Abstract
Tissue clearing has become a powerful technique for studying anatomy and morphology at scales ranging from entire organisms to subcellular features. With the recent proliferation of tissue-clearing methods and imaging options, it can be challenging to determine the best clearing protocol for a particular tissue and experimental question. The fact that so many clearing protocols exist suggests there is no one-size-fits-all approach to tissue clearing and imaging. Even in cases where a basic level of clearing has been achieved, there are many factors to consider, including signal retention, staining (labeling), uniformity of transparency, image acquisition and analysis. Despite reviews citing features of clearing protocols, it is often unknown a priori whether a protocol will work for a given experiment, and thus some optimization is required by the end user. In addition, the capabilities of available imaging setups often dictate how the sample needs to be prepared. After imaging, careful evaluation of volumetric image data is required for each combination of clearing protocol, tissue type, biological marker, imaging modality and biological question. Rather than providing a direct comparison of the many clearing methods and applications available, in this tutorial we address common pitfalls and provide guidelines for designing, optimizing and imaging in a successful tissue-clearing experiment with a focus on light-sheet fluorescence microscopy (LSFM).
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Gradinaru, V., Treweek, J., Overton, K. & Deisseroth, K. Hydrogel-tissue chemistry: principles and applications. Annu. Rev. Biophys. 47, 355–376 (2018).
Susaki, E. A. et al. Versatile whole-organ/body staining and imaging based on electrolyte-gel properties of biological tissues. Nat. Commun. 11, 1982 (2020).
Ueda, H. R. et al. Tissue clearing and its applications in neuroscience. Nat. Rev. Neurosci. 21, 61–79 (2020).
Ertürk, A. et al. Three-dimensional imaging of the unsectioned adult spinal cord to assess axon regeneration and glial responses after injury. Nat. Med. 18, 166–171 (2012).
Lerner, T. N. et al. Intact-brain analyses reveal distinct information carried by SNc dopamine subcircuits. Cell 162, 635–647 (2015).
Adhikari, A. et al. Basomedial amygdala mediates top-down control of anxiety and fear. Nature 527, 179–185 (2015).
Belle, M. et al. Tridimensional visualization and analysis of early human development. Cell 169, 161–173.e12 (2017).
Acar, M. et al. Deep imaging of bone marrow shows non-dividing stem cells are mainly perisinusoidal. Nature 526, 126–130 (2015).
Chen, J. Y. et al. Hoxb5 marks long-term haematopoietic stem cells and reveals a homogenous perivascular niche. Nature 530, 223–227 (2016).
Oshimori, N., Oristian, D. & Fuchs, E. TGF-β promotes heterogeneity and drug resistance in squamous. cell carcinoma. Cell 160, 963–976 (2015).
von Neubeck, B. et al. An inhibitory antibody targeting carbonic anhydrase XII abrogates chemoresistance and significantly reduces lung metastases in an orthotopic breast cancer model in vivo. Int. J. Cancer 143, 2065–2075 (2018).
Tanaka, N. et al. Publisher Correction: whole-tissue biopsy phenotyping of three-dimensional tumours reveals patterns of cancer heterogeneity. Nat. Biomed. Eng. 1, 1 (2018).
Henning, Y., Osadnik, C. & Malkemper, E. P. EyeCi: optical clearing and imaging of immunolabeled mouse eyes using light-sheet fluorescence microscopy. Exp. Eye Res. 180, 137–145 (2019).
Johnson, S. B., Schmitz, H. M. & Santi, P. A. TSLIM imaging and a morphometric analysis of the mouse spiral ganglion. Hear. Res. 278, 34–42 (2011).
Yang, B. et al. Single-cell phenotyping within transparent intact tissue through whole-body clearing. Cell 158, 945–958 (2014).
Spalteholz, W. Über das Durchsichtigmachen von menschlichen und tierischen Präparaten (Leipzig: S. Hierzel). Leipzig (1914).
Costantini, I., Cicchi, R., Silvestri, L., Vanzi, F. & Pavone, F. S. In-vivo and ex-vivo optical clearing methods for biological tissues: review. Biomed. Opt. Express 10, 5251 (2019).
