Abstract
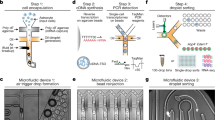
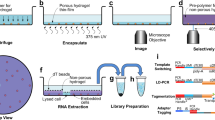
Brain–immune cross-talk and neuroinflammation critically shape brain physiology in health and disease. A detailed understanding of the brain immune landscape is essential for developing new treatments for neurological disorders. Single-cell technologies offer an unbiased assessment of the heterogeneity, dynamics and functions of immune cells. Here we provide a protocol that outlines all the steps involved in performing single-cell multi-omic analysis of the brain immune compartment. This includes a step-by-step description on how to microdissect the border regions of the mouse brain, together with dissociation protocols tailored to each of these tissues. These combine a high yield with minimal dissociation-induced gene expression changes. Next, we outline the steps involved for high-dimensional flow cytometry and droplet-based single-cell RNA sequencing via the 10x Genomics platform, which can be combined with cellular indexing of transcriptomes and epitopes by sequencing (CITE-seq) and offers a higher throughput than plate-based methods. Importantly, we detail how to implement CITE-seq with large antibody panels to obtain unbiased protein-expression screening coupled to transcriptome analysis. Finally, we describe the main steps involved in the analysis and interpretation of the data. This optimized workflow allows for a detailed assessment of immune cell heterogeneity and activation in the whole brain or specific border regions, at RNA and protein level. The wet lab workflow can be completed by properly trained researchers (with basic proficiency in cell and molecular biology) and takes between 6 and 11 h, depending on the chosen procedures. The computational analysis requires a background in bioinformatics and programming in R.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The scRNA-seq data shown in Fig. 4a,b,e are deposited at GEO under GSE128855; Fig. 4f under GSE157480. The CITE-seq data shown in Fig. 5 are deposited at GEO under GSE163120. The gene–cell count matrices of all these datasets can also be downloaded at www.brainimmuneatlas.org. The CITE-seq data shown in Fig. 6 are deposited at GEO under GSE191075.
Code availability
The R codes that were used for scRNA-seq and CITE-seq analyses can be found at Github: https://github.com/Movahedilab/Mouse_brain_borders_NatureProtocols.
References
Salvador, A. F., de Lima, K. A. & Kipnis, J. Neuromodulation by the immune system: a focus on cytokines. Nat. Rev. Immunol. 21, 526–541 (2021).
Hammond, T. R., Robinton, D. & Stevens, B. Microglia and the brain: complementary partners in development and disease. Annu. Rev. Cell Dev. Biol. 34, 523–544 (2018).
Castellani, G. & Schwartz, M. Immunological features of non-neuronal brain cells: implications for Alzheimer’s disease immunotherapy. Trends Immunol. 41, 794–804 (2020).
Sampson, J. H., Gunn, M. D., Fecci, P. E. & Ashley, D. M. Brain immunology and immunotherapy in brain tumours. Nat. Rev. Cancer 20, 12–25 (2020).
Shechter, R., London, A. & Schwartz, M. Orchestrated leukocyte recruitment to immune-privileged sites: absolute barriers versus educational gates. Nat. Rev. Immunol. 13, 206–218 (2013).
Korin, B. et al. High-dimensional, single-cell characterization of the brain’s immune compartment. Nat. Neurosci. 20, 1300–1309 (2017).
Mrdjen, D. et al. High-dimensional single-cell mapping of central nervous system immune cells reveals distinct myeloid subsets in health, aging, and disease. Immunity 48, 599 (2018).
Van Hove, H. et al. A single-cell atlas of mouse brain macrophages reveals unique transcriptional identities shaped by ontogeny and tissue environment. Nat. Neurosci. 22, 1021–1035 (2019).
Brioschi, S. et al. Heterogeneity of meningeal B cells reveals a lymphopoietic niche at the CNS borders. Science 373, eabf9277 (2021).
Cugurra, A. et al. Skull and vertebral bone marrow are myeloid cell reservoirs for the meninges and CNS parenchyma. Science 373, eabf7844 (2021).
Ajami, B. et al. Single-cell mass cytometry reveals distinct populations of brain myeloid cells in mouse neuroinflammation and neurodegeneration models. Nat. Neurosci. 21, 541–551 (2018).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Zheng, G. X. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Peterson, V. M. et al. Multiplexed quantification of proteins and transcripts in single cells. Nat. Biotechnol. 35, 936–939 (2017).
