14 cm with a camera of 5,056 pixels horizontally, a maximum pixel size of 27.7 µm can be achieved in normal acquisition.
-
37
Align the detector with the X-ray beam.
-
38
Adjust the beamline slits by setting their aperture around the field of view.
Critical step
Well-fitted slits reduce the dose deposited on the sample.
-
39
Align the sample with the detector and find the center of rotation.
-
40
Tune the energy and flux to ensure a sufficient penetration through the sample. The optimal average energy depends on the size of the jar containing the organ. Typically, 80 keV for 10 cm, up to 110 keV for 15 cm, especially if dense parts such as cartilage or calcifications are visible on the radiographs. The flux can be tuned to adjust the scanning speed to the experiment requirements while being in the correct energy range and keeping a sufficiently low dose rate to avoid bubbles.
Critical step
Care must be taken regarding the dose deposited on the sample to avoid bubble formation during the scan. This step should be performed by a trained beamline scientist. Furthermore, this step is highly dependent on the type and size of the sample, as well as the characteristics of the beamline and on the quality of the degassing of the sample.
-
41
Set the exposure time and accumulation of the camera. Typically, for a camera signal coded in 16 bits, a maximum pixel value of 42.000 per single subframe enables a strong security margin to avoid saturation, while using the full dynamic of the camera.
Critical step
Increasing the exposure time increases the scan time and therefore the dose deposited on the sample.
-
42
Ensure that the camera dynamic is optimized, and that no saturation of the detector occurs during the scan.
-
43
Perform a scan of the reference container.
-
44
Scan the complete organ.
Critical step
The beamline setup for the reference scan and sample scan must be identical.
-
45
Create a flat field by doing an average of all the projections of the reference scan performed in Step 44.
-
46
Perform a single-slice reconstruction by applying the flat field calculated in Step 46 and using a filtered back-projection algorithm66 coupled with a single-distance phase retrieval to ensure the quality of the scans (reconstruction can be performed using an open-source tomographic reconstruction code, such as PyHST2 (ref. 67) or TomoPy68).
-
47
(Optional) Select features of interest on the reconstructed images.
-
48
(Optional) Calculate motor positions and align the sample for imaging the corresponding region of interest.
-
49
(Optional) Repeat Steps 37–47 for imaging selected regions in the organ.
-
50
Create partial angular integration from the reference scan every 100 projections for flat-field correction of sample scan radiographs.
-
51
Apply the flat-field correction on every 100 projections of the sample scan with the corresponding flat field calculated in Step 50.
-
52
Reconstruct the 3D volume using a filtered back-projection algorithm coupled with a single-distance phase retrieval66 and a 2D unsharp mask (reconstruction can be performed using an open-source tomographic reconstruction code, such as PyHST2 (ref. 67) or TomoPy68).
-
53
(Optional) Correct ring artifacts on reconstructed slices using the updated Lyckegaard et al.69 algorithm (code available on the GitHub indicated in the Code availability section).
Multimodal imaging of the mounted sample
Timing variable depending on the imaging technique
Troubleshooting
Troubleshooting advice can be found in Table 1.
Timing
-
Step 1, organ procurement: ~3 d
-
Step 2, fixation of the organ: 4 d
-
Steps 3–4A or 3–4B, preparation of the organ before mounting: 16–27 d
-
Steps 5–16, preparation of the organ mounting gel: 12 h–2 d
-
Steps 17–32, organ mounting and degassing: ~3 h
-
Steps 33–53, sCT imaging: timing depends on the imaging technique and setup
-
Steps 54–55, imaging and histology: timing depends on the imaging technique and setup
Anticipated results
This protocol provides a method to prepare and stabilize large organs or biological samples for imaging at high resolution using sCT, µCT, clinical CT or MR imaging. Typical images from human organs are shown in Figs. 2 and 5. This sample preparation procedure provides high-contrast images (depending on the imaging modality), prevents sample movement during scanning, is compatible with multiple imaging modalities and preserves the morphological characteristics of the tissue compared with other sample preparation methods such as paraffin embedding. The sample can be stored for at least 1 year without movement or deterioration. This method enables the 3D investigation of large biological structures such as human organs without damaging them. These high-resolution images can provide qualitative and quantitative information on the healthy or pathological characteristics of an organ, for instance the effect of the COVID-19 in human lungs36.