Dodt, H. U. et al. Ultramicroscopy: three-dimensional visualization of neuronal networks in the whole mouse brain. Nat. Methods 4, 331–336 (2007).
Ertürk, A. et al. Three-dimensional imaging of solvent-cleared organs using 3DISCO. Nat. Protoc. 7, 1983–1995 (2012).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Masselink, W. et al. Broad applicability of a streamlined ethyl cinnamate-based clearing procedure. Development 146, dev166884 (2019).
Chung, K. & Deisseroth, K. CLARITY for mapping the nervous system. Nat. Methods 10, 508–513 (2013).
Du, H., Hou, P., Zhang, W. & Li, Q. Advances in CLARITY based tissue clearing and imaging (review). Exp. Ther. Med. 16, 1567–1576 (2018).
Hama, H. et al. Scale: a chemical approach for fluorescence imaging and reconstruction of transparent mouse brain. Nat. Neurosci. 14, 1481–1488 (2011).
Tainaka, K. et al. Whole-body imaging with single-cell resolution by tissue decolorization. Cell 159, 911–924 (2014).
Tainaka, K. et al. Chemical landscape for tissue clearing based on hydrophilic reagents. Cell Rep 24, 2196–2210.e9 (2018).
Ke, M. T., Fujimoto, S. & Imai, T. SeeDB: a simple and morphology-preserving optical clearing agent for neuronal circuit reconstruction. Nat. Neurosci. 16, 1154–1161 (2013).
Hou, B. et al. Scalable and DiI-compatible optical clearance of the mammalian brain. Front. Neuroanat. 9, (2015).
Aoyagi, Y., Kawakami, R., Osanai, H., Hibi, T. & Nemoto, T. A rapid optical clearing protocol using 2,2′-thiodiethanol for microscopic observation of fixed mouse brain. PLoS ONE 10, e0116280 (2015).
Lai, H. M. et al. Next generation histology methods for three-dimensional imaging of fresh and archival human brain tissues. Nat. Commun. 9, 1066 (2018).
Chen, L. et al. UbasM: an effective balanced optical clearing method for intact biomedical imaging. Sci. Rep. 7, 12218 (2017).
Rios, A. C. et al. Intraclonal plasticity in mammary tumors revealed through large-scale single-cell resolution 3D imaging. Cancer Cell 35, 618–632.e6 (2019).
Richardson, D. S. & Lichtman, J. W. SnapShot: tissue clearing. Cell 171, 496–496.e1 (2017).
Tainaka, K., Kuno, A., Kubota, S. I., Murakami, T. & Ueda, H. R. Chemical principles in tissue clearing and staining protocols for whole-body cell profiling. Annu. Rev. Cell Dev. Biol. 32, 713–741 (2016).
Silvestri, L., Costantini, I., Sacconi, L. & Pavone, F. S. Clearing of fixed tissue: a review from a microscopist’s perspective. J. Biomed. Opt. 21, 081205 (2016).
Ariel, P. A beginner’s guide to tissue clearing. Int. J. Biochem. Cell Biol 84, 35–39 (2017).
Seo, J., Choe, M. & Kim, S. Y. Clearing and labeling techniques for large-scale biological tissues. Mol. Cells 39, 439–446 (2016).
Azaripour, A. et al. A survey of clearing techniques for 3D imaging of tissues with special reference to connective tissue. Prog. Histochem. Cytochem. 51, 9–23 (2016).
Yu, T., Qi, Y., Gong, H., Luo, Q. & Zhu, D. Optical clearing for multiscale biological tissues. J. Biophotonics 11, e201700187 (2018).
Richardson, D. S. & Lichtman, J. W. Clarifying tissue clearing. Cell 162, 246–257 (2015).
Li, A. et al. Micro-optical sectioning tomography to obtain a high-resolution atlas of the mouse brain. Science 330, 1404–1408 (2010).
Denk, W., Strickler, J. H. & Webb, W. W. Two-photon laser scanning fluorescence microscopy. Science 248, 73–76 (1990).
