Abstract
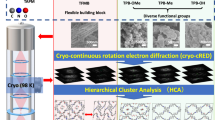
Metal–organic frameworks (MOFs) have attracted considerable interest due to their well-defined pore architecture and structural tunability on molecular dimensions. While single-crystal X-ray diffraction (SCXRD) has been widely used to elucidate the structures of MOFs at the atomic scale, the formation of large and well-ordered crystals is still a crucial bottleneck for structure determination. To alleviate this challenge, three-dimensional electron diffraction (3D ED) has been developed for structure determination of nano- (submicron-)sized crystals. Such 3D ED data are collected from each single crystal using transmission electron microscopy. In this protocol, we introduce the entire workflow for structural analysis of MOFs by 3D ED, from sample preparation, data acquisition and data processing to structure determination. We describe methods for crystal screening and handling of crystal agglomerates, which are crucial steps in sample preparation for single-crystal 3D ED data collection. We further present how to set up a transmission electron microscope for 3D ED data acquisition and, more importantly, offer suggestions for the optimization of data acquisition conditions. For data processing, including unit cell and space group determination, and intensity integration, we provide guidelines on how to use electron and X-ray crystallography software to process 3D ED data. Finally, we present structure determination from 3D ED data and discuss the important features associated with 3D ED data that need to be considered. We believe that this protocol provides critical details for implementing and utilizing 3D ED as a structure determination platform for nano- (submicron-)sized MOFs as well as other crystalline materials.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout














Similar content being viewed by others
Data availability
CCDC 1865990-1865999, 2004306, 2020844 and 2063952 contain the example crystallographic data as part of the supporting primary research publications. These data can be obtained free of charge from www.ccdc.cam.ac.uk/data_request/cif, or by emailing data_request@ccdc.cam.ac.uk, or by contacting The Cambridge Crystallographic Data Centre, 12 Union Road, Cambridge CB2 1EZ, UK; fax: +44 1223 336033. Additional reasonable requests for data supporting this publication should be addressed to the corresponding authors.
Code availability
Reasonable requests for the code and scripts used in this publication should be addressed to the corresponding authors.
References
Yaghi, O. M., Li, G. & Li, H. Selective binding and removal of guests in a microporous metal–organic framework. Nature 378, 703–706 (1995).
Kitagawa, S., Kitaura, R. & Noro, S. Functional porous coordination polymers. Angew. Chem. Int. Ed. 43, 2334–2375 (2004).
Lee, J. et al. Metal–organic framework materials as catalysts. Chem. Soc. Rev. 38, 1450–1459 (2009).
Yoon, M., Srirambalaji, R. & Kim, K. Homochiral metal–organic frameworks for asymmetric heterogeneous catalysis. Chem. Rev. 112, 1196–1231 (2012).
Li, H., Eddaoudi, M., Groy, T. L. & Yaghi, O. M. Establishing microporosity in open metal–organic frameworks: gas sorption isotherms for Zn(BDC) (BDC = 1,4-benzenedicarboxylate). J. Am. Chem. Soc. 120, 8571–8572 (1998).
Li, Q. et al. Docking in metal–organic frameworks. Science 325, 855–859 (2009).
Ma, S. & Zhou, H.-C. Gas storage in porous metal–organic frameworks for clean energy applications. Chem. Commun. 46, 44–53 (2010).
Trickett, C. A. et al. The chemistry of metal–organic frameworks for CO2 capture, regeneration and conversion. Nat. Rev. Mater. 2, 1–16 (2017).
Li, J.-R., Sculley, J. & Zhou, H.-C. Metal–organic frameworks for separations. Chem. Rev. 112, 869–932 (2012).
Duan, J., Jin, W. & Kitagawa, S. Water-resistant porous coordination polymers for gas separation. Coord. Chem. Rev. 332, 48–74 (2017).
