Abstract
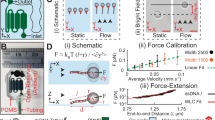
Measuring forces from the piconewton to millinewton range is of great importance for the study of living systems from a biophysical perspective. The use of flexible micropipettes as highly sensitive force probes has become established in the biophysical community, advancing our understanding of cellular processes and microbial behavior. The micropipette force sensor (MFS) technique relies on measurement of the forces acting on a force-calibrated, hollow glass micropipette by optically detecting its deflections. The MFS technique covers a wide micro- and mesoscopic regime of detectable forces (tens of piconewtons to millinewtons) and sample sizes (micrometers to millimeters), does not require gluing of the sample to the cantilever, and allows simultaneous optical imaging of the sample throughout the experiment. Here, we provide a detailed protocol describing how to manufacture and calibrate the micropipettes, as well as how to successfully design, perform, and troubleshoot MFS experiments. We exemplify our approach using the model nematode Caenorhabditis elegans, but by following this protocol, a wide variety of living samples, ranging from single cells to multicellular aggregates and millimeter-sized organisms, can be studied in vivo, with a force resolution as low as 10 pN. A skilled (under)graduate student can master the technique in ~1–2 months. The whole protocol takes ~1–2 d to finish.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout









Similar content being viewed by others
Data availability
The data presented in this protocol are available from the corresponding authors upon request.
References
Kishino, A. & Yanagida, T. Force measurements by micromanipulation of a single actin filament by glass needle. Nature 334, 74–76 (1988).
Houchmandzadeh, B., Marko, J. F., Chatenay, D. & Libchaber, D. Elasticity and structure of eukaryote chromosomes studied by micromanipulation and micropipette aspiration. J. Cell Biol. 139, 1–12 (1997).
Hochmuth, R. M. Micropipette aspiration of living cells. J. Biomech. 33, 15–22 (2000).
Backholm, M., Ryu, W. S. & Dalnoki-Veress, K. Viscoelastic properties of the nematode Caenorhabditis elegans, a self-similar, shear-thinning worm. Proc. Natl Acad. Sci. USA 110, 4528–4533 (2013).
Kamimura, S. & Takahashi, K. Direct measurement of the force of microtubule sliding in flagella. Nature 293, 566–568 (1981).
Marcy, Y., Prost, J., Carlier, M.-F. & Sykes, C. Forces generated during actin-based propulsion: a direct measurement by micromanipulation. Proc. Natl Acad. Sci. USA 101, 5992–5997 (2004).
Schulman, R. D., Backholm, M., Ryu, W. S. & Dalnoki-Veress, K. Dynamic force patterns of an undulatory microswimmer. Phys. Rev. E Stat. Nonlin. Soft Matter Phys. 89, 050701 (2014).
Evans, E. A. Minimum energy analysis of membrane deformation applied to pipet aspiration and surface adhesion of red blood cells. Biophys. J. 30, 265–284 (1980).
Kreis, C. T., Le Blay, M., Linne, C., Makowski, M. M. & Bäumchen, O. Adhesion of Chlamydomonas microalgae to surfaces is switchable by light. Nat. Phys. 14, 45–49 (2018).
Rabets, Y., Backholm, M., Dalnoki-Veress, K. & Ryu, W. S. Direct measurements of drag forces in C. elegans crawling locomotion. Biophys. J. 107, 1980–1987 (2014).
Francis, G. W., Fisher, L. R., Gamble, R. A. & Gingell, D. Direct measurement of cell detachment force on single cells using a new electromechanical method. J. Cell Sci. 87, 519–523 (1987).
Colbert, M.-J., Brochard-Wyart, F., Fradin, C. & Dalnoki-Veress, K. Squeezing and detachment of living cells. Biophys. J. 99, 3555 (2010).
Backholm, M., Kasper, A. K. S., Schulman, R. D., Ryu, W. S. & Dalnoki-Veress, K. The effects of viscosity on the undulatory swimming dynamics of C. elegans. Phys. Fluids 27, 091901 (2015).
Schulman, R. D., Backholm, M., Ryu, W. S. & Dalnoki-Veress, K. Undulatory microswimming near solid boundaries. Phys. Fluids 26, 101902 (2014).
