Abstract
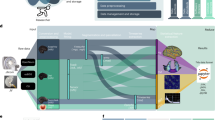
Human intracranial electroencephalography (iEEG) recordings provide data with much greater spatiotemporal precision than is possible from data obtained using scalp EEG, magnetoencephalography (MEG), or functional MRI. Until recently, the fusion of anatomical data (MRI and computed tomography (CT) images) with electrophysiological data and their subsequent analysis have required the use of technologically and conceptually challenging combinations of software. Here, we describe a comprehensive protocol that enables complex raw human iEEG data to be converted into more readily comprehensible illustrative representations. The protocol uses an open-source toolbox for electrophysiological data analysis (FieldTrip). This allows iEEG researchers to build on a continuously growing body of scriptable and reproducible analysis methods that, over the past decade, have been developed and used by a large research community. In this protocol, we describe how to analyze complex iEEG datasets by providing an intuitive and rapid approach that can handle both neuroanatomical information and large electrophysiological datasets. We provide a worked example using an example dataset. We also explain how to automate the protocol and adjust the settings to enable analysis of iEEG datasets with other characteristics. The protocol can be implemented by a graduate student or postdoctoral fellow with minimal MATLAB experience and takes approximately an hour to execute, excluding the automated cortical surface extraction.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Buzsaki, G., Anastassiou, C. A. & Koch, C. The origin of extracellular fields and currents--EEG, ECoG, LFP and spikes. Nat. Rev. Neurosci. 13, 407–420 (2012).
Malmivuo, J. & Plonsey, R. Bioelectromagnetism: Principles and Applications of Bioelectric and Biomagnetic Fields. Bioelectromagnetism: Principles and Applications of Bioelectric and Biomagnetic Fields (Oxford University Press, New York, 2012).
Brunner, P. et al. A practical procedure for real-time functional mapping of eloquent cortex using electrocorticographic signals in humans. Epilepsy Behav. 15, 278–286 (2009).
Ritaccio, A. et al. Proceedings of the fifth international workshop on advances in electrocorticography. Epilepsy Behav. 41, 183–92 (2014)
Lachaux, J.-P., Axmacher, N., Mormann, F., Halgren, E. & Crone, N. E. High-frequency neural activity and human cognition: past, present and possible future of intracranial EEG research. Prog. Neurobiol. 98, 279–301 (2012).
Friston, J. A. & Friston, K. Multimodal image coregistration and partitioning - a unified fFramework. Neuroimage 6, 209–217 (1997).
Jenkinson, M., Bannister, P., Brady, M. & Smith, S. Improved optimization for the robust and accurate linear registration and motion correction of brain images. Neuroimage 17, 825–841 (2002).
Cox, R. W. AFNI: software for analysis and visualization of functional magnetic resonance neuroimages. Comput. Biomed. Res. 29, 162–73 (1996).
Papademetris, X. et al. BioImage suite: an integrated medical image analysis suite: an update. Insight J. 2006, 209 (2006).
Azarion, A. A. et al. An open-source automated platform for three-dimensional visualization of subdural electrodes using CT-MRI coregistration. Epilepsia 55, 2028–2037 (2014).
Blenkmann, A. O. et al. iElectrodes: a comprehensive open-source Toolbox for depth and subdural grid electrode localization. Front. Neuroinform. 11, 14 (2017).
Groppe, D. M. et al. iELVis: an open source MATLAB toolbox for localizing and visualizing human intracranial electrode data. J. Neurosci. Methods 281, 40–48 (2017).
Kubanek, J. & Schalk, G. NeuralAct: a tool to visualize electrocortical (ECoG) activity on a three-dimensional model of the cortex. Neuroinformatics 13, 167–174 (2015).
Qin, C. et al. Automatic and precise localization and cortical labeling of subdural and depth intracranial electrodes. Front. Neuroinform. 11, 1–10 (2017).
Hill, N. J. et al. Recording human electrocorticographic (ECoG) signals for neuroscientific research and real-time functional cortical mapping. J. Vis. Exp. https://doi.org/10.3791/3993(2012).
