Abstract
When used appropriately, a confocal fluorescence microscope is an excellent tool for making quantitative measurements in cells and tissues. The confocal microscope’s ability to block out-of-focus light and thereby perform optical sectioning through a specimen allows the researcher to quantify fluorescence with very high spatial precision. However, generating meaningful data using confocal microscopy requires careful planning and a thorough understanding of the technique. In this tutorial, the researcher is guided through all aspects of acquiring quantitative confocal microscopy images, including optimizing sample preparation for fixed and live cells, choosing the most suitable microscope for a given application and configuring the microscope parameters. Suggestions are offered for planning unbiased and rigorous confocal microscope experiments. Common pitfalls such as photobleaching and cross-talk are addressed, as well as several troubling instrumentation problems that may prevent the acquisition of quantitative data. Finally, guidelines for analyzing and presenting confocal images in a way that maintains the quantitative nature of the data are presented, and statistical analysis is discussed. A visual summary of this tutorial is available as a poster (https://doi.org/10.1038/s41596-020-0307-7).
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout










Similar content being viewed by others
References
Pawley, J. The 39 steps: a cautionary tale of quantitative 3-D fluorescence microscopy. Biotechniques 28, 884–886 (2000). 888.
North, A. J. Seeing is believing? A beginners’ guide to practical pitfalls in image acquisition. J. Cell Biol. 172, 9–18 (2006).
Hell, S., Reiner, G., Cremer, C. & Stelzer, E. H. K. Aberrations in confocal fluorescence microscopy induced by mismatches in refractive index. J. Microsc. 169, 391–405 (1993).
Richardson, D. S. & Lichtman, J. W. Clarifying tissue clearing. Cell 162, 246–257 (2015).
Allan, V. J. Basic immunofluorescence. in Protein Localization by Fluorescence Microscopy: A Practical Approach (ed. Allan, V. J.) 1–26 (Oxford University Press, 1999).
McDonald, K. L., Morphew, M., Verkade, P. & Muller-Reichert, T. Recent advances in high-pressure freezing: equipment- and specimen-loading methods. Methods Mol. Biol. 369, 143–173 (2007).
North, A. J., Chidgey, M. A., Clarke, J. P., Bardsley, W. G. & Garrod, D. R. Distinct desmocollin isoforms occur in the same desmosomes and show reciprocally graded distributions in bovine nasal epidermis. Proc. Natl Acad. Sci. USA 93, 7701–7705 (1996).
Burry, R. W. Immunocytochemistry: A Practical Guide for Biomedical Research (Springer, 2010).
Park, Y. G. et al. Protection of tissue physicochemical properties using polyfunctional crosslinkers. Nat. Biotechnol. 37, 73–83 (2019).
Richter, K. N. et al. Glyoxal as an alternative fixative to formaldehyde in immunostaining and super-resolution microscopy. EMBO J. 37, 139–159 (2018).
Melan, M. A. & Sluder, G. Redistribution and differential extraction of soluble proteins in permeabilized cultured cells. Implications for immunofluorescence microscopy. J. Cell Sci. 101(Pt 4), 731–743 (1992).
Jamur, M. C. & Oliver, C. Permeabilization of cell membranes. Methods Mol. Biol. 588, 63–66 (2010).
Yan, Q. & Bruchez, M. P. Advances in chemical labeling of proteins in living cells. Cell Tissue Res. 360, 179–194 (2015).
Ries, J., Kaplan, C., Platonova, E., Eghlidi, H. & Ewers, H. A simple, versatile method for GFP-based super-resolution microscopy via nanobodies. Nat. Methods 9, 582–584 (2012).
Dolman, N. J., Kilgore, J. A. & Davidson, M. W. A review of reagents for fluorescence microscopy of cellular compartments and structures, part I: BacMam labeling and reagents for vesicular structures. Curr. Protoc. Cytom. 65, 12.30.1–12.30.27 (2013).