Florin, E.-L., Moy, V. & Gaub, H. E. Adhesion forces between individual ligand-receptor pairs. Science 264, 415–417 (1994).
Neuman, K. C. & Nagy, A. Single-molecule force spectroscopy: optical tweezers, magnetic tweezers and atomic force microscopy. Nat. Methods 5, 491–505 (2008).
Dufrene, Y. F., Martinez-Martin, D., Medalsy, I., Alsteens, D. & Müller, D. J. Multiparametric imaging of biological systems by force-distance curve-based AFM. Nat. Methods 10, 847–854 (2013).
Beaussart, A. et al. Quantifying the forces guiding microbial cell adhesion using single-cell force spectroscopy. Nat. Protoc. 9, 1049–1055 (2014).
Iivarinen, J. T., Korhonen, R. K., Julkunen, P. & Jurvelin, J. S. Experimental and computational analysis of soft tissue stiffness in forearm using a manual indentation device. Med. Eng. Phys. 33, 1245–53 (2011).
Parker, D. et al. A device for characterizing the mechanical properties of the plantar soft tissue of the foot. Med. Eng. Phys. 37, 1098–1104 (2015).
Huber, G. et al. Evidence for capillarity contributions to gecko adhesion from single spatula nanomechanical measurements. Proc. Natl Acad. Sci. USA 102, 16293–16296 (2005).
Autumn, K. et al. Adhesive force of a single gecko foot-hair. Nature 405, 681–685 (2000).
Loskill, P. et al. Macroscale adhesion of gecko setae reflects nanoscale differences in subsurface contribution. J.R. Soc. Interface 10, 20120587 (2012).
Shimamoto, Y. & Kapoor, T. M. Microneedle-based analysis of the micromechanics of the metaphase spindle assembled in Xenopus laevis egg extracts. Nat. Protoc. 7, 959–969 (2012).
Sichel, F. J. M. The elasticity of isolated resting skeletal muscle fibers. J. Cell. Physiol. 5, 21–42 (1934).
Norris, C. H. The tension at the surface, and other physical properties of the nucleated erythrocyte. J. Cell. Physiol. 14, 117–133 (1939).
Coman, D. R. Decreased mutual adhesiveness, a property of cells from squamous cell carcinomas. Cancer Res. 4, 625–629 (1944).
Coman, D. R. Adhesiveness and stickiness: two independent properties of cell surfaces. Cancer Res. 21, 1436–1438 (1961).
Yoneda, M. Force exerted by a single cilium of Mytilus edulis. I. J. Exp. Biol. 37, 461–468 (1960).
Meyhofer, E. & Howard, J. The force generated by a single kinesin molecule against an elastic load. Proc. Natl Acad. Sci. USA 92, 574–578 (1995).
Suda, H. & Yamada, S. Force measurements for the movement of a water drop on a surface with a surface tension gradient. Langmuir 19, 529–531 (2002).
Lagubeau, G., Le Merrer, M., Clanet, C. & Quéré, D. Leidenfrost on a ratchet. Nat. Phys. 7, 395–398 (2011).
Pilat, D. W. et al. Dynamic measurement of the force required to move a liquid drop on a solid surface. Langmuir 28, 16812–16820 (2012).
Daniel, D., Timonen, J. V. I., Li, R., Velling, S. J. & Aizenberg, J. Oleoplaning droplets on lubricated surfaces. Nat. Phys. 13, 1020–1025 (2017).
Gao, N. et al. How drops start sliding over solid surfaces. Nat. Phys. 14, 191–196 (2018).
Mitrossilis, D. et al. Single-cell response to stiffness exhibits muscle-like behavior. Proc. Natl Acad. Sci. USA 106, 18243–18248 (2009).
Mitrossilis, D. et al. Real-time single-cell response to stiffness. Proc. Natl Acad. Sci. USA 107, 16518–16523 (2010).
Bowen, W. R., Lovitt, R. W. & Wright, C. J. Atomic force microscopy study of the adhesion of Saccharomyces cerevisiae. J. Colloid Interface Sci. 237, 54–61 (2001).
