Abstract
The glucocorticoid receptor (GR) is a ligand-activated transcription factor that binds DNA and assembles co-regulator complexes to regulate gene transcription. GR agonists are widely prescribed to people with inflammatory and autoimmune diseases. Here we present high-resolution, multidomain structures of GR in complex with ligand, DNA and co-regulator peptide. The structures reveal how the receptor forms an asymmetric dimer on the DNA and provide a detailed view of the domain interactions within and across the two monomers. Hydrogen–deuterium exchange and DNA-binding experiments demonstrate that ligand-dependent structural changes are communicated across the different domains in the full-length receptor. This study demonstrates how GR forms a distinct architecture on DNA and how signal transmission can be modulated by the ligand pharmacophore, provides a platform to build a new level of understanding of how receptor modifications can drive disease progression and offers key insight for future drug design.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The X-ray data and coordinates for the GRΔN–vel–SGK1–PGC1α, GRLBD–vel–PGC1α and GRΔN–FF–SGK1–PGC1α structures are deposited in the PDB (7PRW, 7PRX and 7PRV, respectively). The PDB 3G9O of a GR DBD dimer bound to DNA was used for structural comparison. The degree of sequence conservation of the ER LBD dimer interface was blotted on the PDB structure 3ERD.
All data generated and/or analyzed in the current study are included in this published article (and its supplementary information files). Source data are provided with this paper.
References
Eick, G. N. & Thornton, J. W. Evolution of steroid receptors from an estrogen-sensitive ancestral receptor. Mol. Cell. Endocrinol. 334, 31–38 (2011).
Weikum, E. R., Knuesel, M. T., Ortlund, E. A. & Yamamoto, K. R. Glucocorticoid receptor control of transcription: precision and plasticity via allostery. Nat. Rev. Mol. Cell Biol. 18, 159 (2017).
Köhler, C. et al. Dynamic allosteric communication pathway directing differential activation of the glucocorticoid receptor. Sci. Adv. 6, eabb5277 (2020).
Watson, L. C. et al. The glucocorticoid receptor dimer interface allosterically transmits sequence-specific DNA signals. Nat. Struct. Mol. Biol. 20, 876 (2013).
Zhang, J. et al. DNA binding alters coactivator interaction surfaces of the intact VDR–RXR complex. Nat. Struct. Mol. Biol. 18, 556–563 (2011).
Meijsing, S. H. et al. DNA binding site sequence directs glucocorticoid receptor structure and activity. Science 324, 407–410 (2009).
Schiller, B. J., Chodankar, R., Watson, L. C., Stallcup, M. R. & Yamamoto, K. R. Glucocorticoid receptor binds half sites as a monomer and regulates specific target genes. Genome Biol. 15, 418 (2014).
Hudson, W. H., Youn, C. & Ortlund, E. A. The structural basis of direct glucocorticoid-mediated transrepression. Nat. Struct. Mol. Biol. 20, 53 (2012).
Presman, D. M. et al. DNA binding triggers tetramerization of the glucocorticoid receptor in live cells. Proc. Natl Acad. Sci. USA 113, 8236–8241 (2016).
Presman, D. M. et al. Live cell imaging unveils multiple domain requirements for in vivo dimerization of the glucocorticoid receptor. PLoS Biol. 12, e1001813 (2014).
Johnson, T. A., Paakinaho, V., Kim, S., Hager, G. L. & Presman, D. M. Genome-wide binding potential and regulatory activity of the glucocorticoid receptor’s monomeric and dimeric forms. Nat. Commun. 12, 1987 (2021).
Chandra, V. et al. Structure of the intact PPAR-γ–RXR-α nuclear receptor complex on DNA. Nature 456, 350 (2008).
Chandra, V. et al. Multidomain integration in the structure of the HNF-4α nuclear receptor complex. Nature 495, 394 (2013).
Chandra, V. et al. The quaternary architecture of RARβ–RXRα heterodimer facilitates domain–domain signal transmission. Nat. Commun. 8, 868 (2017).
Lou, X. et al. Structure of the retinoid X receptor α–liver X receptor β (RXRα–LXRβ) heterodimer on DNA. Nat. Struct. Mol. Biol. 21, 277–281 (2014).
Veleiro, A. S., Alvarez, L. D., Eduardo, S. L. & Burton, G. Structure of the glucocorticoid receptor, a flexible protein that can adapt to different ligands. ChemMedChem 5, 649–659 (2010).
Hemmerling, M. et al. Selective nonsteroidal glucocorticoid receptor modulators for the inhaled treatment of pulmonary diseases. J. Med. Chem. 60, 8591–8605 (2017).
Biggadike, K. et al. X-ray crystal structure of the novel enhanced-affinity glucocorticoid agonist fluticasone furoate in the glucocorticoid receptor−ligand binding domain. J. Med. Chem. 51, 3349–3352 (2008).