Kilgore, J. A., Dolman, N. J. & Davidson, M. W. A review of reagents for fluorescence microscopy of cellular compartments and structures, Part II: reagents for non-vesicular organelles. Curr. Protoc. Cytom. 66, 12.31.1–12.31.24 (2013).
Bordeaux, J. et al. Antibody validation. Biotechniques 48, 197–209 (2010).
Pauly, D. & Hanack, K. How to avoid pitfalls in antibody use. F1000Res 4, 691 (2015).
Stadler, C. et al. Systematic validation of antibody binding and protein subcellular localization using siRNA and confocal microscopy. J. Proteomics 75, 2236–2251 (2012).
Stack, R. F. et al. Quality assurance testing for modern optical imaging systems. Microsc. Microanal. 17, 598–606 (2011).
Cordes, T., Maiser, A., Steinhauer, C., Schermelleh, L. & Tinnefeld, P. Mechanisms and advancement of antifading agents for fluorescence microscopy and single-molecule spectroscopy. Phys. Chem. Chem. Phys. 13, 6699–6709 (2011).
Piterburg, M., Panet, H. & Weiss, A. Photoconversion of DAPI following UV or violet excitation can cause DAPI to fluoresce with blue or cyan excitation. J. Microsc. 246, 89–95 (2012).
Frigault, M. M., Lacoste, J., Swift, J. L. & Brown, C. M. Live-cell microscopy—tips and tools. J. Cell Sci. 122, 753–767 (2009).
Ettinger, A. & Wittmann, T. Fluorescence live cell imaging. Methods Cell Biol. 123, 77–94 (2014).
Lambert, T. J. FPbase: a community-editable fluorescent protein database. Nat. Methods 16, 277–278 (2019).
Ai, H. W., Baird, M. A., Shen, Y., Davidson, M. W. & Campbell, R. E. Engineering and characterizing monomeric fluorescent proteins for live-cell imaging applications. Nat. Protoc. 9, 910–928 (2014).
Rodriguez, E. A. et al. The growing and glowing toolbox of fluorescent and photoactive proteins. Trends Biochem. Sci. 42, 111–129 (2017).
Cranfill, P. J. et al. Quantitative assessment of fluorescent proteins. Nat. Methods 13, 557–562 (2016).
Bottanelli, F. et al. Two-colour live-cell nanoscale imaging of intracellular targets. Nat. Commun. 7, 10778 (2016).
Erdmann, R. S. et al. Labeling strategies matter for super-resolution microscopy: a comparison between HaloTags and SNAP-tags. Cell Chem. Biol. 26, 584–592.e6 (2019).
Wang, L. et al. A general strategy to develop cell permeable and fluorogenic probes for multicolour nanoscopy. Nat. Chem. 12, 165–172 (2019).
Grimm, J. B., Brown, T. A., English, B. P., Lionnet, T. & Lavis, L. D. Synthesis of Janelia Fluor HaloTag and SNAP-Tag ligands and their use in cellular imaging experiments. Methods Mol. Biol. 1663, 179–188 (2017).
Ferrando-May, E. et al. Advanced light microscopy core facilities: balancing service, science and career. Microsc. Res. Tech. 79, 463–479 (2016).
Kiepas, A., Voorand, E., Mubaid, F., Siegel, P. M. & Brown, C. M. Optimizing live-cell fluorescence imaging conditions to minimize phototoxicity. J. Cell Sci. 133, jcs242834 (2020).
Laissue, P. P., Alghamdi, R. A., Tomancak, P., Reynaud, E. G. & Shroff, H. Assessing phototoxicity in live fluorescence imaging. Nat. Methods 14, 657 (2017).
Jonkman, J. E., Swoger, J., Kress, H., Rohrbach, A. & Stelzer, E. H. Resolution in optical microscopy. Methods Enzymol. 360, 416–446 (2003).