Ragan, T. et al. Serial two-photon tomography for automated ex vivo mouse brain imaging. Nat. Methods 9, 255–258 (2012).
Jonkman, J., Brown, C. M., Wright, G. D., Anderson, K. I. & North, A. J. Tutorial: guidance for quantitative confocal microscopy. Nat. Protoc. 15, 1585–1611 (2020).
Matryba, P. et al. Systematic evaluation of chemically distinct tissue optical clearing techniques in murine lymph nodes. J. Immunol. 204, 1395–1407 (2020).
Wan, P. et al. Evaluation of seven optical clearing methods in mouse brain. Neurophotonics 5, 1 (2018).
Matryba, P., Kaczmarek, L. & Gołąb, J. Advances in ex situ tissue optical clearing. Laser Photon. Rev. 13, 1800292 (2019).
Gómez-Gaviro, M. V., Sanderson, D., Ripoll, J. & Desco, M. Biomedical applications of tissue clearing and three-dimensional imaging in health and disease. iScience 23, 101432 (2020).
Chung, K. et al. Structural and molecular interrogation of intact biological systems. Nature 497, 332–337 (2013).
Greenbaum, A. et al. Bone CLARITY: clearing, imaging, and computational analysis of osteoprogenitors within intact bone marrow. Sci. Transl. Med. 9, eaah6518 (2017).
Chi, J., Crane, A., Wu, Z. & Cohen, P. Adipo-Clear: a tissue clearing method for three-dimensional imaging of adipose tissue. J. Vis. Exp. 2018, (2018).
Pende, M. et al. A versatile depigmentation, clearing, and labeling method for exploring nervous system diversity. Sci. Adv. 6, eaba0365 (2020).
Chen, F., Tillberg, P. W. & Boyden, E. S. Expansion microscopy. Science 347, 543–548 (2015).
Ku, T. et al. Multiplexed and scalable super-resolution imaging of three-dimensional protein localization in size-adjustable tissues. Nat. Biotechnol. 34, 973–981 (2016).
Renier, N. et al. Mapping of brain activity by automated volume analysis of immediate early genes. Cell 165, 1789–1802 (2016).
Cai, R. et al. Panoptic imaging of transparent mice reveals whole-body neuronal projections and skull–meninges connections. Nat. Neurosci. 22, 317–327 (2019).
Zhao, S. et al. Cellular and molecular probing of intact human organs. Cell 180, 796–812.e19 (2020).
Yun, D. H. et al. Ultrafast immunostaining of organ-scale tissues for scalable proteomic phenotyping. Preprint at bioRxiv https://doi.org/10.1101/660373 (2019).
Park, Y. G. et al. Protection of tissue physicochemical properties using polyfunctional crosslinkers. Nat. Biotechnol. 37, 73 (2019).
Hama, H. et al. ScaleS: an optical clearing palette for biological imaging. Nat. Neurosci. 18, 1518–1529 (2015).
Perin, P., Voigt, F. F., Bethge, P., Helmchen, F. & Pizzala, R. iDISCO+ for the study of neuroimmune architecture of the rat auditory brainstem. Front. Neuroanat. 13, (2019).
Kim, S. Y. et al. Stochastic electrotransport selectively enhances the transport of highly electromobile molecules. Proc. Natl Acad. Sci. USA 112, E6274–E6283 (2015).
Lee, E. et al. ACT-PRESTO: rapid and consistent tissue clearing and labeling method for 3-dimensional (3D) imaging. Sci. Rep. 6, 18631 (2016).
Murray, E. et al. Simple, scalable proteomic imaging for high-dimensional profiling of intact systems. Cell 163, 1500–1514 (2015).
Vogel, A. T., Marqués, G. & Sanders, M. A. Microwave-assisted fixation, labeling and clearing for optical microscopy of thick specimens. Microsc. Microanal. 19, 16–17 (2013).
Treweek, J. B. et al. Whole-body tissue stabilization and selective extractions via tissue-hydrogel hybrids for high-resolution intact circuit mapping and phenotyping. Nat. Protoc. 10, 1860–1896 (2015).