Brown, M. N. et al. Efficacy and safety of AZD7594, an inhaled non-steroidal selective glucocorticoid receptor modulator, in patients with asthma: a phase 2a randomized, double blind, placebo-controlled crossover trial. Respiratory Res. 20, 37 (2019).
Shefrin, A. E. & Goldman, R. D. Use of dexamethasone and prednisone in acute asthma exacerbations in pediatric patients. Can. Fam. Physician 55, 704–706 (2009).
Syed, Y. Y. Fluticasone furoate/vilanterol: a review of its use in patients with asthma. Drugs 75, 407–418 (2015).
Grasso, E. M., Majumdar, A., Wrabl, J. O., Frueh, D. P. & Hilser, V. J. Conserved allosteric ensembles in disordered proteins using TROSY/anti-TROSY R2-filtered spectroscopy. Biophys. J. 120, 2498–2510 (2021).
Shiau, A. K. et al. The structural basis of estrogen receptor/coactivator recognition and the antagonism of this interaction by tamoxifen. Cell 95, 927–937 (1998).
Bledsoe, R. K. et al. Crystal structure of the glucocorticoid receptor ligand binding domain reveals a novel mode of receptor dimerization and coactivator recognition. Cell 110, 93–105 (2002).
Bianchetti, L. et al. Alternative dimerization interfaces in the glucocorticoid receptor-α ligand binding domain. Biochim. Biophys. Acta 1862, 1810–1825 (2018).
Paakinaho, V., Johnson, T. A., Presman, D. M. & Hager, G. L. Glucocorticoid receptor quaternary structure drives chromatin occupancy and transcriptional outcome. Genome Res. 29, 1223–1234 (2019).
Presman, D. M. & Hager, G. L. More than meets the dimer: what is the quaternary structure of the glucocorticoid receptor? Transcription 8, 32–39 (2017).
Kim, D. N., Jacobs, T. M. & Kuhlman, B. Boosting protein stability with the computational design of β-sheet surfaces. Protein Sci. 25, 702–710 (2016).
Timmermans, S. et al. Point mutation I634A in the glucocorticoid receptor causes embryonic lethality by reduced ligand binding. J. Biol. Chem. 298, 101574 (2022).
Robblee, J. P., Miura, M. T. & Bain, D. L. Glucocorticoid receptor–promoter interactions: energetic dissection suggests a framework for the specificity of steroid receptor-mediated gene regulation. Biochemistry 51, 4463–4472 (2012).
Hochberg, G. K. A. et al. A hydrophobic ratchet entrenches molecular complexes. Nature 588, 503–508 (2020).
Tamrazi, A., Carlson, K. E., Daniels, J. R., Hurth, K. M. & Katzenellenbogen, J. A. Estrogen receptor dimerization: ligand binding regulates dimer affinity and dimer dissociation rate. Mol. Endocrinol. 16, 2706–2719 (2002).
Maletta, M. et al. The palindromic DNA-bound USP/EcR nuclear receptor adopts an asymmetric organization with allosteric domain positioning. Nat. Commun. 5, 4139 (2014).
Orlov, I., Rochel, N., Moras, D. & Klaholz, B. P. Structure of the full human RXR/VDR nuclear receptor heterodimer complex with its DR3 target DNA. EMBO J. 31, 291–300 (2012).
Bourguet, W. et al. Crystal structure of a heterodimeric complex of RAR and RXR ligand-binding domains. Mol. Cell 5, 289–298 (2000).
Duda, K., Chi, Y.-I. & Shoelson, S. E. Structural basis for HNF-4α activation by ligand and coactivator binding. J. Biol. Chem. 279, 23311–23316 (2004).
Gampe, R. T. et al. Asymmetry in the PPARγ/RXRα crystal structure reveals the molecular basis of heterodimerization among nuclear receptors. Mol. Cell 5, 545–555 (2000).
Svensson, S. et al. Crystal structure of the heterodimeric complex of LXRα and RXRβ ligand-binding domains in a fully agonistic conformation. EMBO J. 22, 4625–4633 (2003).
Yi, P. et al. Structure of a biologically active estrogen receptor–coactivator complex on DNA. Mol. Cell 57, 1047–1058 (2015).
Wasmuth, E. V. et al. Allosteric interactions prime androgen receptor dimerization and activation. Mol. Cell 82, 2021–2031(2022).
Yu, X. et al. Structural insights of transcriptionally active, full-length androgen receptor coactivator complexes. Mol. Cell 79, 812–823.e814 (2020).
Hegelund Myrbäck, T. et al. Effects of a selective glucocorticoid receptor modulator (AZD9567) versus prednisolone in healthy volunteers: two phase 1, single-blind, randomised controlled trials. Lancet Rheumatol. 2, e31–e41 (2020).
