Abstract
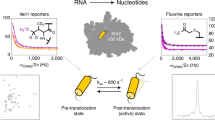
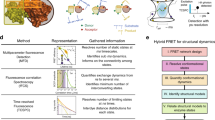
It is important to understand the dynamics and higher energy structures of RNA, called excited states, to achieve better understanding of RNA function. R1ρ relaxation dispersion NMR spectroscopy (RD) determines chemical shift differences between the most stable, ground state and the short-lived, low-populated excited states. We describe a procedure for deducing the excited state structure from these chemical shift differences using the mutate-and-chemical-shift-fingerprint (MCSF) method, which requires ~2–6 weeks and moderate understanding of NMR and RNA structure. We recently applied the MCSF methodology to elucidate the excited state of microRNA 34a targeting the SIRT1 mRNA and use this example to demonstrate the analysis. The protocol comprises the following steps: (i) determination of the secondary structure of the excited state from RD chemical shift data, (ii) design of trapped excited state RNA, (iii) validation of the excited state structure by NMR, and (iv) MCSF analysis comparing the chemical shifts of the trapped excited state with the RD-derived chemical shift differences. MCSF enables observation of the short-lived RNA structures, which can be functionally and structurally characterized by entrapment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Code availability
The custom code used for the local RNA BMRB SQL database query in the Procedure is available at https://github.com/PetzoldLab/mcsf-git along with the SQL database.
References
Mattick, J. S. & Makunin, I. V. Non-coding RNA. Hum. Mol. Genet. 15, R17–R29 (2006).
Cech, T. R. & Steitz, J. A. The noncoding rna revolution—trashing old rules to forge new ones. Cell 157, 77–94 (2014).
Marek, M. S., Johnson-Buck, A. & Walter, N. G. The shape-shifting quasispecies of RNA: one sequence, many functional folds. Phys. Chem. Chem. Phys. 13, 11524–11537 (2011).
Ganser, L. R., Kelly, M. L., Herschlag, D. & Al-Hashimi, H. M. The roles of structural dynamics in the cellular functions of RNAs. Nat. Rev. Mol. Cell Biol. 20, 474–489 (2019).
Mustoe, A. M., Brooks, C. L. & Al-Hashimi, H. M. Hierarchy of RNA functional dynamics. Annu. Rev. Biochem. 83, 441–466 (2014).
Frauenfelder, H., Sligar, S. & Wolynes, P. The energy landscapes and motions of proteins. Science 254, 1598–1603 (1991).
Marušič, M., Schlagnitweit, J. & Petzold, K. RNA dynamics by NMR spectroscopy. ChemBioChem 20, 2685–2710 (2019).
Zhao, B. & Zhang, Q. Measuring residual dipolar couplings in excited conformational states of nucleic acids by CEST NMR spectroscopy. J. Am. Chem. Soc. 137, 13480–13483 (2015).
Al-Hashimi, H. M. & Walter, N. G. RNA dynamics: it is about time. Curr. Opin. Struct. Biol. 18, 321–329 (2008).
Tian, S., Cordero, P., Kladwang, W. & Das, R. High-throughput mutate-map-rescue evaluates SHAPE-directed RNA structure and uncovers excited states. RNA 20, 1815–1826 (2014).
Mulder, F. A. A., Mittermaier, A., Hon, B., Dahlquist, F. W. & Kay, L. E. Studying excited states of proteins by NMR spectroscopy. Nat. Struct. Biol. 8, 932–935 (2001).
Bakolitsa, C. et al. Low-populated folding intermediates of Fyn SH3 characterized by relaxation dispersion NMR. Nature 430, 586–590 (2004).
Nikolova, E. N. et al. Transient Hoogsteen base pairs in canonical duplex DNA. Nature 470, 498–502 (2011).
Kimsey, I. J., Petzold, K., Sathyamoorthy, B., Stein, Z. W. & Al-Hashimi, H. M. Visualizing transient Watson–Crick-like mispairs in DNA and RNA duplexes. Nature 519, 315–320 (2015).
Dethoff, E. A., Petzold, K., Chugh, J., Casiano-Negroni, A. & Al-Hashimi, H. M. Visualizing transient low-populated structures of RNA. Nature 491, 724–728 (2012).
Baisden, J. T., Boyer, J. A., Zhao, B., Hammond, S. M. & Zhang, Q. Visualizing a protonated RNA state that modulates microRNA-21 maturation. Nat. Chem. Biol. 17, 80–88 (2021).
