Abstract
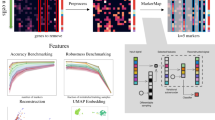
Single-cell RNA-sequencing data have significantly advanced the characterization of cell-type diversity and composition. However, cell-type definitions vary across data and analysis pipelines, raising concerns about cell-type validity and generalizability. With MetaNeighbor, we proposed an efficient and robust quantification of cell-type replicability that preserves dataset independence and is highly scalable compared to dataset integration. In this protocol, we show how MetaNeighbor can be used to characterize cell-type replicability by following a simple three-step procedure: gene filtering, neighbor voting and visualization. We show how these steps can be tailored to quantify cell-type replicability, determine gene sets that contribute to cell-type identity and pretrain a model on a reference taxonomy to rapidly assess newly generated data. The protocol is based on an open-source R package available from Bioconductor and GitHub, requires basic familiarity with Rstudio or the R command line and can typically be run in <5 min for millions of cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
















Similar content being viewed by others
Data availability
The datasets analyzed in the protocol are all previously published and publicly available. Human pancreas datasets were from Baron et al.33 (Gene Expression Omnibus (GEO) accession code GSE84133), Lawlor et al.34 (GEO accession code GSE86473), Muraro et al.35 (GEO accession code GSE85241) and Segerstolpe et al.36 (ArrayExpress accession code E-MTAB-5061). These datasets are accessed through the Bioconductor scRNAseq library in the protocol. The mouse primary visual cortex dataset was from Tasic et al.32 (GEO accession code GSE71585), accessed through the Bioconductor scRNAseq library. The BICCN dataset for the mouse primary motor cortex from Yao et al.4 is available on the Neuroscience Multi-Omic archive (https://assets.nemoarchive.org/dat-ch1nqb7). The subset of the BICCN data necessary to run the protocol is also available on FigShare at https://doi.org/10.6084/m9.figshare.13020569 (R version) and https://doi.org/10.6084/m9.figshare.13034171 (Python version).
Code availability
The code for the procedures (including all figures) is freely available on GitHub at https://github.com/gillislab/MetaNeighbor-Protocol in multiple formats (Rmd, PDF and jupyter notebook for R and Python). The scripts used to generate the protocol data are available in the same repository. The stable R version of MetaNeighbor is available through Bioconductor (https://www.bioconductor.org/install/) at https://www.bioconductor.org/packages/release/bioc/html/MetaNeighbor.html (the protocol was generated by using version 3.12), and the development versions are available on GitHub at https://github.com/gillislab/MetaNeighbor (R version) and https://github.com/gillislab/pyMN (Python version).
References
Hay, S. B., Ferchen, K., Chetal, K., Grimes, H. L. & Salomonis, N. The Human Cell Atlas bone marrow single-cell interactive web portal. Exp. Hematol. 68, 51–61 (2018).
Schaum, N. et al. Single-cell transcriptomics of 20 mouse organs creates a Tabula Muris. Nature 562, 367–372 (2018).
Almanzar, N. et al. A single-cell transcriptomic atlas characterizes ageing tissues in the mouse. Nature 583, 590–595 (2020).
Yao, Z. et al. An integrated transcriptomic and epigenomic atlas of mouse primary motor cortex cell types. Nature (in the press).
Yao, Z. et al. A taxonomy of transcriptomic cell types across the isocortex and hippocampal formation. Cell (in the press).
Bakken, T. E. et al. Evolution of cellular diversity in primary motor cortex of human, marmoset monkey, and mouse. Nature (in the press).
Duò, A., Robinson, M. D. & Soneson, C. A systematic performance evaluation of clustering methods for single-cell RNA-seq data. F1000Res. 7, 1141 (2018).
Haghverdi, L., Lun, A. T. L., Morgan, M. D. & Marioni, J. C. Batch effects in single-cell RNA-sequencing data are corrected by matching mutual nearest neighbors. Nat. Biotechnol. 36, 421–427 (2018).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e21 (2019).
Welch, J. D. et al. Single-cell multi-omic integration compares and contrasts features of brain cell identity. Cell 177, 1873–1887.e17 (2019).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Hie, B., Bryson, B. & Berger, B. Efficient integration of heterogeneous single-cell transcriptomes using Scanorama. Nat. Biotechnol. 37, 685–691 (2019).
Barkas, N. et al. Joint analysis of heterogeneous single-cell RNA-seq dataset collections. Nat. Methods 16, 695–698 (2019).
Luo, C. et al. Single nucleus multi-omics links human cortical cell regulatory genome diversity to disease risk variants. Preprint at bioRxiv https://doi.org/10.1101/2019.12.11.873398 (2019).
Crow, M., Paul, A., Ballouz, S., Huang, Z. J. & Gillis, J. Characterizing the replicability of cell types defined by single cell RNA-sequencing data using MetaNeighbor. Nat. Commun. 9, 884 (2018).
Paul, A. et al. Transcriptional architecture of synaptic communication delineates GABAergic neuron identity. Cell 171, 522–539.e20 (2017).
Hodge, R. D. et al. Conserved cell types with divergent features in human versus mouse cortex. Nature 573, 61–68 (2019).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Forcato, M., Romano, O. & Bicciato, S. Computational methods for the integrative analysis of single-cell data. Brief. Bioinform. 22, 20–29 (2020).