Epp, J. R. et al. Optimization of CLARITY for clearing whole-brain and other intact organs. eNeuro 2, ENEURO.0022-15.2015 (2015).
Magliaro, C. et al. Clarifying CLARITY: quantitative optimization of the diffusion based delipidation protocol for genetically labeled tissue. Front. Neurosci. 10, (2016).
Pan, C. et al. Shrinkage-mediated imaging of entire organs and organisms using uDISCO. Nat. Methods 13, 859–867 (2016).
Feng, G. et al. Imaging neuronal subsets in transgenic mice expressing multiple spectral variants of GFP. Neuron 28, 41–51 (2000).
Bedbrook, C. N., Deverman, B. E. & Gradinaru, V. Viral strategies for targeting the central and peripheral nervous systems. Ann. Rev. Neurosci. 41, 323–348 (2018).
Challis, R. C. et al. Systemic AAV vectors for widespread and targeted gene delivery in rodents. Nat. Protoc. 14, 379–414 (2019).
Livet, J. et al. Transgenic strategies for combinatorial expression of fluorescent proteins in the nervous system. Nature 450, 56–62 (2007).
Sereti, K.-I. et al. Analysis of cardiomyocyte clonal expansion during mouse heart development and injury. Nat. Commun. 9, 754 (2018).
Jing, D. et al. Tissue clearing of both hard and soft tissue organs with the pegasos method. Cell Res 28, 803–818 (2018).
Li, J., Czajkowsky, D. M., Li, X. & Shao, Z. Fast immuno-labeling by electrophoretically driven infiltration for intact tissue imaging. Sci. Rep. 5, 10640 (2015).
Sakaguchi, R., Leiwe, M. N. & Imai, T. Bright multicolor labeling of neuronal circuits with fluorescent proteins and chemical tags. eLlife 7, e40350 (2018).
Gleave, J. A., Lerch, J. P., Henkelman, R. M. & Nieman, B. J. A method for 3D immunostaining and optical imaging of the mouse brain demonstrated in neural progenitor cells. PLoS ONE 8, e72039 (2013).
Sillitoe, R. V. & Hawkes, R. Whole-mount immunohistochemistry: a high-throughput screen for patterning defects in the mouse cerebellum. J. Histochem. Cytochem. 50, 235–244 (2002).
Tomer, R., Ye, L., Hsueh, B. & Deisseroth, K. Advanced CLARITY for rapid and high-resolution imaging of intact tissues. Nat. Protoc. 9, 1682–1697 (2014).
Ryan, D. P. et al. Automatic and adaptive heterogeneous refractive index compensation for light-sheet microscopy. Nat. Commun. 8, 612 (2017).
Boothe, T. et al. A tunable refractive index matching medium for live imaging cells, tissues and model organisms. eLife 6, e27240 (2017).
Glaser, A. K. et al. Multi-immersion open-top light-sheet microscope for high-throughput imaging of cleared tissues. Nat. Commun. 10, 2781 (2019).
Baschong, W., Suetterlin, R. & Hubert Laeng, R. Control of autofluorescence of archival formaldehyde-fixed, paraffin-embedded tissue in confocal laser scanning microscopy (CLSM). J. Histochem. Cytochem. 49, 1565–1571 (2001).
Davis, A. S. et al. Characterizing and diminishing autofluorescence in formalin-fixed paraffin-embedded human respiratory tissue. J. Histochem. Cytochem. 62, 405–423 (2014).
Neumann, M. & Gabel, D. Simple method for reduction of autofluorescence in fluorescence microscopy. J. Histochem. Cytochem. 50, 437–439 (2002).
Zhu, J. et al. MACS: rapid aqueous clearing system for 3D mapping of intact organs. Adv. Sci. 7, 1903185 (2020).
Schnell, S. A., Staines, W. A. & Wessendorf, M. W. Reduction of lipofuscin-like autofluorescence in fluorescently labeled tissue. J. Histochem. Cytochem. 47, 719–730 (1999).