Data availability
Image data used to create the figures present in this protocol paper are publicly available from the ESRF data repository (https://human-organ-atlas.esrf.eu) or from the corresponding authors.
Code availability
The sCT data were reconstructed using a custom code written in MATLAB 2017 available on GitHub (https://github.com/HiPCTProject/Tomo_Recon) and the software package PyHST2 (https://software.pan-data.eu/software/74/pyhst2). VGSTUDIO MAX 3.5 (Volume Graphics) was used for volume rendering.
References
Alho, E. J. L. et al. High thickness histological sections as alternative to study the three-dimensional microscopic human sub-cortical neuroanatomy. Brain Struct. Funct. 223, 1121–1132 (2018).
Kofler, L. et al. Three‐dimensional histology vs. serial section histology in the treatment of primary basal cell carcinoma: a randomized, prospective, blinded study of 569 tumours. J. Eur. Acad. Dermatol. Venereol. 35, 1323–1330 (2021).
Pichat, J., Iglesias, J. E., Yousry, T., Ourselin, S. & Modat, M. A survey of methods for 3D histology reconstruction. Med. Image Anal. 46, 73–105 (2018).
Eberle, A. L. & Zeidler, D. Multi-beam scanning electron microscopy for high-throughput imaging in connectomics research. Front. Neuroanat. 12, 112 (2018).
Hildebrand, D. G. C. et al. Whole-brain serial-section electron microscopy in larval zebrafish. Nature 545, 345–349 (2017).
Miranda, K., Girard-Dias, W., Attias, M., de Souza, W. & Ramos, I. Three dimensional reconstruction by electron microscopy in the life sciences: an introduction for cell and tissue biologists. Mol. Reprod. Dev. 82, 530–547 (2015).
Tainaka, K. et al. Whole-body imaging with single-cell resolution by tissue decolorization. Cell 159, 911–924 (2014).
Zhao, S. et al. Cellular and molecular probing of intact human organs. Cell 180, 796–812.e19 (2020).
Cai, R. et al. Panoptic imaging of transparent mice reveals whole-body neuronal projections and skull–meninges connections. Nat. Neurosci. 22, 317–327 (2019).
Pailhé, R. et al. Qualitative and quantitative assessment of cartilage degeneration using full-field optical coherence tomography ex vivo. Osteoarthr. Cartil. 26, 285–292 (2018).
Raghunathan, R., Singh, M., Dickinson, M. E. & Larin, K. V. Optical coherence tomography for embryonic imaging: a review. J. Biomed. Opt. 21, 1 (2016).
Lefort, C. A review of biomedical multiphoton microscopy and its laser sources. J. Phys. Appl. Phys. 50, 423001 (2017).
Disney, C. M., Lee, P. D., Hoyland, J. A., Sherratt, M. J. & Bay, B. K. A review of techniques for visualising soft tissue microstructure deformation and quantifying strain ex vivo: soft tissue microstructure deformation and quantifying strain. J. Microsc. 272, 165–179 (2018).
Schueder, F. et al. Multiplexed 3D super-resolution imaging of whole cells using spinning disk confocal microscopy and DNA-PAINT. Nat. Commun. 8, 2090 (2017).
Goth, W., Lesicko, J., Sacks, M. S. & Tunnell, J. W. Optical-based analysis of soft tissue structures. Annu. Rev. Biomed. Eng. 18, 357–385 (2016).
Edlow, B. L. et al. 7 Tesla MRI of the ex vivo human brain at 100 micron resolution. Sci. Data 6, 244 (2019).
Bouazizi, K. et al. Differentiation and quantification of fibrosis, fat and fatty fibrosis in human left atrial myocardium using ex vivo MRI. PLoS ONE 13, e0205104 (2018).
