Abstract
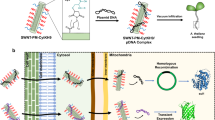
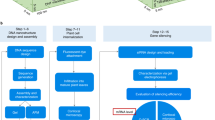
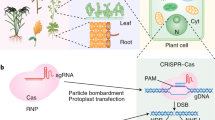
Exogenous biomolecule delivery into plants is difficult because the plant cell wall poses a dominant transport barrier, thereby limiting the efficiency of plant genetic engineering. Traditional DNA delivery methods for plants suffer from host-species limitations, low transformation efficiencies, tissue damage, or unavoidable and uncontrolled DNA integration into the host genome. We have demonstrated efficient plasmid DNA delivery into intact plants of several species with functionalized high-aspect-ratio carbon nanotube (CNT) nanoparticles (NPs), enabling efficient DNA delivery into a variety of non-model plant species (arugula, wheat, and cotton) and resulting in high protein expression levels without transgene integration. Herein, we provide a protocol that can be implemented by plant biologists and adapted to produce functionalized single-walled CNTs (SWNTs) with surface chemistries optimized for delivery of plasmid DNA in a plant species–independent manner. This protocol describes how to prepare, construct, and optimize polyethylenimine (PEI)-functionalized SWNTs and perform plasmid DNA loading. The authors also provide guidance on material characterization, gene expression evaluation, and storage conditions. The entire protocol, from the covalent functionalization of SWNTs to expression quantification, can be completed in 5 d.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All materials are available from commercial sources or can be derived using methods described in this study. All primary data underlying the figures reported in the article can be obtained from the corresponding author upon reasonable request.
References
Goswami, R., Dasgupta, P., Saha, S., Venkatapuram, P. & Nandi, S. Resource integration in smallholder farms for sustainable livelihoods in developing countries. Cogent Food Agric. 2, https://doi.org/10.1080/23311932.2016.1272151 (2016).
Altpeter, F. et al. Advancing crop transformation in the era of genome editing. Plant Cell 28, 1510–1520 (2016).
Himmel, M. E. et al. Biomass recalcitrance: engineering plants and enzymes for biofuels production. Science 315, 804–807 (2007).
Baltes, N. J., Gil-Humanes, J. & Voytas, D. F. Genome engineering and agriculture: opportunities and challenges. Prog. Mol. Biol. Transl. 149, 1–26 (2017).
Herreraestrella, L., Depicker, A., Vanmontagu, M. & Schell, J. Expression of chimaeric genes transferred into plant-cells using a Ti-plasmid-derived vector. Nature 303, 209–213 (1983).
Binns, A. N. Agrobacterium-mediated gene delivery and the biology of host range limitations. Physiol. Plant. 79, 135–139 (1990).
Klein, T. M., Wolf, E. D., Wu, R. & Sanford, J. C. High-velocity microprojectiles for delivering nucleic-acids into living cells. Nature 327, 70–73 (1987).
Cunningham, F. J., Goh, N. S., Demirer, G. S., Matos, J. L. & Landry, M. P. Nanoparticle-mediated delivery towards advancing plant genetic engineering. Trends Biotechnol. 36, 882–897 (2018).
Russell, J. A., Roy, M. K. & Sanford, J. C. Physical trauma and tungsten toxicity reduce the efficiency of biolistic transformation. Plant Physiol. 98, 1050–1056 (1992).
Liu, Q. et al. Carbon nanotubes as molecular transporters for walled plant cells. Nano Lett. 9, 1007–1010 (2009).
Serag, M. F. et al. Trafficking and subcellular localization of multiwalled carbon nanotubes in plant cells. ACS Nano 5, 493–499 (2011).
Chang, F.-P. et al. A simple plant gene delivery system using mesoporous silica nanoparticles as carriers. J. Mater. Chem. B 1, 5279–5287 (2013).
Zhang, H. et al. DNA nanostructures coordinate gene silencing in mature plants. Proc. Natl Acad. Sci. USA 116, 7543–7548 (2019).
Ng, K. K. et al. Intracellular delivery of proteins via fusion peptides in intact plants. PLoS One 11, e0154081 (2016).