Romijn, H. J. et al. Double immunolabeling of neuropeptides in the human hypothalamus as analyzed by confocal laser scanning fluorescence microscopy. J. Histochem. Cytochem. 47, 229–235 (1999).
Whittington, N. C. & Wray, S. Suppression of red blood cell autofluorescence for immunocytochemistry on fixed embryonic mouse tissue. Curr. Protoc. Neurosci. 81, 2.28.1–2.28.12 (2017).
Pende, M. et al. High-resolution ultramicroscopy of the developing and adult nervous system in optically cleared Drosophila melanogaster. Nat. Commun. 9, 4731 (2018).
Clancy, B. & Cauller, L. J. Reduction of background autofluorescence in brain sections following immersion in sodium borohydride. J. Neurosci. Methods 83, 97–102 (1998).
Duong, H. & Han, M. A multispectral LED array for the reduction of background autofluorescence in brain tissue. J. Neurosci. Methods 220, 46–54 (2013).
Croce, A. C. & Bottiroli, G. Autofluorescence spectroscopy and imaging: a tool for biomedical research and diagnosis. Eur. J. Histochem. 58, 320–337 (2014).
Schmid, B. et al. 3Dscript: animating 3D/4D microscopy data using a natural-language-based syntax. Nat. Methods 16, 278–280 (2019).
Voigt, F. F. et al. The mesoSPIM initiative: open-source light-sheet microscopes for imaging cleared tissue. Nat. Methods 16, 1105–1108 (2019).
Huisken, J., Swoger, J., Del Bene, F., Wittbrodt, J. & Stelzer, E. H. K. Optical sectioning deep inside live embryos by selective plane illumination microscopy. Science 305, 1007–1009 (2004).
Voie, A. H., Burns, D. H. & Spelman, F. A. Orthogonal‐plane fluorescence optical sectioning: three‐dimensional imaging of macroscopic biological specimens. J. Microsc. 170, 229–236 (1993).
Glaser, A. K. et al. Light-sheet microscopy for slide-free non-destructive pathology of large clinical specimens. Nat. Biomed. Eng. 1, 0084 (2017).
Chakraborty, T. et al. Light-sheet microscopy of cleared tissues with isotropic, subcellular resolution. Nat. Methods 16, 1109–1113 (2019).
Dodt, H.-U. et al. Ultramicroscopy: development and outlook. Neurophotonics 2, 041407 (2015).
Moatti, A. et al. Three-dimensional imaging of intact porcine cochlea using tissue clearing and custom-built light-sheet microscopy. Biomed. Opt. Express 11, 6181 (2020).
Diel, E. E., Lichtman, J. W. & Richardson, D. S. Tutorial: avoiding and correcting sample-induced spherical aberration artifacts in 3D fluorescence microscopy. Nat. Protoc. 15, 2773–2784 (2020).
Power, R. M. & Huisken, J. A guide to light-sheet fluorescence microscopy for multiscale imaging. Nat. Methods 14, 360–373 (2017).
Glaser, A., Bishop, K., Barner, L., Serafin, R. & Liu, J. A hybrid open-top light-sheet microscope for multi-scale imaging of cleared tissues. Preprint at bioRxiv https://doi.org/10.1101/2020.05.06.081745 (2020).
Sharpe, J. et al. Optical projection tomography as a tool for 3D microscopy and gene expression studies. Science 296, 541–545 (2002).
Sharpe, J. Optical projection tomography. in Advanced Imaging in Biology and Medicine: Technology, Software Environments, Applications (eds. Sensen, C. W. & Hallgrimsson, B.)199–224 (2009).
Remacha, E., Friedrich, L., Vermot, J. & Fahrbach, F. O. How to define and optimize axial resolution in light-sheet microscopy: a simulation-based approach. Biomed. Opt. Express 11, 8 (2020).
Chang, B.-J., Dean, K. M. & Fiolka, R. Systematic and quantitative comparison of lattice and Gaussian light-sheets. Opt. Express 28, 27052 (2020).
Schacht, P., Johnson, S. B. & Santi, P. A. Implementation of a continuous scanning procedure and a line scan camera for thin-sheet laser imaging microscopy. Biomed. Opt. Express 1, 598 (2010).