LaPlante, R. A. et al. The interactive electrode localization utility: software for automatic sorting and labeling of intracranial subdural electrodes. Int. J. Comput. Assist. Radiol. Surg. 12, 1829–1837 (2017).
Branco, M. P. et al. ALICE: a tool for automatic localization of intra-cranial electrodes for clinical and high-density grids. J. Neurosci. Methods 301, 43–51 (2018).
Eglen, S. J. et al. Toward standard practices for sharing computer code and programs in neuroscience. Nat. Neurosci. 20, 770–773 (2017).
Delorme, A. & Makeig, S. EEGLAB: an open source toolbox for analysis of single-trial EEG dynamics including independent component analysis. J. Neurosci. Methods 134, 9–21 (2004).
Zheng, J. et al. Amygdala-hippocampal dynamics during salient information processing. Nat. Commun. 8, 14413 (2017).
Tang, C., Hamilton, L. S. & Chang, E. F. Intonational speech prosody encoding in the human auditory cortex. Science 357, 797–801 (2017).
Martinet, L.-E. et al. Human seizures couple across spatial scales through travelling wave dynamics. Nat. Commun. 8, 14896 (2017).
Gelinas, J. N., Khodagholy, D., Thesen, T., Devinsky, O. & Buzsáki, G. Interictal epileptiform discharges induce hippocampal–cortical coupling in temporal lobe epilepsy. Nat. Med. 22, 641–648 (2016).
Piai, V. et al. Direct brain recordings reveal hippocampal rhythm underpinnings of language processing. Proc. Natl Acad. Sci. USA 113, 11366–11371 (2016).
Hermes, D., Miller, K. J., Noordmans, H. J., Vansteensel, M. J. & Ramsey, N. F. Automated electrocorticographic electrode localization on individually rendered brain surfaces. J. Neurosci. Methods 185, 293–298 (2010).
Dalal, S. S. et al. Localization of neurosurgically implanted electrodes via photograph-MRI-radiograph coregistration. J. Neurosci. Methods 174, 106–115 (2008).
Yang, A. I. et al. Localization of dense intracranial electrode arrays using magnetic resonance imaging. Neuroimage 63, 157–165 (2012).
Onofrey, J. A., Staib, L. H. & Papademetris, X. Learning intervention-induced deformations for non-rigid MR-CT registration and electrode localization in epilepsy patients. NeuroImage Clin. 10, 291–301 (2016).
Pieters, T. A., Conner, C. R. & Tandon, N. Recursive grid partitioning on a cortical surface model: an optimized technique for the localization of implanted subdural electrodes. J. Neurosurg. 118, 1086–1097 (2013).
Stieglitz, L. H. et al. Improved localization of implanted subdural electrode contacts on magnetic resonance imaging with an elastic image fusion algorithm in an invasive electroencephalography recording. Clin. Neurosurg. 10, 506–513 (2014).
Brang, D., Dai, Z., Zheng, W. & Towle, V. L. Registering imaged ECoG electrodes to human cortex: a geometry-based technique. J. Neurosci. Methods 273, 64–73 (2016).
Dykstra, A. R. et al. Individualized localization and cortical surface-based registration of intracranial electrodes. Neuroimage 59, 3563–3570 (2012).
Khodagholy, D. et al. Organic electronics for high-resolution electrocorticography of the human brain. Sci. Adv. 2, 1–9 (2016).
Seo, D. et al. Wireless recording in the peripheral nervous system with ultrasonic neural dust. Neuron 91, 529–539 (2016).
Lauro, P. M. et al. DBSproc: an open source process for DBS electrode localization and tractographic analysis. Hum. Brain Mapp. 37, 422–433 (2016).
Horn, A. & Kühn, A. A. Lead-DBS: a toolbox for deep brain stimulation electrode localizations and visualizations. Neuroimage 107, 127–135 (2015).
Dale, A. M., Fischl, B. & Sereno, M. I. Cortical surface-based analysis: I. Segmentation and surface reconstruction. Neuroimage 9, 179–194 (1999).
Lepore, N. et al. A new combined surface and volume registration. Med. Imaging 2010 Image Process. 7623, 76231E https://doi.org/10.1117/12.844434(2010).
