Abstract
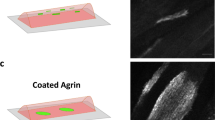
This protocol describes the design, fabrication and use of a 3D physiological and pathophysiological motor unit model consisting of motor neurons coupled to skeletal muscles interacting via the neuromuscular junction (NMJ) within a microfluidic device. This model facilitates imaging and quantitative functional assessment. The ‘NMJ chip’ enables real-time, live imaging of axonal outgrowth, NMJ formation and muscle maturation, as well as synchronization of motor neuron activity and muscle contraction under optogenetic control for the study of normal physiological events. The proposed protocol takes ~2–3 months to be implemented. Pathological behaviors associated with various neuromuscular diseases, such as regression of motor neuron axons, motor neuron death, and muscle degradation and atrophy can also be recapitulated in this system. Disease models can be created by the use of patient-derived induced pluripotent stem cells to generate both the motor neurons and skeletal muscle cells used. This is demonstrated by the use of cells from a patient with sporadic amyotrophic lateral sclerosis but can be applied more generally to models of neuromuscular disease, such as spinal muscular atrophy, NMJ disorder and muscular dystrophy. Models such as this hold considerable potential for applications in precision medicine, drug screening and disease risk assessment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author on reasonable request.
References
Nageshwaran, S., Davies, L. M., Rafi, I. & Radunović, A. Motor neurone disease. BMJ 349, g4052 (2014).
Takahashi, K. et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131, 861–872 (2007).
Metzker, M. L. Sequencing technologies—the next generation. Nat. Rev. Genet. 11, 31–46 (2010).
Andersen, J. K. Oxidative stress in neurodegeneration: cause or consequence? Nat. Med. 10, S18–S25 (2004).
Sreedharan, J. et al. TDP-43 mutations in familial and sporadic amyotrophic lateral sclerosis. Science 319, 1668–1672 (2008).
Williams, K. L. et al. Pathophysiological insights into ALS with C9ORF72 expansions. J. Neurol. Neurosurg. Psychiatry 84, 931–935 (2013).
DeJesus-Hernandez, M. et al. Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS. Neuron 72, 245–256 (2011).
Giuliano, K. A., Haskins, J. R. & Taylor, D. L. Advances in high content screening for drug discovery. Assay. Drug Dev. Technol. 1, 565–577 (2003).
Egawa, N. et al. Drug screening for ALS using patient-specific induced pluripotent stem cells. Sci. Transl. Med. 4, 145ra104 (2012).
Fitzsimonds, R. M. & Poo, M.-M. Retrograde signaling in the development and modification of synapses. Physiol. Rev. 78, 143–170 (1998).
Wyatt, R. M. & Balice-Gordon, R. J. Activity-dependent elimination of neuromuscular synapses. J. Neurocytol. 32, 777–794 (2003).
Uzel, S. G. et al. Microfluidic device for the formation of optically excitable, three-dimensional, compartmentalized motor units. Sci. Adv. 2, e1501429 (2016).
Osaki, T., Uzel, S. G. M. & Kamm, R. D. Microphysiological 3D model of amyotrophic lateral sclerosis (ALS) from human iPS-derived muscle cells and optogenetic motor neurons. Sci. Adv. 4, eaat5847 (2018).
Park, J. W., Vahidi, B., Taylor, A. M., Rhee, S. W. & Jeon, N. L. Microfluidic culture platform for neuroscience research. Nat. Protoc. 1, 2128–2136 (2006).
Qin, D., Xia, Y. & Whitesides, G. M. Soft lithography for micro- and nanoscale patterning. Nat. Protoc. 5, 491–502 (2010).
Sakar, M. S. et al. Formation and optogenetic control of engineered 3D skeletal muscle bioactuators. Lab Chip 12, 4976–4985 (2012).
Lunn, M. R. & Wang, C. H. Spinal muscular atrophy. Lancet 371, 2120–2133 (2008).
Jaretzki, A. et al. Myasthenia gravis: recommendations for clinical research standards. Task Force of the Medical Scientific Advisory Board of the Myasthenia Gravis Foundation of America. Neurology 55, 16–23 (2000).
Vila, O. F. et al. Quantification of human neuromuscular function through optogenetics. Theranostics 9, 1232–1246 (2019).
Emery, A. E. H. The muscular dystrophies. Lancet 359, 687–695 (2002).
Bleicher, K. H., Böhm, H.-J., Müller, K. & Alanine, A. I. Hit and lead generation: beyond high-throughput screening. Nat. Rev. Drug Discov. 2, 369–378 (2003).
Mahmood, T. & Yang, P.-C. Western blot: technique, theory, and trouble shooting. N. Am. J. Med. Sci. 4, 429–434 (2012).