Bobbitt, N. S. et al. Metal–organic frameworks for the removal of toxic industrial chemicals and chemical warfare agents. Chem. Soc. Rev. 46, 3357–3385 (2017).
Lin, R.-B., Xiang, S., Xing, H., Zhou, W. & Chen, B. Exploration of porous metal–organic frameworks for gas separation and purification. Coord. Chem. Rev. 378, 87–103 (2019).
Zhang, T. & Lin, W. Metal–organic frameworks for artificial photosynthesis and photocatalysis. Chem. Soc. Rev. 43, 5982–5993 (2014).
Xia, W., Mahmood, A., Zou, R. & Xu, Q. Metal–organic frameworks and their derived nanostructures for electrochemical energy storage and conversion. Energy Environ. Sci. 8, 1837–1866 (2015).
Sheberla, D. et al. Conductive MOF electrodes for stable supercapacitors with high areal capacitance. Nat. Mater. 16, 220–224 (2017).
Feng, D. et al. Robust and conductive two-dimensional metal–organic frameworks with exceptionally high volumetric and areal capacitance. Nat. Energy 3, 30–36 (2018).
Park, J. et al. High thermopower in a Zn-based 3D semiconductive metal–organic framework. J. Am. Chem. Soc. 142, 20531–20535 (2020).
Della Rocca, J., Liu, D. & Lin, W. Nanoscale metal–organic frameworks for biomedical imaging and drug delivery. Acc. Chem. Res. 44, 957–968 (2011).
Horcajada, P. et al. Metal–organic frameworks in biomedicine. Chem. Rev. 112, 1232–1268 (2012).
Doonan, C., Riccò, R., Liang, K., Bradshaw, D. & Falcaro, P. Metal–organic frameworks at the biointerface: synthetic strategies and applications. Acc. Chem. Res. 50, 1423–1432 (2017).
Cichocka, M. O. et al. A porphyrinic zirconium metal–organic framework for oxygen reduction reaction: tailoring the spacing between active-sites through chain-based inorganic building units. J. Am. Chem. Soc. 142, 15386–15395 (2020).
Ge, M. et al. High-throughput electron diffraction reveals a hidden novel metal–organic framework for electrocatalysis. Angew. Chem. Int. Ed. 60, 11391–11397 (2021).
Dorset, D. L. & Hauptman, H. A. Direct phase determination for quasi-kinematical electron diffraction intensity data from organic microcrystals. Ultramicroscopy 1, 195–201 (1976).
Dorset, D. L. Electron crystallography—accomplishments and challenges. Acta Crystallogr. A 54, 750–757 (1998).
Gemmi, M. et al. 3D electron diffraction: the nanocrystallography revolution. ACS Cent. Sci. 5, 1315–1329 (2019).
Gruene, T., Holstein, J. J., Clever, G. H. & Keppler, B. Establishing electron diffraction in chemical crystallography. Nat. Rev. Chem. 5, 660–668 (2021).
Gruene, T. & Mugnaioli, E. 3D electron diffraction for chemical analysis: instrumentation developments and innovative applications. Chem. Rev. 121, 11823–11834 (2021).
Huang, Z., Willhammar, T. & Zou, X. Three-dimensional electron diffraction for porous crystalline materials: structural determination and beyond. Chem. Sci. 12, 1206–1219 (2021).
Huang, Z., Grape, E. S., Li, J., Inge, A. K. & Zou, X. 3D electron diffraction as an important technique for structure elucidation of metal-organic frameworks and covalent organic frameworks. Coord. Chem. Rev. 427, 213583 (2021).
Kolb, U., Gorelik, T., Kübel, C., Otten, M. T. & Hubert, D. Towards automated diffraction tomography: Part I—data acquisition. Ultramicroscopy 107, 507–513 (2007).
Kolb, U., Gorelik, T. & Otten, M. T. Towards automated diffraction tomography. Part II—cell parameter determination. Ultramicroscopy 108, 763–772 (2008).