Ripa, L. et al. Discovery of a novel oral glucocorticoid receptor modulator (AZD9567) with improved side effect profile. J. Med. Chem. 61, 1785–1799 (2018).
Liu, X. et al. Disruption of a key ligand-H-bond network drives dissociative properties in vamorolone for Duchenne muscular dystrophy treatment. Proc. Natl Acad. Sci. USA 117, 24285–24293 (2020).
Edman, K. et al. Ligand binding mechanism in steroid receptors: from conserved plasticity to differential evolutionary constraints. Structure 23, 2280–2290 (2015).
Weis, D. D. (ed) Hydrogen Exchange Mass Spectrometry of Proteins: Fundamentals, Methods, and Applications (John Wiley & Sons, Ltd, 2016).
Heck, S. et al. A distinct modulating domain in glucocorticoid receptor monomers in the repression of activity of the transcription factor AP-1. EMBO J. 13, 4087–4095 (1994).
Hurley, D. M. et al. Point mutation causing a single amino acid substitution in the hormone binding domain of the glucocorticoid receptor in familial glucocorticoid resistance. J. Clin. Investig. 87, 680–686 (1991).
Benedek, T. G. History of the development of corticosteroid therapy. Clin. Exp. Rheumatol. 29, S-5-12 (2011).
Gehring, U. & Hotz, A. Photoaffinity labeling and partial proteolysis of wild-type and variant glucocorticoid receptors. Biochemistry 22, 4013–4018 (1983).
Simons, S. S. Jr & Thompson, E. B. Dexamethasone 21-mesylate: an affinity label of glucocorticoid receptors from rat hepatoma tissue culture cells. Proc. Natl Acad. Sci. USA 78, 3541–3545 (1981).
Hollenberg, S. M. et al. Primary structure and expression of a functional human glucocorticoid receptor cDNA. Nature 318, 635–641 (1985).
Miesfeld, R. et al. Genetic complementation of a glucocorticoid receptor deficiency by expression of cloned receptor cDNA. Cell 46, 389–399 (1986).
Hudson, W. H. et al. Distal substitutions drive divergent DNA specificity among paralogous transcription factors through subdivision of conformational space. Proc. Natl Acad. Sci. USA 113, 326–331 (2016).
Galliher-Beckley, A. J., Williams, J. G., Collins, J. B. & Cidlowski, J. A. Glycogen synthase kinase 3β-mediated serine phosphorylation of the human glucocorticoid receptor redirects gene expression profiles. Mol. Cell. Biol. 28, 7309–7322 (2008).
Carlsson, P., Koehler, K. F. & Nilsson, L. Glucocorticoid receptor point mutation V571M facilitates coactivator and ligand binding by structural rearrangement and stabilization. Mol. Endocrinol. 19, 1960–1977 (2005).
Kauppi, B. et al. The three-dimensional structures of antagonistic and agonistic forms of the glucocorticoid receptor ligand-binding domain: RU-486 induces a transconformation that leads to active antagonism. J. Biol. Chem. 278, 22748–22754 (2003).
Vonrhein, C. et al. Data processing and analysis with the autoPROC toolbox. Acta Crystallogr. D 67, 293–302 (2011).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Bricogne, G. et al. BUSTER (Global Phasing, 2017).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Kabsch, W. A solution for the best rotation to relate two sets of vectors. Acta Crystallogr. A 32, 922–923 (1976).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Masson, G. R. et al. Recommendations for performing, interpreting and reporting hydrogen deuterium exchange mass spectrometry (HDX-MS) experiments. Nat. Methods 16, 595–602 (2019).
Gluzman, Y. SV40-transformed simian cells support the replication of early SV40 mutants. Cell 23, 175–182 (1981).
Acknowledgements
We acknowledge the European Synchrotron Radiation Facility for provision of synchrotron radiation facilities, and we would like to thank the beamline staff for assistance in using beamline ID30A-1/MASSIF-1 and ID23-1. S.P. and C.K. were supported by the AstraZeneca postdoctoral program. The authors would also like to acknowledge additional support provided by the AstraZeneca Respiratory & Immunology therapeutic area. We thank J. Steele, R. Maciewicz, N.-O. Hermansson, M. Lepistö and R. Neutze for scientific discussions. This work was supported by CNRS, Inserm, Institut National du Cancer (INCa_16099), Fondation pour la Recherche Médicale (FRM), Agence Nationale pour la Recherche (ANR) and the French Infrastructure for Integrated Structural Biology FRISBI ANR-10-INSB-05-01, Instruct-ERIC, and the French Proteomic Infrastructure ProFI ANR-10-INBS-08-03.