Mitter, N. et al. Clay nanosheets for topical delivery of RNAi for sustained protection against plant viruses. Nat. Plants 3, 16207 (2017).
Giraldo, J. P. et al. Plant nanobionics approach to augment photosynthesis and biochemical sensing. Nat. Mater. 13, 400–408 (2014).
Demirer, G. S., Zhang, H., Goh, N. S., Chang, R. & Landry, M. P. Nanotubes effectively deliver siRNA to intact plant cells and protect siRNA against nuclease degradation. Preprint at bioRxiv, https://doi.org/10.1101/564427 (2019).
Demirer, G. S. et al. High aspect ratio nanomaterials enable delivery of functional genetic material without DNA integration in mature plants. Nat. Nanotechnol. 14, 456–464 (2019).
Kwak, S.-Y. et al. Nanosensor technology applied to living plant systems. Annu. Rev. Anal. Chem. 10, 113–140 (2017).
Wong, M. H. et al. Nitroaromatic detection and infrared communication from wild-type plants using plant nanobionics. Nat. Mater. 16, 264 (2017).
Del Bonis-O’Donnell, J. T. et al. Engineering molecular recognition with bio-mimetic polymers on single walled carbon nanotubes. J. Vis. Exp. 2017, e55030 (2017).
Kwak, S.-Y. et al. Chloroplast-selective gene delivery and expression in planta using chitosan-complexed single-walled carbon nanotube carriers. Nat. Nanotechnol. 14, 447–455 (2019).
Zhang, Y. et al. Efficient and transgene-free genome editing in wheat through transient expression of CRISPR/Cas9 DNA or RNA. Nat. Commun. 7, 12617 (2016).
Wang, P., Lombi, E., Zhao, F. J. & Kopittke, P. M. Nanotechnology: a new opportunity in plant sciences. Trends Plant Sci. 21, 699–712 (2016).
Kikkert, J. R., Vidal, J. R. & Reisch, B. I. in Transgenic Plants: Methods and Protocols 61-78 (Springer, 2005).
Liu, Y., Yang, H. & Sakanishi, A. Ultrasound: mechanical gene transfer into plant cells by sonoporation. Biotechnol. Adv. 24, 1–16 (2006).
Asad, S. & Arshad, M. Silicon carbide whisker-mediated plant transformation. in Properties and Applications of Silicon Carbide, 345-358 (InTech Open, 2011).
Lakshmanan, M., Kodama, Y., Yoshizumi, T., Sudesh, K. & Numata, K. Rapid and efficient gene delivery into plant cells using designed peptide carriers. Biomacromolecules 14, 10–16 (2013).
Yoo, S.-D., Cho, Y.-H. & Sheen, J. J. Np Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis. Nat. Protoc. 2, 1565-–1572 (2007).
Vashist, S. K. Comparison of 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide based strategies to crosslink antibodies on amine-functionalized platforms for immunodiagnostic applications. Diagnostics 2, (23–33 (2012).
Dobnik, D., Stebih, D., Blejec, A., Morisset, D. & Zel, J. Multiplex quantification of four DNA targets in one reaction with Bio-Rad droplet digital PCR system for GMO detection. Sci. Rep. 6, 35451 (2016).
Collier, R. et al. Accurate measurement of transgene copy number in crop plants using droplet digital PCR. Plant J. 90, 1014–1025 (2017).
Zhang, J. D., Ruschhaupt, M. & Biczok, R. ddCt method for qRT–PCR data analysis http://bioconductor.jp/packages/2.14/bioc/vignettes/ddCt/inst/doc/rtPCR.pdf (2010).
Zheng, N., Song, Z., Liu, Y., Yin, L. & Cheng, J. Gene delivery into isolated Arabidopsis thaliana protoplasts and intact leaves using cationic, α-helical polypeptide. Front. Chem. Sci. Eng. 11, 521–528 (2017).
Chugh, A. & Eudes, F. Study of uptake of cell penetrating peptides and their cargoes in permeabilized wheat immature embryos. FEBS J. 275, 2403–2414 (2008).