Hie, B. et al. Computational methods for single-cell RNA sequencing. Annu. Rev. Biomed. Data Sci. 3, 339–364 (2020).
Tran, H. T. N. et al. A benchmark of batch-effect correction methods for single-cell RNA sequencing data. Genome Biol. 21, 12 (2020).
Luecken, M. D. et al. Benchmarking atlas-level data integration in single-cell genomics. Preprint at bioRxiv https://doi.org/10.1101/2020.05.22.111161 (2020).
Abdelaal, T. et al. A comparison of automatic cell identification methods for single-cell RNA sequencing data. Genome Biol. 20, 194 (2019).
Kiselev, V. Y., Yiu, A. & Hemberg, M. scmap: projection of single-cell RNA-seq data across data sets. Nat. Methods 15, 359–362 (2018).
Büttner, M., Miao, Z., Wolf, F. A., Teichmann, S. A. & Theis, F. J. A test metric for assessing single-cell RNA-seq batch correction. Nat. Methods 16, 43–49 (2019).
Kapp, A. V. & Tibshirani, R. Are clusters found in one dataset present in another dataset? Biostatistics 8, 9–31 (2007).
Dudoit, S., Fridlyand, J. & Speed, T. P. Comparison of discrimination methods for the classification of tumors using gene expression data. J. Am. Stat. Assoc. 97, 77–87 (2002).
Kiselev, V. Y. et al. SC3: consensus clustering of single-cell RNA-seq data. Nat. Methods 14, 483–486 (2017).
Tasic, B. et al. Shared and distinct transcriptomic cell types across neocortical areas. Nature 563, 72–78 (2018).
gillislab/MetaNeighbor-Protocol. https://github.com/gillislab/MetaNeighbor (2020).
Protocol data (R version). https://doi.org/10.6084/m9.figshare.13020569.v2 (2020).
Tasic, B. et al. Adult mouse cortical cell taxonomy revealed by single cell transcriptomics. Nat. Neurosci. 19, 335–346 (2016).
Baron, M. et al. A single-cell transcriptomic map of the human and mouse pancreas reveals inter- and intra-cell population structure. Cell Syst. 3, 346–360.e4 (2016).
Lawlor, N. et al. Single-cell transcriptomes identify human islet cell signatures and reveal cell-type-specific expression changes in type 2 diabetes. Genome Res. 27, 208–222 (2017).
Muraro, M. J. et al. A single-cell transcriptome atlas of the human pancreas. Cell Syst. 3, 385–394.e3 (2016).
Segerstolpe, Å. et al. Single-cell transcriptome profiling of human pancreatic islets in health and type 2 diabetes. Cell Metab. 24, 593–607 (2016).
Acknowledgements
J.G. was supported by NIH grants R01MH113005 and R01LM012736. S.F. was supported by NIH grant U19MH114821. B.D.H. was supported by the CSHL Crick Cray Fellowship. M.C. was supported by NIH grant K99MH120050.
Author information
Authors and Affiliations
Contributions
S.F., M.C., B.D.H. and J.G. designed the experiments, performed the data analysis and wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks Praneet Chaturvedi, Guoji Guo, Ahmed Mahfouz, Nathan Salomonis and Daniel Schnell for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Crow, M. et al. Nat. Commun. 9, 884 (2018): https://doi.org/10.1038/s41467-018-03282-0
Paul, A. et al. Cell 171, 522–539.e20 (2017): https://doi.org/10.1016/j.cell.2017.08.032
Yao, Z. et al. Preprint at bioRxiv (2020): https://doi.org/10.1101/2020.02.29.970558
Bakken, T. E. et al. Preprint at bioRxiv (2020): https://doi.org/10.1101/2020.03.31.016972
Key data used in this protocol
Yao, Z. et al. Preprint at bioRxiv (2020) https://doi.org/10.1101/2020.02.29.970558
Baron, M. et al. Cell Syst. 3, 346–360.e4 (2016) https://doi.org/10.1016/j.cels.2016.08.011
Lawlor, N. et al. Genome Res. 27, 208–222 (2017) https://doi.org/10.1101/gr.212720.116
Muraro, M. J. et al. Cell Syst. 3, 385–394.e3 (2016) https://doi.org/10.1016/j.cels.2016.09.002
Segerstolpe, Å. et al. Cell Metab. 24, 593–607 (2016) https://doi.org/10.1016/j.cmet.2016.08.020
Tasic, B. et al. Nat. Neurosci. 19, 335–346 (2016) https://doi.org/10.1038/nn.4216
Rights and permissions
About this article
Cite this article
Fischer, S., Crow, M., Harris, B.D. et al. Scaling up reproducible research for single-cell transcriptomics using MetaNeighbor. Nat Protoc 16, 4031–4067 (2021). https://doi.org/10.1038/s41596-021-00575-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-021-00575-5
This article is cited by
-
A spatially-resolved transcriptional atlas of the murine dorsal pons at single-cell resolution
Nature Communications (2024)
-
Comparative single-cell transcriptomic analysis of primate brains highlights human-specific regulatory evolution
Nature Ecology & Evolution (2023)
-
Cell type specific transcriptomic differences in depression show similar patterns between males and females but implicate distinct cell types and genes
Nature Communications (2023)
-
A pan-grass transcriptome reveals patterns of cellular divergence in crops
Nature (2023)
-
Systematic identification of cell-fate regulatory programs using a single-cell atlas of mouse development
Nature Genetics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.