Mugnaioli, E., Gorelik, T. & Kolb, U. “Ab initio” structure solution from electron diffraction data obtained by a combination of automated diffraction tomography and precession technique. Ultramicroscopy 109, 758–765 (2009).
Zhang, D., Oleynikov, P., Hovmöller, S. & Zou, X. Collecting 3D electron diffraction data by the rotation method. Z. Krist. 225, 94–102 (2010).
Wan, W., Sun, J., Su, J., Hovmöller, S. & Zou, X. Three-dimensional rotation electron diffraction: software RED for automated data collection and data processing. J. Appl. Crystallogr. 46, 1863–1873 (2013).
Jiang, J. et al. Synthesis and structure determination of the hierarchical meso-microporous zeolite ITQ-43. Science 333, 1131–1134 (2011).
Willhammar, T. et al. EMM-23: a stable high-silica multidimensional zeolite with extra-large trilobe-shaped channels. J. Am. Chem. Soc. 136, 13570–13573 (2014).
Guo, P. et al. A zeolite family with expanding structural complexity and embedded isoreticular structures. Nature 524, 74–78 (2015).
Birkel, C. S. et al. Solution synthesis of a new thermoelectric Zn1+xSb nanophase and its structure determination using automated electron diffraction tomography. J. Am. Chem. Soc. 132, 9881–9889 (2010).
Feyand, M. et al. Automated diffraction tomography for the structure elucidation of twinned, sub-micrometer crystals of a highly porous, catalytically active bismuth metal–organic framework. Angew. Chem. Int. Ed. 51, 10373–10376 (2012).
Cichocka, M. O., Ångström, J., Wang, B., Zou, X. & Smeets, S. High-throughput continuous rotation electron diffraction data acquisition via software automation. J. Appl. Crystallogr. 51, 1652–1661 (2018).
Nannenga, B. L., Shi, D., Leslie, A. G. W. & Gonen, T. High-resolution structure determination by continuous-rotation data collection in MicroED. Nat. Methods 11, 927–930 (2014).
Plana-Ruiz, S. et al. Fast-ADT: a fast and automated electron diffraction tomography setup for structure determination and refinement. Ultramicroscopy 211, 112951 (2020).
Gemmi, M., La Placa, M. G. I., Galanis, A. S., Rauch, E. F. & Nicolopoulos, S. Fast electron diffraction tomography. J. Appl. Crystallogr. 48, 718–727 (2015).
Nederlof, I., van Genderen, E., Li, Y.-W. & Abrahams, J. P. A Medipix quantum area detector allows rotation electron diffraction data collection from submicrometre three-dimensional protein crystals. Acta Crystallogr. D. 69, 1223–1230 (2013).
van Genderen, E. et al. Ab initio structure determination of nanocrystals of organic pharmaceutical compounds by electron diffraction at room temperature using a Timepix quantum area direct electron detector. Acta Crystallogr. Sect. Found. Adv. 72, 236–242 (2016).
Yuan, S. et al. [Ti8Zr2O12(COO)16] cluster: an ideal inorganic building unit for photoactive metal–organic frameworks. ACS Cent. Sci. 4, 105–111 (2018).
Hynek, J., Brázda, P., Rohlíček, J., Londesborough, M. G. S. & Demel, J. Phosphinic acid based linkers: building blocks in metal–organic framework chemistry. Angew. Chem. Int. Ed. 57, 5016–5019 (2018).
Portolés-Gil, N. et al. Crystalline curcumin bioMOF obtained by precipitation in supercritical CO2 and structural determination by electron diffraction tomography. ACS Sustain. Chem. Eng. 6, 12309–12319 (2018).
Lenzen, D. et al. A metal–organic framework for efficient water-based ultra-low-temperature-driven cooling. Nat. Commun. 10, 3025 (2019).
Leubner, S. et al. Expanding the variety of zirconium-based inorganic building units for metal–organic frameworks. Angew. Chem. Int. Ed. 58, 10995–11000 (2019).