Klein, A. et al. Evaluation of volume-based and surface-based brain image registration methods. Neuroimage 51, 214–220 (2010).
Hill, D. L. G. et al. Measurement of intraoperative brain surface deformation under a craniotomy. Neurosurgery 43, 514–526 (1998).
Roberts, D. W., Hartov, A., Kennedy, F. E., Miga, M. I. & Paulsen, K. D. Intraoperative brain shift and deformation: a quantitative analysis of cortical displacement in 28 cases. Neurosurgery 43, 749–758 (1998).
Miyagi, Y., Shima, F. & Sasaki, T. Brain shift: an error factor during implantation of deep brain stimulation electrodes. J. Neurosurg. 107, 989–97 (2007).
Hastreiter, P. et al. Strategies for brain shift evaluation. Med. Image Anal. 8, 447–464 (2004).
LaViolette, P. S. et al. Three-dimensional visualization of subdural electrodes for presurgical planning. Oper. Neurosurg. 68 https://doi.org/10.1227/NEU.0b013e31820783ba (2011).
Sweet, J. A., Hdeib, A. M., Sloan, A. & Miller, J. P. Depths and grids in brain tumors: Implantation strategies, techniques, and complications. Epilepsia 54, 66–71 (2013).
Kovalev, D. et al. Rapid and fully automated visualization of subdural electrodes in the presurgical evaluation of epilepsy patients. Am. J. Neuroradiol. 26, 1078–1083 (2005).
Wang, P. T. et al. A co-registration approach for electrocorticogram electrode localization using post-implantation MRI and CT of the head. in Proc. International. IEEE/EMBS Conference on Neural Engineering, NER 525–528 https://doi.org/10.1109/NER.2013.6695987(2013).
Schulze-Bonhage, A. H. J. et al. Visualization of subdural strip and grid electrodes using curvilinear reformatting of 3D MR imaging data sets. Am. J. Neuroradiol. 23, 400–403 (2002).
Boatman-Reich, D. et al. Quantifying auditory event-related responses in multichannel human intracranial recordings. Front. Comput. Neurosci. 4, 4 (2010).
Staresina, B. P. et al. Hierarchical nesting of slow oscillations, spindles and ripples in the human hippocampus during sleep. Nat. Neurosci. 18, 1679–1686 (2015).
Manning, J. R., Jacobs, J., Fried, I. & Kahana, M. J. Broadband shifts in local field potential power spectra are correlated with single-neuron spiking in humans. J. Neurosci. 29, 13613–13620 (2009).
Miller, K. J. Broadband spectral change: evidence for a macroscale correlate of population firing rate? J. Neurosci. 30, 6477–6479 (2010).
Ray, S. & Maunsell, J. H. R. Different origins of gamma rhythm and high-gamma activity in macaque visual cortex. PLoS Biol. 9 https://doi.org/10.1371/journal.pbio.1000610 (2011).
Crone, N. E., Miglioretti, D. L., Gordon, B., Lesser, R. P. & Crone, N. Functional mapping of human sensorimotor cortex with electrocorticographic spectral analysis II. Event-related synchronization in the gamma band. Brain 121, 2301–2315 (1998).
Oostenveld, R., Fries, P., Maris, E. & Schoffelen, J. M. FieldTrip: open source software for advanced analysis of MEG, EEG, and invasive electrophysiological data. Comput. Intell. Neurosci. 2011 https://doi.org/10.1155/2011/156869 (2011).
Maris, E. & Oostenveld, R. Nonparametric statistical testing of EEG- and MEG-data. J. Neurosci. Methods 164, 177–190 (2007).
Bastos, A. M. & Schoffelen, J.-M. A tutorial review of functional connectivity analysis methods and their interpretational pitfalls. Front. Syst. Neurosci. 9, 1–23 (2016).
Drury, H. A., Van Essen, D. C., Corbetta, M. & Snyder, A. Z. Brain Warping 337–363 (Elsevier, Cambridge, MA, 1999).
Wells, W. M., Viola, P., Atsumi, H., Nakajima, S. & Kikinis, R. Multi-modal volume registration by maximization of mutual information. Med. Image Anal. 1, 35–51 (1996).