Baronti, L. et al. Base-pair conformational switch modulates miR-34a targeting of Sirt1 mRNA. Nature 583, 139–144 (2020).
Ulrich, E. L. et al. BioMagResBank. Nucleic Acids Res. 36, D402–D408 (2008).
Rangadurai, A., Szymaski, E. S., Kimsey, I. J., Shi, H. & Al-Hashimi, H. M. Characterizing micro-to-millisecond chemical exchange in nucleic acids using off-resonance R 1ρ relaxation dispersion. Prog. Nucl. Mag. Res. Sp. 112–113, 55–102 (2019).
Parisien, M. & Major, F. The MC-Fold and MC-Sym pipeline infers RNA structure from sequence data. Nature 452, 51–55 (2008).
Lukavsky, P. J. & Puglisi, J. D. Structure determination of large biological RNAs. Methods Enzymol. 394, 399–416 (2005).
Karlsson, H., Baronti, L. & Petzold, K. A robust and versatile method for production and purification of large-scale RNA samples for structural biology. RNA 26, 1023–1037 (2020).
Baronti, L., Karlsson, H., Marušič, M. & Petzold, K. A guide to large-scale RNA sample preparation. Anal. Bioanal. Chem. 410, 3239–3252 (2018).
Xue, Y. et al. Chapter two characterizing RNA excited states using NMR relaxation dispersion. Methods Enzymol. 558, 39–73 (2015).
Feyrer, H., Schlagnitweit, J. & Petzold, K. Practical aspects of sample preparation and setup of 1H R1ρ relaxation dispersion experiments of RNA. J. Vis. Exp. https://doi.org/10.3791/62470 (2021).
Beckert, B. & Masquida, B. Synthesis of RNA by in vitro transcription. Methods Mol. Biol. 703, 29–41 (2010).
Milligan, J. F., Groebe, D. R., Witherell, G. W. & Uhlenbeck, O. C. Oligoribonucleotide synthesis using T7 RNA polymerase and synthetic DNA templates. Nucleic Acids Res. 15, 8783–8798 (1987).
Melton, D. A. et al. Efficient in vitro synthesis of biologically active RNA and RNA hybridization probes from plasmids containing a bacteriophage SP6 promoter. Nucleic Acids Res. 12, 7035–7056 (1984).
Feyrer, H., Munteanu, R., Baronti, L. & Petzold, K. One-pot production of RNA in high yield and purity through cleaving tandem transcripts. Molecules 25, 1142 (2020).
Nikonowicz, E. P. et al. Preparation of 13 C and 15 N labelled RNAs for heteronuclear multi-dimensional NMR studies. Nucleic Acids Res. 20, 4507–4513 (1992).
Wyatt, J. R., Chastain, M. & Puglisi, J. D. Synthesis and purification of large amounts of RNA oligonucleotides. Biotechniques 11, 764–769 (1991).
Anderson, A. C., Scaringe, S. A., Earp, B. E. & Frederick, C. A. HPLC purification of RNA for crystallography and NMR. RNA 2, 110–117 (1996).
Murray, J. B., Collier, A. K. & Arnold, J. R. P. A general purification procedure for chemically synthesized oligoribonucleotides. Anal. Biochem. 218, 177–184 (1994).
Strebitzer, E., Nußbaumer, F., Kremser, J., Tollinger, M. & Kreutz, C. Studying sparsely populated conformational states in RNA combining chemical synthesis and solution NMR spectroscopy. Methods 148, 39–47 (2018).
Becette, O., Olenginski, L. T. & Dayie, T. K. Solid-phase chemical synthesis of stable isotope-labeled RNA to aid structure and dynamics studies by NMR spectroscopy. Molecules 24, 3476 (2019).
Strebitzer, E. et al. 5-Oxyacetic acid modification destabilizes double helical stem structures and favors anionic Watson–Crick like cmo5U-G base pairs. Chem. Eur. J. 24, 18903–18906 (2018).
Le, M. T., Brown, R. E., Simon, A. E. & Dayie, T. K. Chapter nineteen in vivo, large-scale preparation of uniformly 15N- and site-specifically 13C-labeled homogeneous, recombinant RNA for NMR studies. Methods Enzymol. 565, 495–535 (2015).
Fürtig, B., Richter, C., Wöhnert, J. & Schwalbe, H. NMR spectroscopy of RNA. ChemBioChem 4, 936–962 (2003).