Verleden, S. E. et al. Small airway loss in the physiologically ageing lung: a cross-sectional study in unused donor lungs. Lancet Respir. Med. 9, 167–174 (2021).
Withers, P. J. et al. X-ray computed tomography. Nat. Rev. Methods Primers 1, 18 (2021).
Mastrogiacomo, M., Campi, G., Cancedda, R. & Cedola, A. Synchrotron radiation techniques boost the research in bone tissue engineering. Acta Biomater. 89, 33–46 (2019).
Longo, E. et al. 3D spatial distribution of nanoparticles in mice brain metastases by X-ray phase-contrast tomography. Front. Oncol. 11, 554668 (2021).
Hwu, Y., Margaritondo, G. & Chiang, A.-S. Q&A: why use synchrotron x-ray tomography for multi-scale connectome mapping? BMC Biol. 15, 122 (2017).
Yokawa, K. et al. Synchrotron radiation-based X-ray phase-contrast imaging of the aortic walls in acute aortic dissection. JVS Vasc. Sci. 1, 81–91 (2020).
Liu, Y., Nelson, J., Holzner, C., Andrews, J. C. & Pianetta, P. Recent advances in synchrotron-based hard x-ray phase contrast imaging. J. Phys. Appl. Phys. 46, 494001 (2013).
Gonzalez-Tendero, A. et al. Whole heart detailed and quantitative anatomy, myofibre structure and vasculature from X-ray phase-contrast synchrotron radiation-based micro computed tomography. Eur. Heart J. Cardiovasc. Imaging 18, 732–741 (2017).
Planinc, I. et al. Comprehensive assessment of myocardial remodeling in ischemic heart disease by synchrotron propagation based X-ray phase contrast imaging. Sci. Rep. 11, 14020 (2021).
Chin, A.-L. et al. A synchrotron X-ray imaging strategy to map large animal brains. Chin. J. Phys. 65, 24–32 (2020).
Endrizzi, M. X-ray phase-contrast imaging. Nucl. Instrum. Methods Phys. Res. Sect. Accel. Spectrometers Detect. Assoc. Equip. 878, 88–98 (2018).
Schulz, G. et al. High-resolution tomographic imaging of a human cerebellum: comparison of absorption and grating-based phase contrast. J. R. Soc. Interface 7, 1665–1676 (2010).
Shinohara, G. et al. Three dimensional visualization of human cardiac conduction tissue in whole heart specimens by high-resolution phase-contrast CT imaging using synchrotron radiation. World J. Pediatr. Congenit. Heart Surg. 7, 700–705 (2016).
Westöö, C. et al. Distinct types of plexiform lesions identified by synchrotron-based phase-contrast micro-CT. Am. J. Physiol. Lung Cell. Mol. Physiol. 321, L17–L28 (2021).
Borisova, E. et al. Micrometer-resolution X-ray tomographic full-volume reconstruction of an intact post-mortem juvenile rat lung. Histochem. Cell Biol. 155, 215–226 (2021).
Broche, L. et al. Individual Airway Closure Characterized In Vivo by Phase-Contrast CT Imaging in Injured Rabbit Lung. Crit. Care Med. 47, e774–e781 (2019).
Dutel, H. et al. Neurocranial development of the coelacanth and the evolution of the sarcopterygian head. Nature 569, 556–559 (2019).
Mansuit, R. et al. Development and growth of the pectoral girdle and fin skeleton in the extant coelacanth Latimeria chalumnae. J. Anat. 236, 493–509 (2020).
Walsh, C. L. et al. Imaging intact human organs with local resolution of cellular structures using hierarchical phase-contrast tomography. Nat. Methods https://doi.org/10.1038/s41592-021-01317-x (2021).
Dias, C. S. B., Neto, D. P. A., Baraldi, G. L. & Fonseca, M. C. Comparative analysis of sample preparation protocols of soft biological tissues for morphometric studies using synchrotron-based X-ray microtomography. J. Synchrotron Radiat. 26, 2013–2023 (2019).
