Abstract
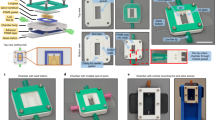
Microengineered biomimetic systems for organ-on-a-chip or tissue engineering purposes often fail as a result of an inability to recapitulate the in vivo environment, specifically the presence of a well-defined vascular system. To address this limitation, we developed an alternative method to cultivate three-dimensional (3D) tissues by incorporating a microfabricated scaffold, termed AngioChip, with a built-in perfusable vascular network. Here, we provide a detailed protocol for fabricating the AngioChip scaffold, populating it with endothelial cells and parenchymal tissues, and applying it in organ-on-a-chip drug testing in vitro and surgical vascular anastomosis in vivo. The fabrication of the AngioChip scaffold is achieved by a 3D stamping technique, in which an intricate microchannel network can be embedded within a 3D scaffold. To develop a vascularized tissue, endothelial cells are cultured in the lumen of the AngioChip network, and parenchymal cells are encapsulated in hydrogels that are amenable to remodeling around the vascular network to form functional tissues. Together, these steps yield a functional, vascularized network in vitro over a 14-d period. Finally, we demonstrate the functionality of AngioChip-vascularized hepatic and cardiac tissues, and describe direct surgical anastomosis of the AngioChip vascular network on the hind limb of a Lewis rat model.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Betts, J. G. et al. Anatomy and Physiology (Open Stax College, Houston, TX, 2013).
Langer, R. & Vacanti, J. P. Tissue engineering. Science 260, 920–926 (1993).
Lovett, M., Lee, K., Edwards, A. & Kaplan, D. L. Vascularization strategies for tissue engineering. Tissue Eng. Part B Rev. 15, 353–370 (2009).
Atala, A., Kasper, F. K. & Mikos, A. G. Engineering complex tissues. Sci. Transl. Med. 4, 160rv112 (2012).
Miller, J. S. The billion cell construct: will three-dimensional printing get us there? PLoS Biol. 12, e1001882 (2014).
Montgomery, M., Zhang, B. & Radisic, M. Cardiac tissue vascularization from angiogenesis to microfluidic blood vessels. J. Cardiovasc. Pharmacol. Ther. 19, 382–393 (2014).
Ingber, D. E. Reverse engineering human pathophysiology with organs-on-chips. Cell 164, 1105–1109 (2016).
Zhang, B. & Radisic, M. Organ-on-a-chip devices advance to market. Lab Chip 17, 2395–2420 (2017).
Esch, E. W., Bahinski, A. & Huh, D. Organs-on-chips at the frontiers of drug discovery. Nat. Rev. Drug Discov. 14, 248–260 (2015).
Takebe, T., Zhang, B. & Radisic, M. Synergistic engineering: organoids meet organs-on-a-chip. Cell Stem Cell 21, 297–300 (2017).
Ahadian, S. et al. Organ-on-a-chip platforms: a convergence of advanced materials, cells, and microscale technologies. Adv. Healthc. Mater. 7, 1700506 (2018).
Young, E. W. K. & Simmons, C. A. Macro- and microscale fluid flow systems for endothelial cell biology. Lab Chip 10, 143–160 (2010).
Zheng, Y. et al. In vitro microvessels for the study of angiogenesis and thrombosis. Proc. Natl. Acad. Sci. USA 109, 9342–9347 (2012).
Morgan, J. P. et al. Formation of microvascular networks in vitro. Nat. Protoc. 8, 1820 (2013).
Hsieh, P. C. H., Davis, M. E., Lisowski, L. K. & Lee, R. T. Endothelial-cardiomyocyte interactions in cardiac development and repair. Annu. Rev. Physiol. 68, 51–66 (2006).
Lai, B. F. L. et al. InVADE: integrated vasculature for assessing dynamic events. Adv. Funct. Mater. 27, 1703524 (2017).
Chiu, L. L., Montgomery, M., Liang, Y., Liu, H. & Radisic, M. Perfusable branching microvessel bed for vascularization of engineered tissues. Proc. Natl. Acad. Sci. USA 109, E3414–E3423 (2012).
Bae, H. et al. Building vascular networks. Sci. Transl. Med. 4, 160ps123 (2012).
Ren, X. et al. Engineering pulmonary vasculature in decellularized rat and human lungs. Nat. Biotechnol. 33, 1097–1102 (2015).