Lee, J., Dethoff, E. A. & Al-Hashimi, H. M. Invisible RNA state dynamically couples distant motifs. Proc. Natl Acad. Sci. USA 111, 9485–9490 (2014).
Ebrahimi, P., Kaur, S., Baronti, L., Petzold, K. & Chen, A. A. A two-dimensional replica-exchange molecular dynamics method for simulating RNA folding using sparse experimental restraints. Methods 162–163, 96–107 (2019).
Berman, H. M. et al. The Protein Data Bank. Nucleic Acids Res. (2000) https://doi.org/10.1093/nar/28.1.235
Aeschbacher, T., Shubert, M. & Allain, F. H.-T. A procedure to validate and correct the C chemical shift calibration of RNA datasets. J. Biomol. NMR 52, 179–190 (2012).
Wishart, D. S. et al. 1H, 13C and 15N chemical shift referencing in biomolecular NMR. J. Biomol. NMR 6, 135–140 (1995).
Popenda, M. et al. RNA FRABASE 2.0: an advanced web-accessible database with the capacity to search the three-dimensional fragments within RNA structures. BMC Bioinformatics 11, 231 (2010).
Gruber, A. R., Bernhart, S. H. & Lorenz, R. RNA bioinformatics. Methods Mol. Biol. 1269, 307–326 (2014).
Jin, X., Zhu, T., Zhang, J. Z. H. & He, X. A systematic study on RNA NMR chemical shift calculation based on the automated fragmentation QM/MM approach. RSC Adv. 6, 108590–108602 (2016).
Hopkins, N. & Riad, M. Mutate-and-Chemical-Shift-Fingerprint (MCSF) to characterize excited states in RNA using NMR spectroscopy. Petzoldlab/mcsf-git, https://doi.org/10.5281/zenodo.4823710 (2021).
Dominguez, C., Schubert, M., Duss, O., Ravindranathan, S. & Allain, F. H.-T. Structure determination and dynamics of protein–RNA complexes by NMR spectroscopy. Prog. Nucl. Mag. Res. Sp. 58, 1–61 (2011).
Farès, C., Amata, I. & Carlomagno, T. 13 C-Detection in RNA bases: revealing structure−chemical shift relationships. J. Am. Chem. Soc. 129, 15814–15823 (2007).
Schlagnitweit, J., Steiner, E., Karlsson, H. & Petzold, K. Efficient detection of structure and dynamics in unlabeled RNAs: the SELOPE approach. Chem. Eur. J. 24, 6067–6070 (2018).
Dingley, A. & Grzesiek, S. Direct observation of hydrogen bonds in nucleic acid base pairs by internucleotide (2)J(NN) couplings. J. Am. Chem. Soc. 120, 8293–8297 (1998).
Wang, Y., Han, G., Jiang, X., Yuwen, T. & Xue, Y. Chemical shift prediction of RNA imino groups: application toward characterizing RNA excited states. Nat. Commun. 12, 1595 (2021).
Nikolova, E. N., Gottardo, F. L. & Al-Hashimi, H. M. Probing transient Hoogsteen hydrogen bonds in canonical duplex DNA using NMR relaxation dispersion and single-atom substitution. J. Am. Chem. Soc. 134, 3667–3670 (2012).
Hansen, A. L., Nikolova, E. N., Casiano-Negroni, A. & Al-Hashimi, H. M. Extending the range of microsecond-to-millisecond chemical exchange detected in labeled and unlabeled nucleic acids by selective carbon R 1ρ NMR spectroscopy. J. Am. Chem. Soc. 131, 3818–3819 (2009).
Steiner, E., Schlagnitweit, J., Lundström, P. & Petzold, K. Capturing excited states in the fast-intermediate exchange limit in biological systems using 1H NMR spectroscopy. Angew. Chem. Int. Ed. 55, 15869–15872 (2016).
Palmer, A. G. & Massi, F. Characterization of the dynamics of biomacromolecules using rotating-frame spin relaxation NMR spectroscopy. Chem. Rev. 106, 1700–1719 (2006).
Yamakuchi, M. & Lowenstein, C. J. MiR-34, SIRT1, and p53: the feedback loop. Cell Cycle 8, 712–715 (2009).
Korzhnev, D. M., Orekhov, V. Y. & Kay, L. E. Off-resonance R 1ρ NMR studies of exchange dynamics in proteins with low spin-lock fields: an application to a Fyn SH3 domain. J. Am. Chem. Soc. 127, 713–721 (2005).