Mitchison, J. M. & Swann, M. M. The mechanical properties of the cell surface. J. Exp. Biol. 31, 443–460 (1954).
Evans, E. A. Analysis of adhesion of large vesicles to surfaces. Biophys. J. 31, 425–431 (1980).
Neher, E. & Sakmann, S. Single-channel currents recorded from membrane of denervated frog muscle fibres. Nature 260, 799–802 (1976).
Sakmann, B. & Neher, E. Patch clamp techniques for studying ionic channels in excitable membranes. Annu. Rev. Physiol. 46, 455–472 (1984).
Basu, R. et al. Cytotoxic T cells use mechanical force to potentiate target cell killing. Cell 165, 100–110 (2016).
Guillou, L., Babataheri, A., Puech, P.-H., Barakat, A. I. & Husson, J. Dynamic monitoring of cell mechanical properties using profile microindentation. Sci. Rep. 6, 21529 (2016).
Sawicka, A. et al. Micropipette force probe to quantify single-cell force generation: application to T-cell activation. Mol. Biol. Cell 28, 3229–3239 (2017).
Houchmandzadeh, B. & Dimitrov, S. Elasticity measurements show the existence of thin rigid cores inside mitotic chromosomes. J. Cell Biol. 145, 215–223 (1999).
Poirier, M., Eroglu, S., Chatenay, D. & Marko, J. F. Reversible and irreversible unfolding of mitotic newt chromosomes by applied force. Mol. Biol. Cell 11, 269–276 (2000).
Poirier, M. G., Eroglu, S. & Marko, J. F. The bending rigidity of mitotic chromosomes. Mol. Biol. Cell 13, 2170–2179 (2002).
Moran, K., Yeung, A. & Masliyah, J. Measuring interfacial tensions of micrometer-sized droplets: a novel micromechanical technique. Langmuir 15, 8497–8504 (1999).
Yeung, A. K. C. & Pelton, R. Micromechanics: a new approach to studying the strength and breakup of flocs. J. Colloid Interface Sci. 184, 579–585 (1996).
Poppele, E. H. & Hozalski, R. M. Micro-cantilever method for measuring the tensile strength of biofilms and microbial flocs. J. Microbiol. Methods 55, 607–615 (2003).
Tsang, P. H., Li, G., Brun, Y. V., Freund, L. B. & Tang, J. X. Adhesion of single bacterial cells in the micronewton range. Proc. Natl Acad. Sci. USA 103, 5764–5768 (2006).
Schulman, R. D. & Dalnoki-Veress, K. Liquid droplets on a highly deformable membrane. Phys. Rev. Lett. 115, 206101 (2015).
’t Mannetje, D. et al. Electrically tunable wetting defects characterized by a simple capillary force sensor. Langmuir 29, 9944–9949 (2013).
Frostad, J. M., Collins, M. C. & Leal, L. G. Cantilevered-capillary force apparatus for measuring multiphase fluid interactions. Langmuir 29, 4715–4725 (2013).
Frostad, J. M., Seth, M., Bernasek, S. M. & Leal, L. G. Direct measurement of interaction forces between charged multilamellar vesicles. Soft Matter 10, 7769–7780 (2014).
Frostad, J. M., Collins, M. C. & Leal, L. G. Direct measurement of the interaction of model food emulsion droplets adhering by arrested coalescence. Colloids Surf. A 441, 459–465 (2014).
Colbert, M.-J., Raegen, A. N., Fradin, C. & Dalnoki-Veress, K. Adhesion and membrane tension of single vesicles and living cells using a micropipette-based technique. Eur. Phys. J. E Soft Matter 30, 117 (2009).
Backholm, M., Ryu, W. S. & Dalnoki-Veress, K. The nematode C. elegans as a complex viscoelastic fluid. Eur. Phys. J. E Soft Matter 38, 36 (2015).
Backholm, M., Schulman, R. D., Ryu, W. S. & Dalnoki-Veress, K. Tangling of tethered swimmers: Interactions between two nematodes. Phys. Rev. Lett. 113, 138101 (2014).
Rüffer, U. & Nultsch, W. Comparison of the beating of cis- and trans-flagella of Chlamydomonas cells held on micropipettes. Cell Motil. Cytoskeleton 7, 87–93 (1987).