Ott, H. C. et al. Perfusion-decellularized matrix: using nature’s platform to engineer a bioartificial heart. Nat. Med. 14, 213–221 (2008).
Ott, H. C. et al. Regeneration and orthotopic transplantation of a bioartificial lung. Nat. Med. 16, 927–933 (2010).
Kolesky, D. B. et al. 3D bioprinting of vascularized, heterogeneous cell-laden tissue constructs. Adv. Mater. 26, 3124–3130 (2014).
Miller, J. S. et al. Rapid casting of patterned vascular networks for perfusable engineered three-dimensional tissues. Nat. Mater. 11, 768–774 (2012).
Vollert, I. et al. In vitro perfusion of engineered heart tissue through endothelialized channels. Tissue Eng. Part A 20, 854–863 (2013).
Tang, M. D., Golden, A. P. & Tien, J. Fabrication of collagen gels that contain patterned, micrometer-scale cavities. Adv. Mater. 16, 1345–1348 (2004).
Golden, A. P. & Tien, J. Fabrication of microfluidic hydrogels using molded gelatin as a sacrificial element. Lab Chip 7, 720–725 (2007).
Zimmermann, W.-H. et al. Engineered heart tissue grafts improve systolic and diastolic function in infarcted rat hearts. Nat. Med. 12, 452–458 (2006).
Nunes, S. S. et al. Biowire: a platform for maturation of human pluripotent stem cell-derived cardiomyocytes. Nat. Methods 10, 781–787 (2013).
Leng, L., McAllister, A., Zhang, B., Radisic, M. & Günther, A. Mosaic hydrogels: one‐step formation of multiscale soft materials. Adv. Mater. 24, 3650–3658 (2012).
Kolesky, D. B. et al. 3D bioprinting of vascularized, heterogeneous cell‐laden tissue constructs. Adv. Mater. 26, 3124–3130 (2014).
Kolesky, D. B., Homan, K. A., Skylar-Scott, M. A. & Lewis, J. A. Three-dimensional bioprinting of thick vascularized tissues. Proc. Natl Acad. Sci. USA 113, 3179–3184 (2016).
Kang, H.-W. et al. A 3D bioprinting system to produce human-scale tissue constructs with structural integrity. Nat. Biotechnol. 34, 312–319 (2016).
Engelmayr, G. C. et al. Accordion-like honeycombs for tissue engineering of cardiac anisotropy. Nat. Mater. 7, 1003–1010 (2008).
Kolewe, M. E. et al. 3D structural patterns in scalable, elastomeric scaffolds guide engineered tissue architecture. Adv. Mater. 25, 4459–4465 (2013).
Xiao, Y. et al. Microfabricated perfusable cardiac biowire: a platform that mimics native cardiac bundle. Lab Chip 14, 869–882 (2014).
Zhang, B. et al. Biodegradable scaffold with built-in vasculature for organ-on-a-chip engineering and direct surgical anastomosis. Nat. Mater. 15, 669–678 (2016).
Zhang, B., Montgomery, M., Davenport-Huyer, L., Korolj, A. & Radisic, M. Platform technology for scalable assembly of instantaneously functional mosaic tissues. Sci. Adv. 1, e1500423 (2015).
Davenport Huyer, L. et al. Highly elastic and moldable polyester biomaterial for cardiac tissue engineering applications. ACS Biomater. Sci. Eng. 2, 780–788 (2016).
Zhang, B., Peticone, C., Murthy, S. K. & Radisic, M. A standalone perfusion platform for drug testing and target validation in micro-vessel networks. Biomicrofluidics 7, 044125 (2013).
Sekine, H. et al. In vitro fabrication of functional three-dimensional tissues with perfusable blood vessels. Nat. Commun. 4, 1399 (2013).
Glaser, A. K. et al. Light-sheet microscopy for slide-free non-destructive pathology of large clinical specimens. Nat. Biomed. Eng. 1, 0084 (2017).
Perbellini, F. et al. Free-of-acrylamide SDS-based tissue clearing (FASTClear) for three dimensional visualization of myocardial tissue. Sci. Rep. 7, 5188 (2017).
Chung, K. & Deisseroth, K. Clarity for mapping the nervous system. Nat. Methods 10, 508–513 (2013).
Chung, K. et al. Structural and molecular interrogation of intact biological systems. Nature 497, 332–337 (2013).