Dou, J.-H. et al. Atomically precise single-crystal structures of electrically conducting 2D metal–organic frameworks. Nat. Mater. 20, 222–228 (2021).
Shi, D. et al. The collection of microED data for macromolecular crystallography. Nat. Protoc. 11, 895–904 (2016).
Wang, B. et al. A porous cobalt tetraphosphonate metal–organic framework: accurate structure and guest molecule location determined by continuous-rotation electron diffraction. Chem. Eur. J. 24, 17429–17433 (2018).
Samperisi, L. et al. Probing molecular motions in metal–organic frameworks by three-dimensional electron diffraction. J. Am. Chem. Soc. 143, 17947–17952 (2021).
Gao, C. et al. Isostructural three-dimensional covalent organic frameworks. Angew. Chem. Int. Ed. 58, 9770–9775 (2019).
Sun, T., Wei, L., Chen, Y., Ma, Y. & Zhang, Y.-B. Atomic-level characterization of dynamics of a 3D covalent organic framework by cryo-electron diffraction tomography. J. Am. Chem. Soc. 141, 10962–10966 (2019).
Kapaca, E. et al. Synthesis and structure of a 22 × 12 × 12 extra-large pore zeolite ITQ-56 determined by 3D electron diffraction. J. Am. Chem. Soc. 143, 8713–8719 (2021).
Fröjdh, E. et al. Discrimination of aluminum from silicon by electron crystallography with the JUNGFRAU detector. Crystals 10, 1148 (2020).
Seo, S. et al. Two aluminophosphate molecular sieves built from pairs of enantiomeric structural building units. Angew. Chem. Int. Ed. 57, 3727–3732 (2018).
Huang, Z. et al. 3D–3D topotactic transformation in aluminophosphate molecular sieves and its implication in new zeolite structure generation. Nat. Commun. 11, 3762 (2020).
Gruene, T. et al. Rapid structure determination of microcrystalline molecular compounds using electron diffraction. Angew. Chem. Int. Ed. 57, 16313–16317 (2018).
Jones, C. G. et al. The cryoEM method microED as a powerful tool for small molecule structure determination. ACS Cent. Sci. 4, 1587–1592 (2018).
Broadhurst, E. T. et al. Polymorph evolution during crystal growth studied by 3D electron diffraction. IUCrJ 7, 5–9 (2020).
Broadhurst, E. T., Xu, H., Parsons, S. & Nudelman, F. Revealing the early stages of carbamazepine crystallization by cryoTEM and 3D electron diffraction. IUCrJ 8, (2021).
Palatinus, L. et al. Hydrogen positions in single nanocrystals revealed by electron diffraction. Science 355, 166–169 (2017).
Brázda, P., Palatinus, L. & Babor, M. Electron diffraction determines molecular absolute configuration in a pharmaceutical nanocrystal. Science 364, 667–669 (2019).
Yang, L. et al. Ligand-directed conformational control over porphyrinic zirconium metal–organic frameworks for size-selective catalysis. J. Am. Chem. Soc. 143, 12129–12137 (2021).
Smeets, S., Wang, B. & Hogenbirk, E. Instamatic (2021); https://doi.org/10.5281/zenodo.1090388
edtools — edtools 1.0.1 documentation; https://edtools.readthedocs.io/en/latest/
Installation - XDSwiki; https://strucbio.biologie.uni-konstanz.de/xdswiki/index.php/Installation
Sheldrick, G. M. SHELXT – Integrated space-group and crystal-structure determination. Acta Crystallogr. Sect. Found. Adv. 71, 3–8 (2015).
Sheldrick, G. M. A short history of SHELX. Acta Crystallogr. A 64, 112–122 (2008).
Oracle VM VirtualBox; https://www.virtualbox.org/
Install WSL; https://docs.microsoft.com/en-us/windows/wsl/install
Palatinus, L. et al. Specifics of the data processing of precession electron diffraction tomography data and their implementation in the program PETS2.0. Acta Crystallogr. B 75, 512–522 (2019).