Li, T., Schreibmann, E., Yang, Y. & Xing, L. Motion correction for improved target localization with on-board cone-beam computed tomography. Phys. Med. Biol. 51, 253–267 (2006).
Sun, T., Kim, J.-H., Fulton, R. & Nuyts, J. An iterative projection-based motion estimation and compensation scheme for head x-ray CT. Med. Phys. 43, 5705–5716 (2016).
Topal, E. et al. Multi-scale X-ray tomography and machine learning algorithms to study MoNi4 electrocatalysts anchored on MoO2 cuboids aligned on Ni foam. BMC Mater. 2, 5 (2020).
Topal, E., Löffler, M. & Zschech, E. Deep learning-based inaccuracy compensation in reconstruction of high resolution XCT data. Sci. Rep. 10, 7682 (2020).
Burger, M. et al. A variational reconstruction method for undersampled dynamic x-ray tomography based on physical motion models. Inverse Probl. 33, 124008 (2017).
Patzelt, M. et al. Ethanol fixation method for heart and lung imaging in micro-CT. Jpn. J. Radiol. 37, 500–510 (2019).
Ackermann, M. et al. The bronchial circulation in COVID-19 pneumonia. Am. J. Respir. Crit. Care Med. 205, 121–125 (2022).
Ackermann, M. et al. The fatal trajectory of pulmonary COVID-19 is driven by lobular ischemia and fibrotic remodelling. eBioMedicine 85, 104296 (2022).
Rodgers, G. et al. Impact of fixation and paraffin embedding on mouse brain morphology: a synchrotron radiation-based tomography study. In Proc. SPIE Developments in X-Ray Tomography XIII (eds Müller, B. & Wang, G.) 27 (SPIE, 2021); https://doi.org/10.1117/12.2595144
Rodgers, G. et al. Virtual histology of an entire mouse brain from formalin fixation to paraffin embedding. Part 1: data acquisition, anatomical feature segmentation, tracking global volume and density changes. J. Neurosci. Methods 364, 109354 (2021).
Gusnard, D. & Kirschner, R. H. Cell and organelle shrinkage during preparation for scanning electron microscopy: effects of fixation, dehydration and critical point drying. J. Microsc. 110, 51–57 (1977).
Metscher, B. D. MicroCT for developmental biology: a versatile tool for high-contrast 3D imaging at histological resolutions. Dev. Dyn. 238, 632–640 (2009).
Shirai, R. et al. Enhanced renal image contrast by ethanol fixation in phase-contrast X-ray computed tomography. J. Synchrotron Radiat. 21, 795–800 (2014).
Wilke, H.-J., Krischak, S. & Claes, L. E. Formalin fixation strongly influences biomechanical properties of the spine. J. Biomech. 29, 1629–1631 (1996).
Rouleau, L., Tremblay, D., Cartier, R., Mongrain, R. & Leask, R. L. Regional variations in canine descending aortic tissue mechanical properties change with formalin fixation. Cardiovasc. Pathol. 21, 390–397 (2012).
Vesper, E. O., Hammond, M. A., Allen, M. R. & Wallace, J. M. Even with rehydration, preservation in ethanol influences the mechanical properties of bone and how bone responds to experimental manipulation. Bone 97, 49–53 (2017).
Madi, K. et al. In situ characterization of nanoscale strains in loaded whole joints via synchrotron X-ray tomography. Nat. Biomed. Eng. 4, 343–354 (2020).
Gignac, P. M. et al. Diffusible iodine‐based contrast‐enhanced computed tomography (diceCT): an emerging tool for rapid, high‐resolution, 3‐D imaging of metazoan soft tissues. J. Anat. 228, 889–909 (2016).
Koç, M. M., Aslan, N., Kao, A. P. & Barber, A. H. Evaluation of X‐ray tomography contrast agents: a review of production, protocols, and biological applications. Microsc. Res. Tech. 82, 812–848 (2019).
Metscher, B. D. MicroCT for comparative morphology: simple staining methods allow high-contrast 3D imaging of diverse non-mineralized animal tissues. BMC Physiol. 9, 11 (2009).