Coppeta, J. et al. A portable and reconfigurable multi-organ platform for drug development with onboard microfluidic flow control. Lab Chip 17, 134–144 (2017).
Huh, D. et al. Reconstituting organ-level lung functions on a chip. Science 328, 1662–1668 (2010).
Shin, Y. et al. Microfluidic assay for simultaneous culture of multiple cell types on surfaces or within hydrogels. Nat. Protoc. 7, 1247–1259 (2012).
Tran, R. T. et al. Synthesis and characterization of a biodegradable elastomer featuring a dual crosslinking mechanism. Soft Matter. 6, 2449–2461 (2010).
Yang, J., Webb, A. R. & Ameer, G. A. Novel citric acid-based biodegradable elastomers for tissue engineering. Adv. Mater. 16, 511–516 (2004).
Montgomery, M. et al. Flexible shape-memory scaffold for minimally invasive delivery of functional tissues. Nat. Mater. 16, 1038–1046 (2017).
Khan, O. F., Voice, D. N., Leung, B. M. & Sefton, M. V. A novel high‐speed production process to create modular components for the bottom‐up assembly of large‐scale tissue‐engineered constructs. Adv. Healthc. Mater. 4, 113–120 (2015).
Chamberlain, M. D., Gupta, R. & Sefton, M. V. Bone marrow-derived mesenchymal stromal cells enhance chimeric vessel development driven by endothelial cell-coated microtissues. Tissue Eng. A 18, 285–294 (2012).
Radisic, M. et al. Functional assembly of engineered myocardium by electrical stimulation of cardiac myocytes cultured on scaffolds. Proc. Natl. Acad. Sci. USA 101, 18129–18134 (2004).
Zhang, B., Green, J. V., Murthy, S. K. & Radisic, M. Label-free enrichment of functional cardiomyocytes using microfluidic deterministic lateral flow displacement. PLoS ONE 7, e37619 (2012).
Ogawa, M. et al. Directed differentiation of cholangiocytes from human pluripotent stem cells. Nat. Biotechnol. 33, 853–861 (2015).
Ogawa, S. et al. Three-dimensional culture and cAMP signaling promote the maturation of human pluripotent stem cell-derived hepatocytes. Development 140, 3285–3296 (2013).
Lian, X. et al. Directed cardiomyocyte differentiation from human pluripotent stem cells by modulating Wnt/β-catenin signaling under fully defined conditions. Nat. Protoc. 8, 162–175 (2013).
Levario, T. J., Zhan, M., Lim, B., Shvartsman, S. Y. & Lu, H. Microfluidic trap array for massively parallel imaging of Drosophila embryos. Nat. Protoc. 8, 721–736 (2013).
Huh, D. et al. Microfabrication of human organs-on-chips. Nat. Protoc. 8, 2135–2157 (2013).
Kim, J. et al. A microfluidic technique for quantification of steroids in core needle biopsies. Anal. Chem. 87, 4688–4695 (2015).
Di, L. et al. Optimization of a higher throughput microsomal stability screening assay for profiling drug discovery candidates. J. Biomol. Screen. 8, 453–462 (2003).
Hiebert, L. M., Ping, T. & Wice, S. M. Repeated doses of oral and subcutaneous heparins have similar antithrombotic effects in a rat carotid arterial model of thrombosis. J. Cardiovasc. Pharmacol. Ther. 17, 110–116 (2012).
Ross, M. H. & Pawlina, W. Histology (Lippincott Williams & Wilkins, Philadelphia, 2006).
Zhang, K., Rekhter, M. D., Gordon, D. & Phan, S. H. Myofibroblasts and their role in lung collagen gene expression during pulmonary fibrosis. A combined immunohistochemical and in situ hybridization study. Am. J. Pathol. 145, 114 (1994).
JoVE Science Education Database. General laboratory techniques: histological sample preparation for light microscopy, https://www.jove.com/science-education/5039/histological-sample-preparation-for-light-microscopy (JoVE, Cambridge, MA, 2018).