Jacques, S. L. Optical properties of biological tissues: a review. Phys. Med. Biol. 58, R37–R61 (2013).
Bolte, S. & Cordelieres, F. P. A guided tour into subcellular colocalization analysis in light microscopy. J. Microsc. 224, 213–232 (2006).
Dunn, K. W., Kamocka, M. M. & McDonald, J. H. A practical guide to evaluating colocalization in biological microscopy. Am. J. Physiol. Cell Physiol. 300, C723–C742 (2011).
Wallace, W., Schaefer, L. H. & Swedlow, J. R. A workingperson’s guide to deconvolution in light microscopy. Biotechniques 31, 1076–1078 (2001). 1080, 1082 passim.
Jonkman, J. & Brown, C. M. Any way you slice it—a comparison of confocal microscopy techniques. J. Biomol. Tech. 26, 54–65 (2015).
Korobchevskaya, K., Lagerholm, B. C., Colin-York, H. & Fritzsche, M. Exploring the potential of Airyscan microscopy for live cell imaging. Photonics 4, 41 (2017).
Zipfel, W. R., Williams, R. M. & Webb, W. W. Nonlinear magic: multiphoton microscopy in the biosciences. Nat. Biotechnol. 21, 1369–1377 (2003).
Axelrod, D. Total internal reflection fluorescence microscopy in cell biology. Traffic 2, 764–774 (2001).
Power, R. M. & Huisken, J. A guide to light-sheet fluorescence microscopy for multiscale imaging. Nat. Methods 14, 360–373 (2017).
Strobl, F., Schmitz, A. & Stelzer, E. H. K. Improving your four-dimensional image: traveling through a decade of light-sheet-based fluorescence microscopy research. Nat. Protoc. 12, 1103–1109 (2017).
Sigal, Y. M., Zhou, R. & Zhuang, X. Visualizing and discovering cellular structures with super-resolution microscopy. Science 361, 880–887 (2018).
Wu, Y. & Shroff, H. Faster, sharper, and deeper: structured illumination microscopy for biological imaging. Nat. Methods 15, 1011–1019 (2018).
Ishikawa-Ankerhold, H. C., Ankerhold, R. & Drummen, G. P. Advanced fluorescence microscopy techniques—FRAP, FLIP, FLAP, FRET and FLIM. Molecules 17, 4047–4132 (2012).
Lippincott-Schwartz, J. & Patterson, G. H. Development and use of fluorescent protein markers in living cells. Science 300, 87–91 (2003).
Lippincott-Schwartz, J., Altan-Bonnet, N. & Patterson, G. H. Photobleaching and photoactivation: following protein dynamics in living cells. Nat. Cell Biol. Suppl, S7–S14 (2003).
Elson, E. L. Fluorescence correlation spectroscopy: past, present, future. Biophys. J. 101, 2855–2870 (2011).
Kim, S. A., Heinze, K. G. & Schwille, P. Fluorescence correlation spectroscopy in living cells. Nat. Methods 4, 963–973 (2007).
Brown, C. M. et al. Raster image correlation spectroscopy (RICS) for measuring fast protein dynamics and concentrations with a commercial laser scanning confocal microscope. J. Microsc. 229, 78–91 (2008).
Sprague, B. L. & McNally, J. G. FRAP analysis of binding: proper and fitting. Trends Cell Biol. 15, 84–91 (2005).
Padilla-Parra, S. & Tramier, M. FRET microscopy in the living cell: different approaches, strengths and weaknesses. Bioessays 34, 369–376 (2012).
Broussard, J. A., Rappaz, B., Webb, D. J. & Brown, C. M. Fluorescence resonance energy transfer microscopy as demonstrated by measuring the activation of the serine/threonine kinase Akt. Nat. Protoc. 8, 265–281 (2013).
Bacia, K. & Schwille, P. Practical guidelines for dual-color fluorescence cross-correlation spectroscopy. Nat. Protoc. 2, 2842–2856 (2007).