McConnell, H. M. Reaction rates by nuclear magnetic resonance. J. Chem. Phys. 28, 430–431 (1958).
Trott, O. & Palmer, A. G. R1ρ relaxation outside of the fast-exchange limit. J. Magn. Reson. 154, 157–160 (2002).
Dejaegere, A. P. & Case, D. A. Density functional study of ribose and deoxyribose chemical shifts. J. Phys. Chem. 102, 5280–5289 (1998).
Ebrahimi, M., Rossi, P., Rogers, C. & Harbison, G. S. Dependence of 13C NMR chemical shifts on conformations of RNA nucleosides and nucleotides. J. Magn. Reson. 150, 1–9 (2001).
Xu, X.-P. & Au−Yeung, S. C. F. Investigation of chemical shift and structure relationships in nucleic acids using NMR and density functional theory methods. J. Phys. Chem. B 104, 5641–5650 (2000).
Clay, M. C., Ganser, L. R., Merriman, D. K. & Al-Hashimi, H. M. Resolving sugar puckers in RNA excited states exposes slow modes of repuckering dynamics. Nucleic Acids Res. 45, e134 (2017).
Leontis, N. B. & Westhof, E. Geometric nomenclature and classification of RNA base pairs. RNA 7, 499–512 (2001).
Antao, V. P., Lai, S. Y. & Tinoco, I. A thermodynamic study of unusually stable RNA and DNA hairpins. Nucleic Acids Res. 19, 5901–5905 (1991).
Nozinovic, S., Fürtig, B., Jonker, H. R. A., Richter, C. & Schwalbe, H. High-resolution NMR structure of an RNA model system: the 14-mer cUUCGg tetraloop hairpin RNA. Nucleic Acids Res. 38, 683–694 (2010).
Varani, G., Cheong, C. & Tinoco, I. Solution structure of an unusually stable RNA hairpin, 5GGAC(UUCG)GUCC. Nature 346, 680–682 (1990).
Ghose, R., Marino, J. P., Wiberg, K. B. & Prestegard, J. H. Dependence of 13C chemical shifts on glycosidic torsional angles in ribonucleic acids. J. Am. Chem. Soc. 116, 8827–8828 (1994).
Acknowledgements
K.P. acknowledges funding from the Swedish Research Council (grant numbers 2014-04303 and 2018-00250), the Swedish Foundation for Strategic Research (project number ICA14-0023), Harald och Greta Jeansson Stiftelse (JS20140009), Carl Tryggers stiftelse (CTS14-383 and 15-383), Eva och, Oscar Ahréns Stiftelse, Åke Wiberg Stiftelse (467080968 and M14-0109), Cancerfonden (CAN 2015/388), the Karolinska Institute Department of Medical Biochemistry and Biophysics (grant number KID 2-3707/2013 and support for the purchase of a 600-MHz Bruker NMR spectrometer) and Ragnar Söderberg Stiftelse (M91/14). J.S. acknowledges funding through a Marie Sklodowska-Curie Individual Fellowship (EU H2020/project number 747446).
Author information
Authors and Affiliations
Contributions
M.R. and K.P. wrote the manuscript. N.H. and M.R. wrote the scripts based on earlier work done by N.H., H.K. and L.B. L.B. carried out most of the experiments of the original manuscript and data analysis, with assistance from K.P. and J.S. All authors contributed to the final version.
Corresponding author
Ethics declarations
Competing interests
K.P. is a consultant to Arrakis Therapeutics, an RNA-targeting drug-discovery company.
Additional information
Peer review information Nature Protocols thanks Diego Carnevale, Christoph Kreutz and the other, anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Baronti, L. et al. Nature 583, 139–144 (2020): https://doi.org/10.1038/s41586-020-2336-3
Schlagnitweit, J. et al. Chem. European J. 24, 6067–6070 (2018): https://doi.org/10.1002/chem.201800992
Steiner, E. et al. Angew. Chem. Int. Ed. 55, 15869–15872 (2016): https://doi.org/10.1002/anie.201609102
Rights and permissions
About this article
Cite this article
Riad, M., Hopkins, N., Baronti, L. et al. Mutate-and-chemical-shift-fingerprint (MCSF) to characterize excited states in RNA using NMR spectroscopy. Nat Protoc 16, 5146–5170 (2021). https://doi.org/10.1038/s41596-021-00606-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-021-00606-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.