Stoeckius, M. et al. Simultaneous epitope and transcriptome measurement in single cells. Nat. Methods 14, 865–868 (2017).
Shemer, A. et al. Interleukin-10 prevents pathological microglia hyperactivation following peripheral endotoxin challenge. Immunity 53, 1033–1049 e1037 (2020).
Pombo Antunes, A. R. et al. Single-cell profiling of myeloid cells in glioblastoma across species and disease stage reveals macrophage competition and specialization. Nat. Neurosci. 24, 595–610 (2021).
Kierdorf, K., Masuda, T., Jordao, M. J. C. & Prinz, M. Macrophages at CNS interfaces: ontogeny and function in health and disease. Nat. Rev. Neurosci. 20, 547–562 (2019).
Korin, B., Dubovik, T. & Rolls, A. Mass cytometry analysis of immune cells in the brain. Nat. Protoc. 13, 377–391 (2018).
Hammond, T. R. et al. Single-cell RNA sequencing of microglia throughout the mouse lifespan and in the injured brain reveals complex cell-state changes. Immunity 50, 253–271 e256 (2019).
Baruch, K. et al. Breaking immune tolerance by targeting Foxp3(+) regulatory T cells mitigates Alzheimer’s disease pathology. Nat. Commun. 6, 7967 (2015).
Movahedi, K. et al. Different tumor microenvironments contain functionally distinct subsets of macrophages derived from Ly6C(high) monocytes. Cancer Res. 70, 5728–5739 (2010).
Peptan, I. A., Hong, L. & Evans, C. A. Multiple differentiation potentials of neonatal dura mater-derived cells. Neurosurgery. 60, 346–352 (2007). discussion 352.
Ryg-Cornejo, V., Ioannidis, L. J. & Hansen, D. S. Isolation and analysis of brain-sequestered leukocytes from Plasmodium berghei ANKA-infected mice. J. Vis. Exp. 71, e50112 (2013).
Van Damme, H. et al. Therapeutic depletion of CCR8(+) tumor-infiltrating regulatory T cells elicits antitumor immunity and synergizes with anti-PD-1 therapy. J. Immunother. Cancer 9, e001749 (2021).
Wu, Y. E., Pan, L., Zuo, Y., Li, X. & Hong, W. Detecting activated cell populations using single-cell RNA-seq. Neuron 96, 313–329 e316 (2017).
Rustenhoven, J. et al. Functional characterization of the dural sinuses as a neuroimmune interface. Cell 184, 1000–1016 e1027 (2021).
Dani, N. et al. A cellular and spatial map of the choroid plexus across brain ventricles and ages. Cell 184, 3056–3074 e3021 (2021).
Golomb, S. M. et al. Multi-modal single-cell analysis reveals brain immune landscape plasticity during aging and gut microbiota dysbiosis. Cell Rep. 33, 108438 (2020).
Masuda, T. et al. Spatial and temporal heterogeneity of mouse and human microglia at single-cell resolution. Nature 566, 388–392 (2019).
Zelco, A. et al. Single-cell atlas reveals meningeal leukocyte heterogeneity in the developing mouse brain. Genes Dev. 35, 1190–1207 (2021).
Li, Q. et al. Developmental heterogeneity of microglia and brain myeloid cells revealed by deep single-cell RNA sequencing. Neuron 101, 207–223 e210 (2019).
Jordao, M. J. C. et al. Single-cell profiling identifies myeloid cell subsets with distinct fates during neuroinflammation. Science 363, eaat7554 (2019).
Schafflick, D. et al. Single-cell profiling of CNS border compartment leukocytes reveals that B cells and their progenitors reside in non-diseased meninges. Nat. Neurosci. 24, 1225–1234 (2021).
Wang, Y. et al. Early developing B cells undergo negative selection by central nervous system-specific antigens in the meninges. Immunity 54, 2784–2794 e2786 (2021).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Brummelman, J. et al. Development, application and computational analysis of high-dimensional fluorescent antibody panels for single-cell flow cytometry. Nat. Protoc. 14, 1946–1969 (2019).
Park, L. M., Lannigan, J. & Jaimes, M. C. OMIP-069: forty-color full spectrum flow cytometry panel for deep immunophenotyping of major cell subsets in human peripheral blood. Cytometry A 97, 1044–1051 (2020).