Collignon, A. & Maes, F. Automated multi-modality image registration based on information theory. Proc. Inf. Process. Med. Imaging 263–274 (1995).
Schaer, M. et al. A surface-based approach to quantify local cortical gyrification. IEEE Trans. Med. Imaging 27, 161–170 (2008).
Lancaster, J. L. et al. Automated labeling of the human brain: a preliminary report on the development and evaluation of a forward-transform method. Hum. Brain Mapp. 5, 238–242 (1997).
Tzourio-Mazoyer, N. et al. Automated anatomical labeling of activations in SPM using a macroscopic anatomical parcellation of the MNI MRI single-subject brain. Neuroimage 15, 273–289 (2002).
Cocosco, C. A., Kollokian, V., Kwan, R. K., Pike, G. B. & Evans, A. C. BrainWeb : online interface to a 3D MRI simulated brain database. Proc. 3rd Int. Conf. Funct. Mapp. Hum. Brain 5, S425 (1997). in.
Eickhoff, S. B. et al. A new SPM toolbox for combining probabilistic cytoarchitectonic maps and functional imaging data. Neuroimage 25, 1325–1335 (2005).
Wang, L., Mruczek, R. E. B., Arcaro, M. J. & Kastner, S. Probabilistic maps of visual topography in human cortex. Cereb. Cortex 25, 3911–3931 (2015).
Fan, L. et al. The human brainnetome atlas: a new brain atlas based on connectional architecture. Cereb. Cortex 26, 3508–3526 (2016).
Desikan, R. S. et al. An automated labeling system for subdividing the human cerebral cortex on MRI scans into gyral based regions of interest. Neuroimage 31, 968–980 (2006).
Destrieux, C., Fischl, B., Dale, A. & Halgren, E. Automatic parcellation of human cortical gyri and sulci using standard anatomical nomenclature. Neuroimage 53, 1–15 (2010).
Ashburner, J. & Friston, K. J. Nonlinear spatial normalization using basis functions. Hum. Brain Mapp. 7, 254–266 (1999).
Bigdely-Shamlo, N., Mullen, T., Kothe, C., Su, K.-M. & Robbins, K. A. The PREP pipeline: standardized preprocessing for large-scale EEG analysis. Front. Neuroinform. 9, 16 (2015).
Liu, Y., Coon, W. G., Pesters, A., de Brunner, P. & Schalk, G. The effects of spatial filtering and artifacts on electrocorticographic signals. J. Neural Eng. 12, 56008 (2015).
Dien, J. Issues in the application of the average reference: review, critiques, and recommendations. Behav. Res. Methods, Instrum., Comput. 30, 34–43 (1998).
Ludwig, K. A. et al. Using a common average reference to improve cortical neuron recordings from microelectrode arrays. J. Neurophysiol. 101, 1679–1689 (2009).
Trongnetrpunya, A. et al. Assessing Granger causality in electrophysiological data: removing the adverse effects of common signals via bipolar derivations. Front. Syst. Neurosci. 9, 189 (2015).
Shirhatti, V., Borthakur, A. & Ray, S. Effect of reference scheme on power and phase of the local field potential. Neural Comput. 882–913 https://doi.org/10.1162/NECO (2016).
Arnulfo, G., Hirvonen, J., Nobili, L., Palva, S. & Palva, J. M. Phase and amplitude correlations in resting-state activity in human stereotactical EEG recordings. Neuroimage 112, 114–127 (2015).
Zaveri, H. P., Duckrow, R. B. & Spencer, S. S. On the use of bipolar montages for time-series analysis of intracranial electroencephalograms. Clin. Neurophysiol. 117, 2102–2108 (2006).
Mercier, M. R. et al. Evaluation of cortical local field potential diffusion in stereotactic electro-encephalography recordings: a glimpse on white matter signal. Neuroimage 147, 219–232 (2017).