Buytaert, J. A. N. & Dirckx, J. J. J. Design and quantitative resolution measurements of an optical virtual sectioning three-dimensional imaging technique for biomedical specimens, featuring two-micrometer slicing resolution. J. Biomed. Opt. 12, 014039 (2007).
Dean, K. M., Roudot, P., Welf, E. S., Danuser, G. & Fiolka, R. Deconvolution-free subcellular imaging with axially swept light sheet microscopy. Biophys. J. 108, 2807–2815 (2015).
Fu, Q., Martin, B. L., Matus, D. Q. & Gao, L. Imaging multicellular specimens with real-time optimized tiling light-sheet selective plane illumination microscopy. Nat. Commun. 7, 11088 (2016).
Huisken, J. & Stainier, D. Y. R. Even fluorescence excitation by multidirectional selective plane illumination microscopy (mSPIM). Opt. Lett. 32, 2608 (2007).
Keller, P. J., Schmidt, A. D., Wittbrodt, J. & Stelzer, E. H. K. Reconstruction of zebrafish early embryonic development by scanned light sheet microscopy. Science 322, 1065–1069 (2008).
Keller, P. J. et al. Fast, high-contrast imaging of animal development with scanned light sheet-based structured-illumination microscopy. Nat. Methods 7, 637–642 (2010).
Silvestri, L., Bria, A., Sacconi, L., Iannello, G. & Pavone, F. S. Confocal light sheet microscopy: micron-scale neuroanatomy of the entire mouse brain. Opt. Express 20, 20582 (2012).
Baumgart, E. & Kubitscheck, U. Scanned light sheet microscopy with confocal slit detection. Opt. Express 20, 21805 (2012).
Fahrbach, F. O., Simon, P. & Rohrbach, A. Microscopy with self-reconstructing beams. Nat. Photonics 4, 780–785 (2010).
Planchon, T. A. et al. Rapid three-dimensional isotropic imaging of living cells using Bessel beam plane illumination. Nat. Methods 8, 417–423 (2011).
Vettenburg, T. et al. Light-sheet microscopy using an Airy beam. Nat. Methods 11, 541–544 (2014).
Chen, B.-C. et al. Lattice light-sheet microscopy: imaging molecules to embryos at high spatiotemporal resolution. Science 346, 1257998 (2014).
Chang, B. J. et al. Universal light-sheet generation with field synthesis. Nat. Methods 16, 235–238 (2019).
Ji, N. Adaptive optical fluorescence microscopy. Nat. Methods 14, 374–380 (2017).
Hörl, D. et al. BigStitcher: reconstructing high-resolution image datasets of cleared and expanded samples. Nat. Methods 16, 870–874 (2019).
Bria, A. & Iannello, G. TeraStitcher—a tool for fast automatic 3D-stitching of teravoxel-sized microscopy images. BMC Bioinform. 13, 316 (2012).
Kirst, C. et al. Mapping the fine-scale organization and plasticity of the brain vasculature. Cell 180, 780–795.e25 (2020).
Andreev, A. & Koo, D. E. S. Practical guide to storage of large amounts of microscopy data. Micros. Today 28, 42–45 (2020).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res 19, 1639–1645 (2009).
Kumar, A. et al. Dual-view plane illumination microscopy for rapid and spatially isotropic imaging. Nat. Protoc. 9, 2555–2573 (2014).
Tanaka, N. et al. Whole-tissue biopsy phenotyping of three-dimensional tumours reveals patterns of cancer heterogeneity. Nat. Biomed. Eng. 1, 796–806 (2017).
Costantini, I. et al. A versatile clearing agent for multi-modal brain imaging. Sci. Rep. 5, 9808 (2015).
Susaki, E. A. et al. Advanced CUBIC protocols for whole-brain and whole-body clearing and imaging. Nat. Protoc. 10, 1709–1727 (2015).
Matsumoto, K. et al. Advanced CUBIC tissue clearing for whole-organ cell profiling. Nat. Protoc. 14, 3506–3537 (2019).