Krieger, J. W. et al. Imaging fluorescence (cross-) correlation spectroscopy in live cells and organisms. Nat. Protoc. 10, 1948–1974 (2015).
Soderberg, O. et al. Direct observation of individual endogenous protein complexes in situ by proximity ligation. Nat. Methods 3, 995–1000 (2006).
Holman, L., Head, M. L., Lanfear, R. & Jennions, M. D. Evidence of experimental bias in the life sciences: why we need blind data recording. PLoS Biol. 13, e1002190 (2015).
Kaptchuk, T. J. The double-blind, randomized, placebo-controlled trial: gold standard or golden calf? J. Clin. Epidemiol. 54, 541–549 (2001).
Bankhead, P. et al. QuPath: open source software for digital pathology image analysis. Sci. Rep. 7, 16878 (2017).
Komura, D. & Ishikawa, S. Machine learning methods for histopathological image analysis. Comput. Struct. Biotechnol. J. 16, 34–42 (2018).
Robertson, S., Azizpour, H., Smith, K. & Hartman, J. Digital image analysis in breast pathology—from image processing techniques to artificial intelligence. Transl. Res. 194, 19–35 (2018).
Howard, V. & Reed, M. G. Unbiased Stereology: Three-Dimensional Measurement in Microscopy (Springer, 1998).
Kipanyula, M. J. & Sife, A. S. Global trends in application of stereology as a quantitative tool in biomedical research. Biomed. Res. Int. 2018, 1825697 (2018).
Jonkman, J. E. et al. An introduction to the wound healing assay using live-cell microscopy. Cell Adh. Migr. 8, 440–451 (2014).
Zimmermann, T., Marrison, J., Hogg, K. & O’Toole, P. Clearing up the signal: spectral imaging and linear unmixing in fluorescence microscopy. Methods Mol. Biol. 1075, 129–148 (2014).
Jonkman, J., Brown, C. M. & Cole, R. W. Quantitative confocal microscopy: beyond a pretty picture. Methods Cell Biol. 123, 113–134 (2014).
Oreopoulos, J., Berman, R. & Browne, M. Chapter 9—Spinning-disk confocal microscopy: present technology and future trends. in Methods in Cell Biology: Quantitative Imaging in Cell Biology Vol. 123 (eds Waters, J. C. & Wittman, T.) 153–175 (Academic Press, 2014).
Model, M. A. & Blank, J. L. Concentrated dyes as a source of two-dimensional fluorescent field for characterization of a confocal microscope. J. Microsc. 229, 12–16 (2008).
International Organization for Standardization. Microscopes—Confocal microscopes—Optical data of fluorescence confocal microscopes for biological imaging. ISO Standard No. 21073:2019 (2019).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Arena, E. T. et al. Quantitating the cell: turning images into numbers with ImageJ. Wiley Interdiscip. Rev. Dev. Biol. 6, e260 (2017).
Linkert, M. et al. Metadata matters: access to image data in the real world. J. Cell Biol. 189, 777–782 (2010).
Arganda-Carreras, I. et al. Trainable Weka Segmentation: a machine learning tool for microscopy pixel classification. Bioinformatics 33, 2424–2426 (2017).
Tinevez, J. Y. et al. TrackMate: an open and extensible platform for single-particle tracking. Methods 115, 80–90 (2017).
McQuin, C. et al. CellProfiler 3.0: next-generation image processing for biology. PLoS Biol. 16, e2005970 (2018).
Bray, M. A. et al. Cell Painting, a high-content image-based assay for morphological profiling using multiplexed fluorescent dyes. Nat. Protoc. 11, 1757–1774 (2016).
Weigert, M. et al. Content-aware image restoration: pushing the limits of fluorescence microscopy. Nat. Methods 15, 1090–1097 (2018).