Nolan, T., Hands, R. E. & Bustin, S. A. Quantification of mRNA using real-time RT-PCR. Nat. Protoc. 1, 1559–1582 (2006).
Ran, F. A. et al. Genome engineering using the CRISPR–Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Brooks, B. R. Natural history of ALS: symptoms, strength, pulmonary function, and disability. Neurology 47, 71S–82S (1996).
Chen, Y. et al. The relationship between four GWAS-identified loci in Alzheimer’s disease and the risk of Parkinson’s disease, amyotrophic lateral sclerosis, and multiple system atrophy. Neurosci. Lett. 686, 205–210 (2018).
Laing, N. G. Genetics of neuromuscular disorders. Crit. Rev. Clin. Lab Sci. 49, 33–48 (2012).
Rudnick, N. D. et al. Distinct roles for motor neuron autophagy early and late in the SOD1G93A mouse model of ALS. Proc. Natl Acad. Sci. USA 114, E8294–E8303 (2017).
Lutz, C. Mouse models of ALS: past, present and future. Brain Res. 1693, 1–10 (2018).
Hsieh-Li, H. M. et al. A mouse model for spinal muscular atrophy. Nat. Genet. 24, 66–70 (2000).
Daub, A., Sharma, P. & Finkbeiner, S. High-content screening of primary neurons: ready for prime time. Curr. Opin. Neurobiol. 19, 537–543 (2009).
Ribas, J., Pawlikowska, J. & Rouwkema, J. Microphysiological systems: analysis of the current status, challenges and commercial future. Microphysiol. Syst. 2, 10 (2018).
Southam, K. A., King, A. E., Blizzard, C. A., McCormack, G. H. & Dickson, T. C. Microfluidic primary culture model of the lower motor neuron-neuromuscular junction circuit. J. Neurosci. Methods 218, 164–169 (2013).
Ionescu, A., Zahavi, E. E., Gradus, T., Ben-Yaakov, K. & Perlson, E. Compartmental microfluidic system for studying muscle-neuron communication and neuromuscular junction maintenance. Eur. J. Cell Biol. 95, 69–88 (2016).
Fujimori, K. et al. Modeling sporadic ALS in iPSC-derived motor neurons identifies a potential therapeutic agent. Nat. Med. 24, 1579–1589 (2018).
Smith, A. S. T., Long, C. J., Pirozzi, K. & Hickman, J. J. A functional system for high-content screening of neuromuscular junctions in vitro. Technology 1, 37–48 (2013).
Yizhar, O., Fenno, L. E., Davidson, T. J., Mogri, M. & Deisseroth, K. Optogenetics in neural systems. Neuron 71, 9–34 (2011).
Maffioletti, S. M. et al. Three-dimensional human iPSC-derived artificial skeletal muscles model muscular dystrophies and enable multilineage tissue engineering. Cell Rep. 23, 899–908 (2018).
Afshar Bakooshli, M. et al. A 3D culture model of innervated human skeletal muscle enables studies of the adult neuromuscular junction. eLife 8, e44530 (2019).
Wilson, M. H. & Deschenes, M. R. The neuromuscular junction: anatomical features and adaptations to various forms of increased, or decreased neuromuscular activity. Int. J. Neurosci. 115, 803–828 (2005).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Gilestro, G. F. Video tracking and analysis of sleep in Drosophila melanogaster. Nat. Protoc. 7, 995–1007 (2012).
Afshar, M. E. et al. A 96-well culture platform enables longitudinal analyses of engineered human skeletal muscle microtissue strength. Preprint at https://www.biorxiv.org/content/10.1101/562819v1 (2019).
Davie, J. T. et al. Dendritic patch-clamp recording. Nat. Protoc. 1, 1235–1247 (2006).
Spira, M. E. & Hai, A. Multi-electrode array technologies for neuroscience and cardiology. Nat. Nanotechnol. 8, 83–94 (2013).
Pearce, T. M., Wilson, J. A., Oakes, S. G., Chiu, S.-Y. & Williams, J. C. Integrated microelectrode array and microfluidics for temperature clamp of sensory neurons in culture. Lab Chip 5, 97–101 (2005).
Hochbaum, D. R. et al. All-optical electrophysiology in mammalian neurons using engineered microbial rhodopsins. Nat. Methods 11, 825–833 (2014).
Scanziani, M. & Häusser, M. Electrophysiology in the age of light. Nature 461, 930–939 (2009).
Jeong, J.-W. et al. Wireless optofluidic systems for programmable in vivo pharmacology and optogenetics. Cell 162, 662–674 (2015).
Psaltis, D., Quake, S. R. & Yang, C. Developing optofluidic technology through the fusion of microfluidics and optics. Nature 442, 381–386 (2006).
Albert-Smet, I. et al. Applications of light-sheet microscopy in microdevices. Front. Neuroanat. 13, 1 (2019).
