Abstract
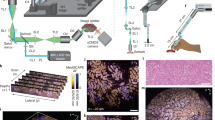
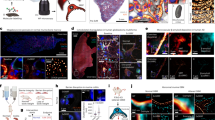
In pathology, microscopy is an important tool for the analysis of human tissues, both for the scientific study of disease states and for diagnosis. However, the microscopes commonly used in pathology are limited in resolution by diffraction. Recently, we discovered that it was possible, through a chemical process, to isotropically expand preserved cells and tissues by 4–5× in linear dimension. We call this process expansion microscopy (ExM). ExM enables nanoscale resolution imaging on conventional microscopes. Here we describe protocols for the simple and effective physical expansion of a variety of human tissues and clinical specimens, including paraffin-embedded, fresh frozen and chemically stained human tissues. These protocols require only inexpensive, commercially available reagents and hardware commonly found in a routine pathology laboratory. Our protocols are written for researchers and pathologists experienced in conventional fluorescence microscopy. The conventional protocol, expansion pathology, can be completed in ~1 d with immunostained tissue sections and 2 d with unstained specimens. We also include a new, fast variant, rapid expansion pathology, that can be performed on <5-µm-thick tissue sections, taking <4 h with immunostained tissue sections and <8 h with unstained specimens.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
Part of the primary data underlying the figures presented in this article can be found as examples in https://github.com/zhao-biophotonics/ExPath-reg; the rest of the primary data can be provided upon reasonable request from the corresponding authors.
References
Hell, S. W. Far-field optical nanoscopy. In 2010 23rd Annual Meeting of the IEEE Photonics Society, PHOTINICS 2010. 3–4 (IEEE, 2010) https://doi.org/10.1109/PHOTONICS.2010.5698725
Zhuang, X. Nano-imaging with STORM. Nat. Photonics 3, 365–367 (2009).
Huang, B., Bates, M. & Zhuang, X. Super-resolution fluorescence microscopy. Annu. Rev. Biochem. 78, 993–1016 (2009).
Betzig, E. Proposed method for molecular optical imaging. Opt. Lett. 20, 237–239 (1995).
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313, 1642–1645 (2006).
Chen, F., Tillberg, P. W. & Boyden, E. S. Expansion microscopy. Science 347, 543–548 (2015).
Tillberg, P. W. et al. Protein-retention expansion microscopy of cells and tissues labeled using standard fluorescent proteins and antibodies. Nat. Biotechnol. 34, 987–992 (2016).
Zhao, Y. et al. Nanoscale imaging of clinical specimens using pathology-optimized expansion microscopy. Nat. Biotechnol. 35, 757–764 (2017).
Chen, F. et al. Nanoscale imaging of RNA with expansion microscopy. Nat. Methods 13, 679–684 (2016).
Hausen, P. & Dreyer, C. The use of polyacrylamide as an embedding medium for immunohistochemical studies of embryonic tissues. Biotech. Histochem. 56, 287–293 (1981).
Wassie, A. T., Zhao, Y. & Boyden, E. S. Expansion microscopy: principles and uses in biological research. Nat. Methods 16, 33–41 (2019).
Chozinski, T. J. et al. Expansion microscopy with conventional antibodies and fluorescent proteins. Nat. Methods 13, 1–7 (2016).
Ku, T. et al. Multiplexed and scalable super-resolution imaging of three-dimensional protein localization in size-adjustable tissues. Nat. Biotechnol. 34, 973–981 (2016).
Chang, J.-B. et al. Iterative expansion microscopy. Nat. Methods 14, 593–599 (2017).
Asano, S. M. et al. Expansion microscopy: protocols for imaging proteins and RNA in cells and tissues. Curr. Protoc. Cell Biol. 80, e56 (2018).
Karagiannis, E. & Boyden, E. Expansion microscopy: development and neuroscience applications. Curr. Opin. Neurobiol. 50, 56–63 (2018).
Gao, R., Asano, S. M. & Boyden, E. S. Q&A: expansion microscopy. BMC Biol. 15, 50 (2017).
Hell, S. W. & Wichmann, J. Breaking the diffraction resolution limit by stimulated emission: stimulated-emission-depletion fluorescence microscopy. Opt. Lett. 19, 780–782 (1994).
Klar, T. A., Jakobs, S., Dyba, M., Egner, A. & Hell, S. W. Fluorescence microscopy with diffraction resolution barrier broken by stimulated emission. Proc. Natl Acad. Sci. USA 97, 8206–8210 (2000).
Gustafsson, M. G. Surpassing the lateral resolution limit by a factor of two using structured illumination microscopy. J. Microsc. 198, 82–87 (2000).
Heintzmann, R. & Cremer, C. G. Laterally modulated excitation microscopy: improvement of resolution by using a diffraction grating. In SPIE Proceedings Vol. 3568: Optical Biopsies and Microscopic Techniques III (eds Bigio, I. J., Schneckenburger, H., Slavik, J., Svanberg, K. & Viallet, P. M.) 185–196 (International Society for Optics and Photonics, 1999) https://spie.org/Publications/Proceedings/Paper/10.1117/12.336833
Rust, M. J., Bates, M. & Zhuang, X. W. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat. Methods 3, 793–795 (2006).
Jungmann, R. et al. Multiplexed 3D cellular super-resolution imaging with DNA-PAINT and Exchange-PAINT. Nat. Methods 11, 313–318 (2014).
Ertürk, A. et al. Three-dimensional imaging of solvent-cleared organs using 3DISCO. Nat. Protoc. 7, 1983–1995 (2012).
Renier, N. et al. iDISCO: a simple, rapid method to immunolabel large tissue samples for volume imaging. Cell 159, 896–910 (2014).