Winter, G. et al. DIALS: implementation and evaluation of a new integration package. Acta Crystallogr. Sect. Struct. Biol. 74, 85–97 (2018).
CrysAlis Pro | Rigaku global website; https://www.rigaku.com/products/crystallography/crysalis
Burla, M. C. et al. Crystal structure determination and refinement via SIR2014. J. Appl. Crystallogr. 48, 306–309 (2015).
Petříček, V., Dušek, M. & Palatinus, L. Crystallographic computing system JANA2006: general features. Z. Krist. Cryst. Mater. 229, 345–352 (2014).
Palatinus, L., Petříček, V. & Corrêa, C. A. Structure refinement using precession electron diffraction tomography and dynamical diffraction: theory and implementation. Acta Crystallogr. Sect. Found. Adv. 71, 235–244 (2015).
Roslova, M. et al. InsteaDMatic: towards cross-platform automated continuous rotation electron diffraction. J. Appl. Crystallogr. 53, 1217–1224 (2020).
Mugnaioli, E. & Gemmi, M. Single-crystal analysis of nanodomains by electron diffraction tomography: mineralogy at the order-disorder borderline. Z. Krist. Cryst. Mater. 233, 163–178 (2018).
Nannenga, B. L., Shi, D., Leslie, A. G. W. & Gonen, T. High-resolution structure determination by continuous-rotation data collection in MicroED. Nat. Methods 11, 927–930 (2014).
Smeets, S., Wang, B. & Hogenbirk, E. instamatic-dev/instamatic: 1.7.0 (Zenodo, 2021); https://doi.org/10.5281/zenodo.5175957
Wang, B., Zou, X. & Smeets, S. Automated serial rotation electron diffraction combined with cluster analysis: an efficient multi-crystal workflow for structure determination. IUCrJ 6, 854–867 (2019).
de la Cruz, M. J., Martynowycz, M. W., Hattne, J. & Gonen, T. MicroED data collection with SerialEM. Ultramicroscopy 201, 77–80 (2019).
Yonekura, K., Ishikawa, T. & Maki-Yonekura, S. A new cryo-EM system for electron 3D crystallography by eEFD. J. Struct. Biol. 206, 243–253 (2019).
Carragher, B. et al. Leginon: An automated system for acquisition of images from vitreous ice specimens. J. Struct. Biol. 132, 33–45 (2000).
Electron Microscopy | Software Updates - SE; http://www.thermofisher.com/uk/en/home/electron-microscopy/products/software-em-3d-vis/software-updates.html
cctbx - Determination of lattice symmetry; http://cci.lbl.gov/cctbx/lattice_symmetry.html
Grosse-Kunstleve, R. W., Sauter, N. K., Moriarty, N. W. & Adams, P. D. The Computational Crystallography Toolbox: crystallographic algorithms in a reusable software framework. J. Appl. Crystallogr. 35, 126–136 (2002).
Xu, H. XDS input template for 3D ED/MicroED data processing. (2021); https://doi.org/10.5281/zenodo.5574261
Doyle, P. A. & Turner, P. S. Relativistic Hartree–Fock X-ray and electron scattering factors. Acta Crystallogr. A 24, 390–397 (1968).
Peng, L.-M. Electron scattering factors of ions and their parameterization. Acta Crystallogr. A 54, 481–485 (1998).
Dolomanov, O. V., Bourhis, L. J., Gildea, R. J., Howard, J. A. K. & Puschmann, H. OLEX2: a complete structure solution, refinement and analysis program. J. Appl. Crystallogr. 42, 339–341 (2009).
FAES: Factors of Atomic Electron Scattering; https://srv.mbi.ucla.edu/faes/
Hübschle, C. B., Sheldrick, G. M. & Dittrich, B. ShelXle: a Qt graphical user interface for SHELXL. J. Appl. Crystallogr. 44, 1281–1284 (2011).