Acknowledgements
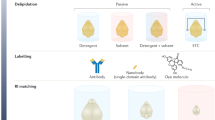
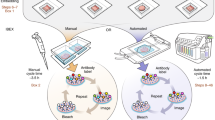
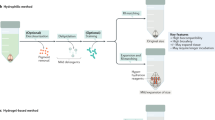
We thank J. Liu, A. Glaser, P. Ariel, D. Richardson, F. Helmchen, R. Power and the Huisken lab for their comments and discussion on the manuscript. Illustrations for Figs. 1 and 3 were done by M. Neufeld (madyrose.com). The Circos133 data visualization tool was used to generate Fig. 2. Blender (www.blender.org) was used for rendering Fig. 4. Brain tissue samples displayed in Fig. 3 were provided by L. Egolf, D. Kirschenbaum, F. Catto, P. Perin, K. Le Corf and A. Frick. We also thank the UW-Madison tissue clearing group and all those who provided inspiration, insight and samples: M. Taylor, R. Sulllivan, E. Dent, R. Taylor, K. Chan, R. Salamon and N. Iyer. We acknowledge funding by the Morgridge Institute for Research and the NIH (R01OD026219).
Author information
Authors and Affiliations
Contributions
K.W. and J.H. conceived of the manuscript and assembled the team. K.W., F.F.V. and D.P.S. constructed light-sheet microscopes designed for imaging cleared tissue and developed the manuscript content over years of troubleshooting tissue clearing and imaging. K.W. wrote the manuscript. K.W. and J.H. designed the figures. J.H., F.F.V. and D.P.S. provided feedback and extensive editing of the manuscript. F.F.V. provided images for Fig. 3.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks Hans-Ulrich Dodt and the other, anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Tainaka, K. et al. Annu. Rev. Cell Dev. Biol. 32, 713–714 (2016) (https://www.annualreviews.org/doi/abs/10.1146/annurev-cellbio-111315-125001) introduces the key strategies for clearing tissue and provides an overview of protocols developed up to 2016.
Gradinaru, V. et al. Annu. Rev. Biophys. 45, 355–376 (2018) (https://www.annualreviews.org/doi/10.1146/annurev-biophys-070317-032905) provides an introduction into the principles underlying hydrogel-based tissue processing techniques.
The Mesoscale Selective Plane Illumination Initiative (mesoSPIM) (http://www.mesospim.org) is an open-hardware project aimed at making instructions and software to set up and operate versatile light-sheet microscopes for large cleared samples more accessible.
Jonkman, J. et al. Nat. Protoc. 15, 1585–1611 (2020) (https://www.nature.com/articles/s41596-020-0313-9) is an excellent resource for optimizing fluorescent imaging experiments relevant to many imaging modalities.
The Confocal Microscopy List (http://confocal-microscopy-list.588098.n2.nabble.com/) has, since the early 1990s, been a go-to resource for microscopy and imaging-related questions.
Microform (https://forum.microlist.org/) was recently established by Jennifer Waters and Tally Lambert. Currently, most questions are on hardware, fluorophores and other practical aspects of microscopy.
TeraStitcher (https://abria.github.io/TeraStitcher/) is a stitching tool for large microscopy datasets developed by Alessandro Bria, Giulio Iannello and Roberto Valenti.
Napari (https://github.com/napari/napari) is a rapidly evolving multidimensional image viewer especially well suited for microscopy datasets written in Python. The development is spearheaded by the Chan-Zuckerberg Biohub.
BigStitcher (https://imagej.net/BigStitcher) is a stitching toolbox integrated into the ImageJ/Fiji ecosystem capable of stitching and fusing large multiview lightsheet datasets. The BigStitcher software was developed in the laboratory of Stephan Preibisch.
Supplementary information
Rights and permissions
About this article
Cite this article
Weiss, K.R., Voigt, F.F., Shepherd, D.P. et al. Tutorial: practical considerations for tissue clearing and imaging. Nat Protoc 16, 2732–2748 (2021). https://doi.org/10.1038/s41596-021-00502-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-021-00502-8
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.