Rüffer, U. & Nultsch, W. Flagellar photoresponses of Chlamydomonas cells held on micropipettes: II. Change in flagellar beat pattern. Cell Motil. Cytoskeleton 18, 269–278 (1991).
Wan, K. Y. & Goldstein, R. E. Coordinated beating of algal flagella is mediated by basal coupling. Proc. Natl Acad. Sci. USA 113, E2784–E2793 (2016).
Drescher, K., Goldstein, R. E., Michel, N., Polin, M. & Tuval, I. Direct measurement of the flow field around swimming microorganisms. Phys. Rev. Lett. 105, 168101 (2010).
Harz, H. & Hegemann, P. Rhodopsin-regulated calcium currents in Chlamydomonas. Nature 351, 489–491 (1991).
Petit, J. et al. A modular approach for multifunctional polymersomes with controlled adhesive properties. Soft Matter 14, 894–900 (2018).
Kee, Y. S. & Robinson, D. N. Micropipette aspiration for studying cellular mechanosensory responses and mechanics. Methods Mol. Biol. 983, 367–382 (2013).
Biro, M. & Maître, J. L. Dual pipette aspiration: a unique tool for studying intercellular adhesion. Methods Cell Biol. 125, 255–267 (2015).
Guevorkian, K. & Maître, J. L. Micropipette aspiration: a unique tool for exploring cell and tissue mechanics in vivo. Methods Cell Biol. 139, 187–201 (2017).
Acknowledgements
M.B. gratefully acknowledges support from the Academy of Finland (Centres of Excellence Programme (2014–2019, grant agreement no. 272361) and the Postdoctoral Researcher Project (grant agreement no. 309237)). O.B. acknowledges funding from the German Research Foundation (DFG) under grant BA3406/2. The authors are deeply grateful to K. Dalnoki-Veress for inspiring discussions and support. R.D. Schulman, C.T. Kreis, M.M. Makowski, T. Böddeker, and Q. Magdelaine are acknowledged for sharing their hands-on experiences and for valuable technical suggestions regarding improvements to the protocol.
Author information
Authors and Affiliations
Contributions
M.B. and O.B. developed the protocol and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key reference using this protocol
Backholm, M., Ryu, W. S. & Dalnoki-Veress, K. Proc. Natl Acad. Sci. USA 110, 4528–4533 (2013): http://www.pnas.org/content/110/12/4528
Backholm, M., Kasper, A. K. S., Schulman, R. D., Ryu, W. S. & Dalnoki-Veress, K. Phys. Fluids 27, 091901 (2015): https://aip.scitation.org/doi/10.1063/1.4931795
Kreis, C. T., Le Blay, M., Linne, C., Makowski, M. M. & Bäumchen, O. Nat. Phys. 14, 45–49 (2018): https://www.nature.com/articles/nphys4258
Supplementary information
Supplementary Video 1
Real-time video from an MFS measurement of a swimming C. elegans nematode (in a 10% (wt/vol) PEO-M9 buffer solution) in which a three-dimensional micropipette is used to measure both lateral and propulsive drag forces.
Supplementary Software 1
Matlab code ‘calibration.m’ for determining the pipette spring constant in Calibration Option A.
Supplementary Software 2
Matlab code ‘deflection.m’ for analysing the pipette deflection via a cross-correlation approach.
Rights and permissions
About this article
Cite this article
Backholm, M., Bäumchen, O. Micropipette force sensors for in vivo force measurements on single cells and multicellular microorganisms. Nat Protoc 14, 594–615 (2019). https://doi.org/10.1038/s41596-018-0110-x
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-018-0110-x
This article is cited by
-
Droplet slipperiness despite surface heterogeneity at molecular scale
Nature Chemistry (2024)
-
Probing surface wetting across multiple force, length and time scales
Communications Physics (2023)
-
Long-term stability of aerophilic metallic surfaces underwater
Nature Materials (2023)
-
3D mechanical characterization of single cells and small organisms using acoustic manipulation and force microscopy
Nature Communications (2021)
-
Water droplet friction and rolling dynamics on superhydrophobic surfaces
Communications Materials (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.