Dodt, H.-U. et al. Ultramicroscopy: three-dimensional visualization of neuronal networks in the whole mouse brain. Nat. Methods 4, 331–336 (2007).
Ke, M.-T., Fujimoto, S. & Imai, T. SeeDB: a simple and morphology-preserving optical clearing agent for neuronal circuit reconstruction. Nat. Neurosci. 16, 1154–1161 (2013).
Hama, H. et al. Scale: a chemical approach for fluorescence imaging and reconstruction of transparent mouse brain. Nat. Neurosci. 14, 1481–1488 (2011).
Susaki, E. A. et al. Whole-brain imaging with single-cell resolution using chemical cocktails and computational analysis. Cell 157, 726–739 (2014).
Chung, K. et al. Structural and molecular interrogation of intact biological systems. Nature 497, 332–337 (2013).
Yang, B. et al. Single-cell phenotyping within transparent intact tissue through whole-body clearing. Cell 158, 945–958 (2014).
Gao, R. et al. Cortical column and whole-brain imaging with molecular contrast and nanoscale resolution. Science 363, eaau8302 (2019).
Halpern, A. R., Alas, G. C. M., Chozinski, T. J., Paredez, A. R. & Vaughan, J. C. Hybrid structured illumination expansion microscopy reveals microbial cytoskeleton organization. ACS Nano 11, 12677–12686 (2017).
Wang, Y. et al. Combined expansion microscopy with structured illumination microscopy for analyzing protein complexes. Nat. Protoc. 13, 1869–1895 (2018).
Cahoon, C. K. et al. Superresolution expansion microscopy reveals the three-dimensional organization of the Drosophila synaptonemal complex. Proc. Natl Acad. Sci. USA 114, E6857–E6866 (2017).
Tong, Z. et al. Ex-STORM: expansion single molecule super-resolution microscopy. Preprint at https://www.biorxiv.org/content/10.1101/049403v2 (2016).
Gao, M. et al. Expansion stimulated emission depletion microscopy (ExSTED). ACS Nano 12, 4178–4185 (2018).
Gambarotto, D. et al. Imaging cellular ultrastructures using expansion microscopy (U-ExM). Nat. Methods 16, 71–74 (2019).
Truckenbrodt, S., Sommer, C., Rizzoli, S. O. & Danzl, J. G. A practical guide to optimization in X10 expansion microscopy. Nat. Protoc. 14, 832–863 (2019).
Acknowledgements
Human lymph node specimens were from the pathology archives of the Harvard University Center for AIDS Research, obtained under IRB protocol #2010P000632 to B.D.W. For funding, E.S.B. acknowledges L. Yang, Schmidt Futures, the MIT Media Lab, the Chan Zuckerberg Initiative, NIH U01MH114819, NIH 1U19MH114821, the Ludwig Foundation, NIH 1R01NS102727, John Doerr, NIH 1R01EB024261, the Open Philanthropy project, the HHMI-Simons Faculty Scholars Program, the US Army Research Laboratory and the US Army Research Office under contract/grant number W911NF1510548, NIH 1R01MH110932 and NIH 1RM1HG008525. O.B. acknowledges support from the Ludwig Center at Harvard and from Harvard Catalyst (the Harvard Clinical and Translational Science Center (National Center for Research Resources and the National Center for Advancing Translational Sciences, National Institutes of Health Award UL1 TR001102)). Y.Z. acknowledges support from Carnegie Mellon University and NIH Director’s New Innovator Award (DP2 OD025926-01).
Author information
Authors and Affiliations
Contributions
Y.Z., O.B. and E.S.B. wrote the manuscript. Y.Z., O.B. and F.F. conducted the experiments. Y.Z., O.B., F.F. and E.S.B. analyzed the data. M.C. and S.W. conducted STED imaging for validation of rExPath. J.D. filmed and photographed the ExPath process. G.H.M., N.L.L. and B.D.W. provided de-identified human clinical specimens and helped with the experiments. Y.Z. and E.S.B. oversaw the research.
Corresponding authors
Ethics declarations
Competing interests
The authors have filed and obtained patent protection on a subset of the technologies here described (US provisional application no. 62/299,754, 62/463,265 and 62/463,251). E.S.B. helped cofound a company to help disseminate ExM to the community. O.B. is the Co-Founder and CEO of QPathology LLC, Boston, MA.
Additional information
Peer review information Nature Protocols thanks Sven Truckenbrodt and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Zhao, Y. et al. Nat. Biotechnol. 35, 757–764 (2017): https://doi.org/10.1038/nbt.3892
Gao, R. et al. Science 363, eaau8302 (2019): https://doi.org/10.1126/science.aau8302
Wassie, A. T. et al. Nat. Methods 16, 33–41 (2019): https://doi.org/10.1038/s41592-018-0219-4
Supplementary information
Rights and permissions
About this article
Cite this article
Bucur, O., Fu, F., Calderon, M. et al. Nanoscale imaging of clinical specimens using conventional and rapid-expansion pathology. Nat Protoc 15, 1649–1672 (2020). https://doi.org/10.1038/s41596-020-0300-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-020-0300-1
This article is cited by
-
Microglia-mediated demyelination protects against CD8+ T cell-driven axon degeneration in mice carrying PLP defects
Nature Communications (2023)
-
Magnify is a universal molecular anchoring strategy for expansion microscopy
Nature Biotechnology (2023)
-
Nanoscale fluorescence imaging of biological ultrastructure via molecular anchoring and physical expansion
Nano Convergence (2022)
-
Fluorescent labeling of abundant reactive entities (FLARE) for cleared-tissue and super-resolution microscopy
Nature Protocols (2022)
-
Super-resolved fluorescence imaging of peripheral nerve
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.