Takahashi, C. et al. Mass cytometry panel optimization through the designed distribution of signal interference. Cytometry A 91, 39–47 (2017).
Giesen, C. et al. Highly multiplexed imaging of tumor tissues with subcellular resolution by mass cytometry. Nat. Methods 11, 417–422 (2014).
Goltsev, Y. et al. Deep profiling of mouse splenic architecture with CODEX multiplexed imaging. Cell 174, 968–981 e915 (2018).
Waylen, L. N., Nim, H. T., Martelotto, L. G. & Ramialison, M. From whole-mount to single-cell spatial assessment of gene expression in 3D. Commun. Biol. 3, 602 (2020).
Grieshaber-Bouyer, R. et al. The neutrotime transcriptional signature defines a single continuum of neutrophils across biological compartments. Nat. Commun. 12, 2856 (2021).
Monaco, G. et al. RNA-Seq signatures normalized by mRNA abundance allow absolute deconvolution of human immune cell types. Cell Rep. 26, 1627–1640 e1627 (2019).
Giordani, L. et al. High-dimensional single-cell cartography reveals novel skeletal muscle-resident cell populations. Mol. Cell 74, 609–621 e606 (2019).
Dahlin, J. S. et al. A single-cell hematopoietic landscape resolves 8 lineage trajectories and defects in Kit mutant mice. Blood 131, e1–e11 (2018).
Stoeckius, M. et al. Cell Hashing with barcoded antibodies enables multiplexing and doublet detection for single cell genomics. Genome Biol. 19, 224 (2018).
McGinnis, C. S. et al. MULTI-seq: sample multiplexing for single-cell RNA sequencing using lipid-tagged indices. Nat. Methods 16, 619–626 (2019).
Cossarizza, A. et al. Guidelines for the use of flow cytometry and cell sorting in immunological studies. Eur. J. Immunol. 47, 1584–1797 (2017).
Millard, S. M. et al. Fragmentation of tissue-resident macrophages during isolation confounds analysis of single-cell preparations from mouse hematopoietic tissues. Cell Rep. 37, 110058 (2021).
Quintelier, K. et al. Analyzing high-dimensional cytometry data using FlowSOM. Nat. Protoc. 16, 3775–3801 (2021).
Louis, K. S. & Siegel, A. C. Cell viability analysis using trypan blue: manual and automated methods. Methods Mol. Biol. 740, 7–12 (2011).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Lun, A. T., Bach, K. & Marioni, J. C. Pooling across cells to normalize single-cell RNA sequencing data with many zero counts. Genome Biol. 17, 75 (2016).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 e1821 (2019).
Klein, A. M. et al. Droplet barcoding for single-cell transcriptomics applied to embryonic stem cells. Cell 161, 1187–1201 (2015).
Zappia, L. & Oshlack, A. Clustering trees: a visualization for evaluating clusterings at multiple resolutions. Gigascience 7, giy083 (2018).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Polanski, K. et al. BBKNN: fast batch alignment of single cell transcriptomes. Bioinformatics 36, 964–965 (2020).
Hie, B., Bryson, B. & Berger, B. Efficient integration of heterogeneous single-cell transcriptomes using Scanorama. Nat. Biotechnol. 37, 685–691 (2019).
Germain, P., Lun, A., Macnair, W. & Robinson, M. Doublet identification in single-cell sequencing data using scDblFinder [version 2; peer review: 2 approved]. F1000Research 10, 979 (2022).
Xi, N. M. & Li, J. J. Protocol for executing and benchmarking eight computational doublet-detection methods in single-cell RNA sequencing data analysis. STAR Protoc. 2, 100699 (2021).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 8, 329–337 e324 (2019).
Young, M. D. & Behjati, S. SoupX removes ambient RNA contamination from droplet-based single-cell RNA sequencing data. Gigascience 9, giaa151 (2020).
Yang, S. et al. Decontamination of ambient RNA in single-cell RNA-seq with DecontX. Genome Biol. 21, 57 (2020).
Tung, P. Y. et al. Batch effects and the effective design of single-cell gene expression studies. Sci. Rep. 7, 39921 (2017).
Welch, J. D. et al. Single-cell multi-omic integration compares and contrasts features of brain cell identity. Cell 177, 1873–1887 e1817 (2019).
Tran, H. T. N. et al. A benchmark of batch-effect correction methods for single-cell RNA sequencing data. Genome Biol. 21, 12 (2020).