Mittone, A. et al. Multiscale pink-beam microCT imaging at the ESRF-ID17 biomedical beamline. J. Synchrotron Radiat. 27, 1347–1357 (2020).
Xian, R. P. et al. A multiscale X-ray phase-contrast tomography dataset of a whole human left lung. Sci. Data 9, 264 (2022).
Saccomano, M. et al. Synchrotron inline phase contrast µCT enables detailed virtual histology of embedded soft-tissue samples with and without staining. J. Synchrotron Radiat. 25, 1153–1161 (2018).
Strotton, M. C. et al. Optimising complementary soft tissue synchrotron X-ray microtomography for reversibly-stained central nervous system samples. Sci. Rep. 8, 12017 (2018).
Wereszczyńska, B. Alcohol-fixed specimens for high-contrast post-mortem MRI. Forensic Imaging 25, 200449 (2021).
Roebroeck, A., Miller, K. L. & Aggarwal, M. Ex vivo diffusion MRI of the human brain: technical challenges and recent advances. NMR Biomed. 32, e3941 (2019).
Shepherd, T. M., Thelwall, P. E., Stanisz, G. J. & Blackband, S. J. Aldehyde fixative solutions alter the water relaxation and diffusion properties of nervous tissue: aldehyde fixation alters tissue mri properties. Magn. Reson. Med. 62, 26–34 (2009).
Wilke, J. et al. Appraising the methodological quality of cadaveric studies: validation of the QUACS scale. J. Anat. 226, 440–446 (2015).
Paganin, D., Mayo, S. C., Gureyev, T. E., Miller, P. R. & Wilkins, S. W. Simultaneous phase and amplitude extraction from a single defocused image of a homogeneous object. J. Microsc. 206, 33–40 (2002).
Mirone, A., Brun, E., Gouillart, E., Tafforeau, P. & Kieffer, J. The PyHST2 hybrid distributed code for high speed tomographic reconstruction with iterative reconstruction and a priori knowledge capabilities. Nucl. Instrum. Methods Phys. Res. Sect. B Beam Interact. Mater. 324, 41–48 (2014).
Gürsoy, D., De Carlo, F., Xiao, X. & Jacobsen, C. TomoPy: a framework for the analysis of synchrotron tomographic data. J. Synchrotron Radiat. 21, 1188–1193 (2014).
Lyckegaard, A., Johnson, G. & Tafforeau, P. Correction of ring artifacts in X-ray tomographic images. Int J. Tomo Stat. 18, 1–9 (2011).
Oishi, H. et al. Ex vivo lung CT findings may predict the outcome of the early phase after lung transplantation. PLoS One 15, e0233804 (2020).
Verleden, S. E. et al. Radiological analysis of unused donor lungs: a tool to improve donor acceptance for transplantation? Am. J. Transplant. 17, 1912–1921 (2017).
Ross, M. H. & Pawlina, W. Histology: A Text and Atlas: With Correlated Cell and Molecular Biology (Wolters Kluwer, 2020).
Acknowledgements
We thank S. Bayat (INSERM) for help during the test phase, P. Masson (LADAF) for dissections of donors’ bodies, H. Reichert (ESRF) and R. Tori for general support of the project and E. Boller, C. Muzelle, R. Homs, C. Jarnias, F. Cianciosi, P. Vieux, P. Cook, L. Capasso and A. Mirone for their help in setup developments and improvements. We also thank R. Engelhardt, A. Muller Brechlin, C. Petzold, N. Kroenke and M. Kuhel for help with histology and autopsies. This project has been made possible in part by grant number 2020-225394 from the Chan Zuckerberg Initiative DAF, an advised fund of Silicon Valley Community Foundation and grant number CZIF2021-006424 from the Chan Zuckerberg Initiative Foundation, The ESRF funding proposals md1252 and md1290, the Royal Academy of Engineering (CiET1819/10). P.D.L. and C.L.W. gratefully acknowledge funding from the MRC (MR/R025673/1). M.A. acknowledges grants from the National Institutes of Health (HL94567 and HL134229). This work was supported by the German Registry of COVID-19 Autopsies (DeRegCOVID, www.DeRegCOVID.ukaachen.de; supported by the Federal Ministry of Health: ZMVI1-2520COR201) and the Federal Ministry of Education and Research as part of the Network of University Medicine (DEFEAT PANDEMIcs, 01KX2021).