Acknowledgements
This work was made possible by a National Sciences and Engineering Research Council of Canada (NSERC) Postgraduate Scholarship–Doctoral, awarded to B.F.L.L.; a Canadian Institutes of Health Research (CIHR) Vanier Canada Graduate Scholarship, awarded to L.D.H.; and a CIHR Banting Postdoctoral Fellowship, awarded to B.Z. This work was also funded by CIHR Operating Grants (MOP-126027 and MOP-137107), an NSERC Discovery Grant (RGPIN-2015-05952), an NSERC Steacie Fellowship (SMFSU 4620), a Heart and Stroke Foundation Grant-in-Aid (G-16-00012), an NSERC-CIHR Collaborative Health Research Grant (CHRPJ 4937), an NSERC Strategic Grant (STPGP 5066) to M.R., and National Institutes of Health grant 2R01 HL076485.
Author information
Authors and Affiliations
Contributions
B.Z. and M.R. designed the research; B.Z., L.D.H., and M.M. performed the research; B.Z. and M.R. analyzed the data; B.Z. and R.X. prepared the figures for the paper; B.Z., B.F.L.L., and R.X. prepared the supplementary video; B.Z., B.F.L.L., and M.R. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
M.R. and B.Z. are among the co-founders of TARA Biosystems and they hold equity in this company. The AngioChip is licensed to TARA Biosystems. The remaining authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Zhang, B. et al. Nat. Mater. 15, 669–678 (2016) https://doi.org/10.1038/nmat4570
Lai., B. F. L. et al. Adv. Funct. Mater. 27, 1703524 (2017) https://doi.org/10.1002/adfm.201703524
Zhang, B. et al. Sci. Adv. 1, e1500423 (2015) https://doi.org/10.1126/sciadv.1500423
Integrated supplementary information
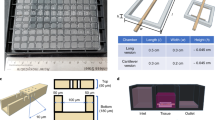
Supplementary Figure 1 Bioreactor design.
All units are in millimeters unless specified otherwise. All bioreactor components are made of polycarbonate.
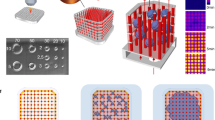
Supplementary Figure 2 Endothelialized AngioChip network.
Confocal fluorescent images of various location of the AngioChip network populated with human umbilical cord endothelial cells stained for VE-cadherin (red) to identify the intercellular junctions (red) on day 2 after cell seeding. Scale bar, 50 μm. Reproduced with permission from Zhang et al.36, Macmillan Publishers Limited.
Supplementary Figure 3 Angiogenic vascular sprouting from AngioChip.
Confocal fluorescent images of GFP-labeled human umbilical cord endothelial cells (green) sprouting from the AngioChip networks through the built-in micro-holes on the channel walls in the presence of thymosin β4 on day 2 after cell seeding. Scale bar, 50μm.
Supplementary Figure 4 Tissue integration of AngioChip implants.
Smooth muscle actin (SMA) stained histology cross-section of the AngioChip cardiac tissue implants after 1 week with the direct surgical anastomosis in the configuration of artery-to-vein graft. Scale bar: 200μm (top image) and 100μm (bottom image). Reproduced with permission from Zhang et al.36, Macmillan Publishers Limited.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4
Supplementary Data
CAD file containing AngioChip design. Dimensions of four different AngioChips with increasing complexity. The default unit for dimensions in AutoCAD is the millimeter. The thickness of all layers is specified in micrometers
Supplementary Video 1
Step-by-step video protocol on the fabrication of AngioChip scaffolds. Step number in the video corresponds to the step number in the main protocol
Rights and permissions
About this article
Cite this article
Zhang, B., Lai, B.F.L., Xie, R. et al. Microfabrication of AngioChip, a biodegradable polymer scaffold with microfluidic vasculature. Nat Protoc 13, 1793–1813 (2018). https://doi.org/10.1038/s41596-018-0015-8
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-018-0015-8
This article is cited by
-
Biodegradable elastomeric circuit boards from citric acid-based polyesters
npj Flexible Electronics (2023)
-
Opportunities and challenges in cardiac tissue engineering from an analysis of two decades of advances
Nature Biomedical Engineering (2022)
-
Microvascularized tumor organoids-on-chips: advancing preclinical drug screening with pathophysiological relevance
Nano Convergence (2021)
-
A well plate–based multiplexed platform for incorporation of organoids into an organ-on-a-chip system with a perfusable vasculature
Nature Protocols (2021)
-
Vascularisation of pluripotent stem cell–derived myocardium: biomechanical insights for physiological relevance in cardiac tissue engineering
Pflügers Archiv - European Journal of Physiology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.