Belthangady, C. & Royer, L. A. Applications, promises, and pitfalls of deep learning for fluorescence image reconstruction. Nat. Methods 16, 1215–1225 (2019).
Royer, L. A. et al. ClearVolume: open-source live 3D visualization for light-sheet microscopy. Nat. Methods 12, 480–481 (2015).
Cromey, D. W. Avoiding twisted pixels: ethical guidelines for the appropriate use and manipulation of scientific digital images. Sci. Eng. Ethics 16, 639–667 (2010).
Cromey, D. W. Digital images are data: and should be treated as such. Methods Mol. Biol. 931, 1–27 (2013).
Goodwin, P. C. Quantitative deconvolution microscopy. Methods Cell Biol. 123, 177–192 (2014).
Allan, C. et al. OMERO: flexible, model-driven data management for experimental biology. Nat. Methods 9, 245–253 (2012).
Ellenberg, J. et al. A call for public archives for biological image data. Nat. Methods 15, 849–854 (2018).
Fay, D. S. & Gerow, K. A biologist’s guide to statistical thinking and analysis. WormBook Jul 9, 1–54 (2013).
Vaux, D. L. Basic statistics in cell biology. Annu. Rev. Cell Dev. Biol. 30, 23–37 (2014).
Lacoste, J., Young, K. & Brown, C. M. Live-cell migration and adhesion turnover assays. Methods Mol. Biol. 931, 61–84 (2013).
Krzywinski, M. & Altman, N. Visualizing samples with box plots. Nat. Methods 11, 119–120 (2014).
Krzywinski, M. & Altman, N. Significance, P values and t-tests. Nat. Methods 10, 1041–1042 (2013).
Wasserstein, R. L., Schirm, A. L. & Lazar, N. A. Moving to a world beyond “p < 0.05”. Am. Stat. 73, 1–19 (2019).
Hibbs, A. R., MacDonald, G. & Garsha, K. Chapter 36: Practical confocal microscopy. in Handbook of Biological Confocal Microscopy 3rd edn (ed. Pawley, J. B.) (Springer, 2006).
Wang, H., Lacoche, S., Huang, L., Xue, B. & Muthuswamy, S. K. Rotational motion during three-dimensional morphogenesis of mammary epithelial acini relates to laminin matrix assembly. Proc. Natl Acad. Sci. USA 110, 163–168 (2013).
Cole, R. W., Jinadasa, T. & Brown, C. M. Measuring and interpreting point spread functions to determine confocal microscope resolution and ensure quality control. Nat. Protoc. 6, 1929–1941 (2011).
Acknowledgements
J.J. thanks the AOMF staff for helpful discussions, Courtney McIntosh for the images in Supplementary Fig. 1, and the Princess Margaret Foundation for ongoing financial support of the AOMF. G.D.W. thanks A*STAR and the National Research Foundation’s Shared Infrastructure Support Grant for continued support of the A*STAR Microscopy Platform and John Common for samples (Box 2). C.M.B. acknowledges Alex Kiepas (McGill University), who collected the adhesion dynamics data for the statistics section of the paper including Fig. 10, and the ABIF for general support and access to the Diskovery spinning disk TIRF microscope for collecting the adhesion dynamics data. K.I.A. thanks the Francis Crick Institute for their CALM support, and facility colleagues for helpful discussion. A.J.N. thanks the Rockefeller University for its continued support of the Frits and Rita Markus Bio-Imaging Resource Center (BIRC), the Sohn Conference Foundation for funding the Leica SP8 confocal microscope used to generate Figs. 3 and 4 and the facility staff and users for stimulating discussions.