International Tables for Crystallography. C: Mathematical, Physical and Chemical Tables (ed. Prince, E.) (Kluwer Academic, 2004).
CheckCIF; https://checkcif.iucr.org/
Spek, A. L. Single-crystal structure validation with the program PLATON. J. Appl. Crystallogr. 36, 7–13 (2003).
Spek, A. L. checkCIF validation ALERTS: what they mean and how to respond. Acta Crystallogr. E 76, 1–11 (2020).
Case: Electron Diffraction Data in the CSD - The Cambridge Crystallographic Data Centre (CCDC); https://www.ccdc.cam.ac.uk/support-and-resources/support/case/?caseid=ceb813f7-aca7-e911-8cce-005056975d8a
Wang, Y., Yang, T., Xu, H., Zou, X. & Wan, W. On the quality of the continuous rotation electron diffraction data for accurate atomic structure determination of inorganic compounds. J. Appl. Crystallogr. 51, 1094–1101 (2018).
Huang, Z. et al. Can 3D electron diffraction provide accurate atomic structures of metal–organic frameworks? Faraday Discuss. 225, 118–132 (2021).
Latitude D software | Gatan; https://www.gatan.com/products/tem-imaging-spectroscopy/latitude-d-software
Ge, M. et al. On the completeness of three-dimensional electron diffraction data for structural analysis of metal–organic frameworks. Faraday Discuss. 231, 66–80 (2021).
Acknowledgements
This work was supported by the Swedish research council Formas (2020-00831 to Z.H.), the Swedish Research Council (VR 2016-04625 to Z.H., VR 2017-04321 and 2019-00815 to X.Z., VR 2017-05333 to H.X. and VR 2019-05465 to T.W.), the Knut and Alice Wallenberg Foundation (2012.0112 and 2018.0237 to X.Z.) and the MicroED@SciLifeLab project (H.X.)
Author information
Authors and Affiliations
Contributions
T.Y., T.W., H.X., X.Z. and Z.H. wrote the manuscript. Z.H. initiated this manuscript. All authors read, discussed and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks Jessica Bruhn, Russell E. Morris and Enrico Mugnaioli for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Ge, M. et al. Angew. Chem. Int. Ed. 60, 11391–11397 (2021): https://doi.org/10.1002/anie.202016882
Samperisi, L. et al. J. Am. Chem. Soc. 143, 17947–17952 (2021): https://doi.org/10.1021/jacs.1c08354
Cichocka, M. O. et al. J. Am. Chem. Soc. 142, 15386–15395 (2020): https://doi.org/10.1021/jacs.0c06329
Yuan, S. et al. ACS Cent. Sci. 4, 105–111 (2018): https://doi.org/10.1021/acscentsci.7b00497
Wang, B. et al. Chem. Eur. J. 24, 17429–17433 (2018): https://doi.org/10.1002/chem.201804133
Key data used in this protocol
Wang, B. et al. Chem. Eur. J. 24, 17429–17433 (2018): https://doi.org/10.1002/chem.201804133
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yang, T., Willhammar, T., Xu, H. et al. Single-crystal structure determination of nanosized metal–organic frameworks by three-dimensional electron diffraction. Nat Protoc 17, 2389–2413 (2022). https://doi.org/10.1038/s41596-022-00720-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-022-00720-8
This article is cited by
-
Synthesis of microcrystalline indium (III)-MOF and adsorptive and selective removal of dyes
Research on Chemical Intermediates (2024)
-
Structure determination of a low-crystallinity covalent organic framework by three-dimensional electron diffraction
Communications Chemistry (2023)
-
Materials for a changing planet
Nature Materials (2023)
-
Crystal formation, structural, optical, and dielectric measurements of l-histidine hydrochloride hydrate (LHHCLH) crystals for optoelectronic applications
Journal of Materials Science: Materials in Electronics (2023)
-
Delocalization state-induced selective bond breaking for efficient methanol electrosynthesis from CO2
Nature Catalysis (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.