Author information
Authors and Affiliations
Contributions
P.T., P.D.L., D.D.J., M.A., C.L.W. and W.L.W. conceptualized the project and designed experiments. M.A., C.W., P.T., A.B., S.E.V., C.L.W. and J.B. performed and contributed to autopsies and sample preparation. P.T. designed and built instrumentation and performed HiP-CT imaging. S.M. designed sample holders. P.T. designed and implemented tomographic reconstruction methods. J.B., P.T., C.L.W. and P.D.L. wrote the paper. All authors assisted in reviewing and revising the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks Ali Erturk, Stuart Stock and Hiroki Ueda for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Walsh, C. L. et al. Nat. Methods 18, 1532–1541 (2021): https://doi.org/10.1038/s41592-021-01317-x
Ackermann, M. et al. Am. J. Respir. Crit. Care Med. 205, 121–125 (2022): https://doi.org/10.1164/rccm.202103-0594IM
Ackermann, M. et al. eBioMedicine 85, 104296 (2022): https://doi.org/10.1016/j.ebiom.2022.104296
Extended data
Extended Data Fig. 1 Challenges faced during the development of the technique and their solutions.
This protocol was developed in an iterative manner overcoming all the different challenges related to soft tissue imaging, dose deposition and local tomography.
Extended Data Fig. 2 Basic information on human organs used in this protocol paper.
The heart, kidney, and brain data are present in the Human Organ Atlas (https://human-organ-atlas.esrf.eu). The liver data are not included in the Human Organ Atlas because of the large number of artifacts present in the images due to bubble formation during scanning. However, the images are available on request from the corresponding authors. The formation of these bubbles occurred because a crash in the beamline software caused the beam to remain in the same position for several hours, exceeding the organ’s dose threshold. Other livers have been scanned without artifacts.
Extended Data Fig. 3 List of human organs and biological samples compatible with this method and the preparation time for each step.
The process described in this protocol paper should work for all major organs, but only the organs listed here have been tested. The maximum degassing times listed here are specific to our vacuum degassing setup and are subject to change depending on the pump, the volume to be pumped, the pumping section, the quantity of gas to be evacuated, and the paths that the gas can take to leave the sample. As such, they should be adapted to each user vacuum setup by looking at the bubbling intensity as explained in Step 4A(ii) of the protocol.
Extended Data Fig. 4 The two types of large leak-proof container in PET in the custom-made container holders.
The left leak-proof container is a Medline Scientific container (cat. no. 129-0592) of 3 L. The right leak-proof container is a Lock & Lock container (cat. no. INL-403) of 2.1 L. Both are inside a custom-made container holder described in Extended Data Fig. 5.
Extended Data Fig. 5 Custom-made container holder drawings and 3D rendering.
Two containers are represented inside the custom-made container holder. The bottom container contains the sample while the top container, referred to as the reference container, is used for the flat-field correction.
Extended Data Fig. 6 Beamline parameters of all the organ scans presented in this paper.
The optimal scanning settings are strongly dependent on the sample and on the beamline setup and equipment. The parameters presented here are given as examples. Quarter acquisition means one scan in half-acquisition plus one annular scan to increase the lateral field of view. Mo = molybdenum.
Supplementary information
Supplementary Video 1
Example of a sample mounted with insufficiently compacted agar, allowing rotation upon a slight movement.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Brunet, J., Walsh, C.L., Wagner, W.L. et al. Preparation of large biological samples for high-resolution, hierarchical, synchrotron phase-contrast tomography with multimodal imaging compatibility. Nat Protoc 18, 1441–1461 (2023). https://doi.org/10.1038/s41596-023-00804-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-023-00804-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.