Saelens, W., Cannoodt, R., Todorov, H. & Saeys, Y. A comparison of single-cell trajectory inference methods. Nat. Biotechnol. 37, 547–554 (2019).
Cannoodt, R. et al.. SCORPIUS improves trajectory inference and identifies novel modules in dendritic cell development. Preprint at bioRxiv https://www.biorxiv.org/content/10.1101/079509v2 (2016).
Haghverdi, L., Buttner, M., Wolf, F. A., Buettner, F. & Theis, F. J. Diffusion pseudotime robustly reconstructs lineage branching. Nat. Methods 13, 845–848 (2016).
Setty, M. et al. Wishbone identifies bifurcating developmental trajectories from single-cell data. Nat. Biotechnol. 34, 637–645 (2016).
Street, K. et al. Slingshot: cell lineage and pseudotime inference for single-cell transcriptomics. BMC Genomics 19, 477 (2018).
Wolf, F. A. et al. PAGA: graph abstraction reconciles clustering with trajectory inference through a topology preserving map of single cells. Genome Biol. 20, 59 (2019).
Jin, S. et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. CellPhoneDB: inferring cell–cell communication from combined expression of multi-subunit ligand–receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Browaeys, R., Saelens, W. & Saeys, Y. NicheNet: modeling intercellular communication by linking ligands to target genes. Nat. Methods 17, 159–162 (2020).
Cha, J. & Lee, I. Single-cell network biology for resolving cellular heterogeneity in human diseases. Exp. Mol. Med. 52, 1798–1808 (2020).
Pratapa, A., Jalihal, A. P., Law, J. N., Bharadwaj, A. & Murali, T. M. Benchmarking algorithms for gene regulatory network inference from single-cell transcriptomic data. Nat. Methods 17, 147–154 (2020).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Habib, N. et al. Massively parallel single-nucleus RNA-seq with DroNc-seq. Nat. Methods 14, 955–958 (2017).
Acknowledgements
This work was supported by Innoviris (Attract grant BB2B 2015−2), Fonds Wetenschappelijk Onderzoek (grant no. 1506316 N) to K.M., a VLAIO grant (no. ImmCyte HBC.2016.0889) to K.M. and JAVG. K.M. is a Collen-Franqui research professor. I.S. is supported by an FWO postdoctoral fellowship. H.V.H. is supported by an FWO predoctoral fellowship. We thank Y. Elkrim, G. van Isterdael, the VIB Flow Core and the VIB Nucleomics Core for technical assistance, VIB Tech Watch and the single-cell accelerator program for support regarding single-cell technologies.
Author information
Authors and Affiliations
Contributions
I.S., H.V.H., K.D.V. and K.M. developed and optimized the protocol with input and help from C.L.S., J.A.V.G., Y.S., M.G., N.V. I.S., H.V.H., K.D.V., J.B., M.V.P., A.R.P.A. and N.V. performed wet-lab work and acquired scRNA-seq and/or CITE-seq data. I.S., D.K., L.M. and Y.S. analyzed the data and/or generated analysis pipelines. I.S., H.V.H., D.K. and K.M. wrote the manuscript. K.M. conceptualized and directed the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks Naomi Habib and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Van Hove, H. et al. Nat. Neurosci. 22, 1021–1035 (2019): https://doi.org/10.1038/s41593-019-0393-4
Shemer, A. et al. Immunity 54, 1033–1049.E7 (2020): https://doi.org/10.1016/j.immuni.2020.09.018
Pombo Antunes, A. R. et al. Nat. Neurosci. 24, 595–610 (2021): https://doi.org/10.1038/s41593-020-00789-y
Supplementary information
Supplementary Information
Supplementary Fig. 1.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Scheyltjens, I., Van Hove, H., De Vlaminck, K. et al. Single-cell RNA and protein profiling of immune cells from the mouse brain and its border tissues. Nat Protoc 17, 2354–2388 (2022). https://doi.org/10.1038/s41596-022-00716-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-022-00716-4
This article is cited by
-
Experimental hepatic encephalopathy causes early but sustained glial transcriptional changes
Journal of Neuroinflammation (2023)
-
Proteomic interrogation of the meninges reveals the molecular identities of structural components and regional distinctions along the CNS axis
Fluids and Barriers of the CNS (2023)
-
Meningeal immunity and neurological diseases: new approaches, new insights
Journal of Neuroinflammation (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.