Compston, A. & Coles, A. Multiple sclerosis. Lancet 372, 1502–1517 (2008).
Montanez-Sauri, S. I., Sung, K. E., Puccinelli, J. P., Pehlke, C. & Beebe, D. J. Automation of three-dimensional cell culture in arrayed microfluidic devices. J. Lab. Autom. 16, 171-185 (2011).
Chu, L. & Robinson, D. K. Industrial choices for protein production by large-scale cell culture. Curr. Opin. Biotechnol. 12, 180–187 (2001).
Miller, J. D. et al. Human iPSC-based modeling of late-onset disease via progerin-induced aging. Cell Stem Cell 13, 691–705 (2013).
Ziff, O. J. & Patani, R. Harnessing cellular aging in human stem cell models of amyotrophic lateral sclerosis. Aging Cell 18, e12862 (2019).
Hargus, G. et al. Origin-dependent neural cell identities in differentiated human iPSCs in vitro and after transplantation into the mouse brain. Cell Rep. 8, 1697–1703 (2014).
Mao, T. et al. Long-range neuronal circuits underlying the interaction between sensory and motor cortex. Neuron 72, 111–123 (2011).
Lake, M. et al. Microfluidic device design, fabrication, and testing protocols. Protocol Exchange https://doi.org/10.1038/protex.2015.069 (2015).
Acknowledgements
AAV-CAG-hChR2H134R-tdTomato was a gift from K. Svoboda at Howard Hughes Medical Institute (Addgene viral prep no. 28017-AAVrg; http://n2t.net/addgene:28017; RRID: Addgene_28017). pMD2.G was a gift from D. Trono at EPLF (Addgene plasmid no. 12259; http://n2t.net/addgene:12259; RRID: Addgene_12259). psPAX2 was a gift from D. Trono (Addgene plasmid no. 12260; http://n2t.net/addgene:12260; RRID: Addgene_12260). The pLEX307-EF1a-ChR2[H134R]-mCherry-Puro-WPRE plasmid can be obtained from Addgene (no. 125256). T.O. was supported by an overseas research fellowship (from the Japan Society for the Promotion of Science). T.O., S.G.M.U. and R.D.K. also acknowledge support from the National Science Foundation for a Science and Technology Center on Emergent Behaviors of Integrated Cellular Systems grant (CBET-0939511) and partially from the Cancer Center Support (core) grant P30-CA14051 from the NCI.
Author information
Authors and Affiliations
Contributions
T.O., S.G.M.U. and R.D.K. conceived and designed the study. T.O. conducted the experiment, and collected and analyzed the data. T.O. and S.G.M.U. designed the microfluidic devices for the human NMJ model. T.O. modified the microfluidic devices for the human NMJ model. T.O. prepared entire procedures and all of the figures. All authors wrote and revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
A part of this study was funded by Biogen Inc.
Additional information
Peer review information Nature Protocols thanks Eran Perlson, Francesco Saverio Tedesco and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Osaki, T., Uzel, S. G. M. & Kamm, R. D. Sci. Adv. 4, eaat5847 (2018): https://doi.org/10.1126/sciadv.aat5847.
Uzel, S. G. et al. Sci. Adv. 2, e1501429 (2016): https://doi.org/10.1126/sciadv.1501429.
Vila, O. F. et al. Theranostics 9, 1232–1246 (2019): https://doi.org/10.7150/thno.25735.
Supplementary information
Supplementary Data 1
Photomask-1 for photolithography (Step 5)
Supplementary Data 2
Photomask-2 for photolithography (Step 11)
Supplementary Data 3
Source code of TTL control for optogenetics in Arduino
Supplementary Data 4
Description: ImageJ macro for calculating pillar deflection by Method A (Steps 77B)
Supplementary Data 5
ImageJ macro for calculating pillar deflection by Method B (Steps 77B)
Rights and permissions
About this article
Cite this article
Osaki, T., Uzel, S.G.M. & Kamm, R.D. On-chip 3D neuromuscular model for drug screening and precision medicine in neuromuscular disease. Nat Protoc 15, 421–449 (2020). https://doi.org/10.1038/s41596-019-0248-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-019-0248-1
This article is cited by
-
Generation of functional posterior spinal motor neurons from hPSCs-derived human spinal cord neural progenitor cells
Cell Regeneration (2023)
-
A novel 3D bilayer hydrogel tri-culture system for studying functional motor units
Cell & Bioscience (2023)
-
Functional bioengineered models of the central nervous system
Nature Reviews Bioengineering (2023)
-
Genetics of amyotrophic lateral sclerosis: seeking therapeutic targets in the era of gene therapy
Journal of Human Genetics (2023)
-
Intramuscular delivery of neural crest stem cell spheroids enhances neuromuscular regeneration after denervation injury
Stem Cell Research & Therapy (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.