Author information
Authors and Affiliations
Contributions
No section of this manuscript was untouched by all five authors. J.J. drafted the outline, assembled the team and wrote the Introduction, ‘Removing bias’, ‘Troubleshooting instrumentation issues’ and ‘Analyzing and presenting quantitative images’. He also generated Figs. 1, 2, 6, 7, 8 and 9; Table 1; Box 3; and Supplementary Figs. 1 and 2 and contributed to general editing. C.M.B. analyzed adhesion dynamics data, generated the statistics figure (Fig. 10), wrote the Statistics section of the manuscript and contributed to ‘General considerations for preparing samples for quantitative fluorescence microscopy’, ‘Preparing fixed cells and tissues’ and ‘Preparing live cells’ and significantly to general editing of the manuscript. G.D.W. worked on ‘Choosing the right microscope’ and ‘Setting up the microscope’, performed general editing of the manuscript and generated Fig. 5, the images for Box 2 and Supplementary Video 1. K.I.A. worked on ‘Choosing the right microscope’ and ‘Planning your experiment’ and performed general editing of the manuscript. A.J.N. worked on ‘General considerations for preparing samples for quantitative fluorescence microscopy’ and ‘Setting up the microscope’, contributed extensively to general editing and generated Figs. 3 and 4 and Boxes 1 and 2.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks Gary Laevsky, Timo Zimmermann and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Guidance for Quantitative Confocal Microscopy (https://doi.org/10.1038/s41596-020-0307-7): This poster is a visual summary of this paper.
Molecular Expressions Optical Microscopy Primer (https://micro.magnet.fsu.edu/primer/): Developed by Michael W. Davidson and his team at The Florida State University, this website hosts a vast amount of knowledge on all aspects of optical microscopy from the physics of light and color and the anatomy of a basic transmitted-light microscope to advanced techniques such as FRET and TIRF.
iBiology Microscopy Series (https://www.ibiology.org/online-biology-courses/microscopy-series/): Directed by Ron Vale, Nico Stuurman and Kurt Thorn, this 72-video series (and growing!) starts with the basics of optical microscopy and concludes with some of the latest techniques, such as super-resolution microscopy. Their Short Microscopy series of just 14 videos may be more manageable for some.
iBiology BioImage Analysis Course (https://www.ibiology.org/online-biology-courses/bioimage-analysis-course/): Created by Anne Carpenter (CellProfiler project lead) and Kevin Eliceiri (ImageJ project lead), this series focuses on general steps of image processing as well as introductions to CellProfiler and ImageJ specifically.
Microscope Vendor websites: Many microscope vendors have educational material on their websites. Some of them license sections from Molecular Expressions, but supplemented with their own microscopes’ unique attributes. Others have written their own material from scratch.
The Confocal Microscopy Listserv (https://lists.umn.edu/cgi-bin/wa?A0=confocalmicroscopy): With an active subscriber base of nearly 4,000 participants and an archive that goes back to 1991, the Confocal Listserv is a valuable resource for confocal and related questions. Registration (free) is required.
Microforum (https://forum.microlist.org/): This new web forum led by Jennifer Waters and Tally Lambert was established in 2019. Its focus is on hardware, acquisition and specimen-related aspects of scientific imaging, particularly (but not limited to) the theory and practical use of optical microscopes and detectors, fluorescent proteins and probes and specimen preparation.
Integrated supplementary information
Supplementary Fig. 1 How objective magnification affects the brightness of a CLSM image.