Chuah, J.-A., Yoshizumi, T., Kodama, Y. & Numata, K. J. Sr Gene introduction into the mitochondria of Arabidopsis thaliana via peptide-based carriers. Sci. Rep. 5, 7751 (2015).
Acknowledgements
We acknowledge support from a Burroughs Wellcome Fund Career Award at the Scientific Interface (CASI), a Stanley Fahn PDF Junior Faculty Grant under award no. PF-JFA-1760, a Beckman Foundation Young Investigator Award, a USDA AFRI award, a grant from the Gordon and Betty Moore Foundation, a USDA NIFA award, a USDA-BBT EAGER award, support from the Chan-Zuckerberg Foundation, and an FFAR New Innovator Award (to M.P.L.). G.S.D. was supported by a Schlumberger Foundation Faculty for the Future Fellowship. H.Z. acknowledges the support of the National Natural Science Foundation of China (21605153). We also acknowledge support from the UC Berkeley Molecular Imaging Center (supported by the Gordon and Betty Moore Foundation), the QB3 Shared Stem Cell Facility, and the Innovative Genomics Institute (IGI). We are grateful to the Staskawicz lab at UC Berkeley for the N. benthamiana seeds.
Author information
Authors and Affiliations
Contributions
G.S.D. and M.P.L. designed the experiments. G.S.D., H.Z., and N.S.G. performed the experiments. G.S.D. analyzed the data and created the figures. G.S.D., H.Z., N.S.G., and E.G.G. wrote the manuscript. All authors approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks Ardemis Boghossian, Mohammad Ramezani and other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key reference using this protocol
Demirer, G. S. et al. Nat. Nanotechnol. 14, 456–464 (2019): https://www.nature.com/articles/s41565-019-0382-5
Integrated supplementary information
Supplementary Figure 1 Characterization and comparison of heat reaction and EDC/NHS coupling for the synthesis of PEI-SWNTs.
a, XPS nitrogen peaks for the heat reacted PEI-SWNTs: amine peak at 399 eV and amide peak at 401 eV. b, XPS nitrogen peaks for the EDC/NHS reacted PEI-SWNTs. c, Zeta potential measurements of COOH-SWNTs, PEI-SWNTs via heat reaction, and DNA loaded PEI-SWNTs. ****P < 0.0001 and *P = 0.0191 in two-way ANOVA. N=3 and error bars are standard deviation. d, Zeta potential measurements of COOH-SWNTs, PEI-SWNTs via EDC/NHS coupling, and DNA loaded PEI-SWNTs. ****P < 0.0001and **P = 0.0025 in two-way ANOVA. N=3 and error bars are standard deviation. e, Representative confocal images at Day 3 and 10 of wild-type Nicotiana benthamiana leaves infiltrated with DNA-PEI-SWNTs prepared via heat reaction. e, Representative confocal images at Day 3 and 10 of wild-type Nicotiana benthamiana leaves infiltrated with DNA-PEI-SWNTs prepared via EDC/NHS coupling. All scale bars are 50 µm.
Supplementary information
Supplementary Information
Supplementary Figure 1
Rights and permissions
About this article
Cite this article
Demirer, G.S., Zhang, H., Goh, N.S. et al. Carbon nanotube–mediated DNA delivery without transgene integration in intact plants. Nat Protoc 14, 2954–2971 (2019). https://doi.org/10.1038/s41596-019-0208-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-019-0208-9
This article is cited by
-
Applications of CRISPR/Cas genome editing in economically important fruit crops: recent advances and future directions
Molecular Horticulture (2023)
-
Promoting genotype-independent plant transformation by manipulating developmental regulatory genes and/or using nanoparticles
Plant Cell Reports (2023)
-
Engineering plants with carbon nanotubes: a sustainable agriculture approach
Journal of Nanobiotechnology (2022)
-
A cationic lipid mediated CRISPR/Cas9 technique for the production of stable genome edited citrus plants
Plant Methods (2022)
-
Polymer-coated carbon nanotube hybrids with functional peptides for gene delivery into plant mitochondria
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.