Acknowledgements
The authors thank the patient for participation and C.R. Holdgraf, V. Rangarajan, C.W. Hoy, J. Kam, L. Bellier, R. Helfrich, R. Jimenez, E. Gerber, A. Blenkmann, J. Lubell, and M. Pereira for fruitful discussions. The authors are also grateful to the present and former FieldTrip core developers, as well as the greater FieldTrip community, for contributing code, documentation, and expertise that have made this protocol possible. A.S. was supported by Rubicon grant 446-14-007 from NWO and Marie Sklodowska-Curie Global Fellowship 658868 from the European Union; R.v.d.M. by R01 MH095984-03S1 from the NIMH; J.-M.S. by VIDI 864-14-011 from NWO, R.T.K. by R37 NS21135 from NINDS, and R.O. by Marie Skłodowska-Curie Innovative Training Networks 641652 from the European Union.
Author information
Authors and Affiliations
Contributions
A.S., S.G., R.v.d.M., J.-M.S., and R.O. developed the protocol. G.P. contributed the algorithm for brain-shift compensation. J.J.L. provided access and guidance in the data acquisition. A.S., S.G., J.-M.S., R.T.K., and R.O. wrote the paper, and R.v.d.M., C.D., I.S., G.P., and J.J.L., provided substantial editorial revisions.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
1. Zheng, J. et al. Nat. Commun. 8, 14413 (2017): http://dx.doi.org/10.1038/ncomms14413
2. Piai et al. Proc. Natl. Acad. Sci. USA 113, 11366–11371 (2016): http://dx.doi.org/10.1073/pnas.1603312113
Supplementary information
Supplementary Text and Figures
Supplementary Figure 1
Supplementary Data 1
Example code for a start-to-end implementation of the anatomical and functional workflow for SubjectUCI29
Supplementary Data 2
Code for the automatic DICOM series search and visualization tool, search_dicomseries.m
Supplementary Data 3
Code for the automatic electrode-labeling tool, generate_electable.m
Supplementary Video 1
Tutorial video showing the first stage of preprocessing the anatomical MRI (corresponding to Step 3)
Supplementary Video 2
Tutorial video showing the second stage of preprocessing of the anatomical MRI (corresponding to Step 4)
Supplementary Video 3
Tutorial video showing the preprocessing of the anatomical CT (corresponding to Step 11)
Supplementary Video 4
Tutorial video showing the placement of electrodes in the MRI-fused anatomical CT (corresponding to Step 17)
Supplementary Video 5
Tutorial video showing the interactive manipulation of anatomically informed graphical representations of time–frequency resolved neural data
Supplementary Video 6
Spatiotemporal dynamics of task-modulated high-frequency-band activity at surface electrodes overlaid on left parietal and temporal cortex. It can be observed that processing occurs in the temporal lobe at hearing the target tone followed by the sensorimotor system contralateral to the hand used for the button press. Warm and cold colors represent increases and decreases in high-frequency-band power, respectively
Supplementary Video 7
Spatiotemporal dynamics of epileptiform activity recorded from depth electrodes targeting the bilateral hippocampus and amygdala. It can be observed that the (interictal) epileptiform discharges first occur in the left hippocampus and amygdala and then spread to their right-hemisphere homologs during this particular episode. Warm and cold colors represent positive and negative deflections in raw signal amplitude, respectively. The size of each point cloud is scaled according to signal amplitude. This video is created using data obtained from a different subject
Rights and permissions
About this article
Cite this article
Stolk, A., Griffin, S., van der Meij, R. et al. Integrated analysis of anatomical and electrophysiological human intracranial data. Nat Protoc 13, 1699–1723 (2018). https://doi.org/10.1038/s41596-018-0009-6
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-018-0009-6
This article is cited by
-
Ramping dynamics and theta oscillations reflect dissociable signatures during rule-guided human behavior
Nature Communications (2024)
-
Electrocorticographic Activation Patterns of Electroencephalographic Microstates
Brain Topography (2024)
-
Asymmetric coding of reward prediction errors in human insula and dorsomedial prefrontal cortex
Nature Communications (2023)
-
Internal states as a source of subject-dependent movement variability are represented by large-scale brain networks
Nature Communications (2023)
-
Functional specialization and interaction in the amygdala-hippocampus circuit during working memory processing
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.