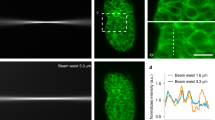
a, Schematic showing illumination and light collection for 40×/1.4 NA and 63×/1.4 NA objective lenses. The same objective is used for both illumination and collection in an epifluorescence geometry, but for clarity, the beampath has been unfolded. The incoming laser beam is focused to a small point in the specimen plane: the PSF. The shape of the PSF is determined by the NA of the objective. The intensity of the PSF depends on the size of the back aperture of the objective, which changes with the magnification of the lens. The incoming laser beam overfills the back aperture of the objective, with the result that the back aperture crops the outer rays of the laser beam, reducing the intensity of the incoming laser beam accordingly. For example, when switching from a 40×/1.4 NA objective to a 63×/1.4 NA objective, the aperture area is reduced by a factor of 402/632 = 0.40. Hence, we would expect a CLSM image through the 63× objective to be ~40% of the intensity compared to using the 40× objective, if the NA and quality of the lenses are identical. Does this mean that a 40×/1.4 NA objective is more sensitive than a 63×/1.4 NA objective because of the increased brightness through the 40× lens? No. The increased brightness for the 40× lens stems from the fact that more of the laser beam passes through the aperture and hits the sample: this change in intensity by means of a physical aperture is no more helpful than changing the laser power in the software. On the other hand, a higher NA is always helpful for confocal microscopy since the collection efficiency of the objective increases with the NA2 independent of magnification. b, CLSM image of a mouse kidney slide labeled with Alexa Fluor 488-WGA (Molecular Probes Prepared Slide #3) taken with a 40×/1.4 NA objective. c, CLSM image of the same field of view as b, taken with a 63×/1.4 NA objective using the same imaging parameters as b. The mean intensity of the image is reduced by ~36% in the 63× lens compared to the 40× lens. However, the power of the laser beam, which was measured using an oil immersion–compatible power meter (Thorlabs PM400 console with S170C sensor), was also reduced by 34%. This demonstrates that the dominant effect of changing magnification is to crop out a portion of the laser beam, for which one could easily compensate by increasing the laser power accordingly. The scale bar is 10 μm.
Supplementary Fig. 2 Focus drift.
Focus drift affects most microscope stands for 2–3 h after turning the microscope on, even when no incubators are used. A thin, fixed fluorescently labeled slide (Fluocells Prepared Slide #1, Molecular Probes) was placed on the stage of several confocal microscopes that had been turned off overnight (≥12 h). An optimized image was captured, and the focus position was recorded 10 min after turning power on to the instrument. For subsequent time points, the focus was adjusted manually so that the new image exactly matched the first image of the time series (the saturation LUT was helpful for evaluating when the images matched). There were no microscope incubators installed on these microscopes. a, Focus drift measured three times on the same Leica SP8 equipped with STED superresolution and a Super Galvo Z-stage demonstrates that the stand should be turned on 2–3 h before beginning confocal acquisition (particularly if STED is employed, as the acquisition times tend to be longer than regular confocal imaging). The Leica DMi8 microscope stand’s closed-loop focus feedback was enabled. b, Focus drift on a similar Leica SP8, both with and without the Super Galvo Z-stage, and both with and without the DMi8 closed-loop focus enabled. c, Focus drift on four other microscopes, demonstrating that the problem is not limited to any particular brand but is widespread.
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2.
Supplementary Video 1
DIC complements fluorescence for live-cell confocal microscopy. Live-cell confocal fluorescence (left) and DIC (right) timelapse imaging of keratinocytes expressing Keratin5-GFP. DIC can produce sharp images of cell and organelle boundaries without the need for labeling with additional fluorophores.
Rights and permissions
About this article
Cite this article
Jonkman, J., Brown, C.M., Wright, G.D. et al. Tutorial: guidance for quantitative confocal microscopy. Nat Protoc 15, 1585–1611 (2020). https://doi.org/10.1038/s41596-020-0313-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-020-0313-9
This article is cited by
-
Signal Processing and Artificial Intelligence for Dual-Detection Confocal Probes
International Journal of Precision Engineering and Manufacturing (2024)
-
A rapid and sensitive, multiplex, whole mount RNA fluorescence in situ hybridization and immunohistochemistry protocol
Plant Methods (2023)
-
Single-shot isotropic differential interference contrast microscopy
Nature Communications (2023)
-
Sensitivity of endogenous autofluorescence in HeLa cells to the application of external magnetic fields
Scientific Reports (2023)
-
Three-dimensional operando optical imaging of particle and electrolyte heterogeneities inside Li-ion batteries
Nature Nanotechnology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.