Author information
Authors and Affiliations
Contributions
S.P. purified the proteins and crystallized the GRΔN complexes. C.K. purified the wild-type GRLBD protein. L.W. crystallized the GRLBD. S.P. and K.E. solved the structures and wrote the manuscript with input from all authors. S.P. and C.A.J. performed HDX experiments and analyzed the data. S.P. and A.G. performed fluorescence polarization assays. S.P., M.C. and E.G. performed SEC-MALS and analyzed the data. P.J. analyzed the sequence conservation of GR and ER. B.C., D.Ö., L.F.R., S.P., K.E., I.D. and S.D. designed and cloned constructs and performed cell assays. B.B., B.P.K. and I.M.L.B. helped to conceive and conceptualize the study and to interpret structural data.
Corresponding author
Ethics declarations
Competing interests
S.P., L.W., C.A.J., A.G., E.G., B.C., C.K., M.C., D.Ö., P.J., L.F.R., I.D., S.D. and K.E. were employed by AstraZeneca at the time of the study. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks Keith Yamamoto and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Editor recognition statement (if applicable to your journal): Primary Handling Editors: Florian Ullrich, Katarzyna Ciazynska and Carolina Perdigoto, in collaboration with the Nature Structural & Molecular Biology team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
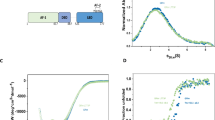
Extended Data Fig. 1 SEC-MALS of GRΔN-vel, GRΔN-FF, GRΔN-vel-SGK1-PGC1α, GRΔN-FF-SGK1-PGC1α, GRΔN-vel-SGK1-half1-PGC1α, GRΔN-vel)-SGK1-half2-PGC1α and GR-vel.
a, The monomeric GR proteins and GR complexes eluted as single peaks. b, The experimentally determined and expected molecular weights. c, GRΔN proteins and GRΔN-SGK1-PGC1α complexes separated on a native PAGE. The experiments were repeated at least three times.
Extended Data Fig. 2 The GRΔN-vel-SGK1-PGC1α crystal lattice.
The crystal lattice a, orthogonal to the DNA and b, turned by 90 degrees. c The crystal contact mediated by H1.
Extended Data Fig. 3 GRΔN-vel-SGK1-PGC1α domains overlaid on structures of the isolated domains.
a, DBD1 (purple), DBD2 (green) and dsDNA (red) overlaid on the structure of the DBD dimer alone on the same GBS (PDB: 3G9O, all in white). Zn atoms are denoted as gray and white spheres, respectively. b, LBD1 (blue) with velsecorat (magenta) and coactivator peptide PGC1α134–154 (yellow) overlaid on the structure of GRLBD (white) in complex with velsecorat (black) and coactivator peptide PGC1α134–154 (black). c, LBD2 (green) with velsecorat (magenta) overlayed on the structure of GRLBD (white) in complex with velsecorat (black) and coactivator peptide PGC1α134–154 (black).
Extended Data Fig. 4 Position of I628 in the GRΔN-vel-SGK1-PGC1α complex.
The location of the residue I628 in the quaternary complex is shown a, in LBD1 and b, in LBD2. GR LBD helix numbering is annotated within the circles.
Extended Data Fig. 5 Sequence conservation of the GR LBD.
Alignment of a set of diverse GR related vertebrate sequences with a pairwise identity of 37–84%. The mean pairwise column identity of each residue as calculated by Geneious Prime is shown as bars in the top graph. The residues involved in the LBD:LBD and LBD:DBD interfaces are indicated by orange and green boxes, respectively.
Extended Data Fig. 6 Sequence conservation of the ER LBD.
Alignment of a set of diverse ER related vertebrate sequences with a pairwise identity of 38–90%. The mean pairwise column identity of each residue as calculated by Geneious Prime is shown as bars in the top graph. LBD dimer interface residues are highlighted with orange boxes.
Extended Data Fig. 7 Sequence conservation of key interfaces in the GRΔNvel-SGK1-PGC1α complex.
Conservation of GR LBD1 residues colored according to column identity between 0.4 (white) and 1.0 (blue) and shown as a surface highlighting the LBD1 interface with a, the DBD and DNA and b, the PGC1α peptide (yellow).
Extended Data Fig. 8 Fluticasone furoate rearranges the region where H6 and H7 meet.
a, Overlay of the GRΔN-FF-SGK1-PGC1α LBD1 (white) with PGC1α peptide in black on the GRΔN-vel-SGK1-PGC1α LBD1 (blue) with PGC1α peptide in yellow. Fluticasone furoate and velsecorat are shown in black and magenta, respectively. b, Fluticasone furoate repositions Q642 and pushes on D638 and M639, rearranging the H6-H7 loop. Helix numbering is annotated within the circles.
Extended Data Fig. 9 Relative deuteration of specific peptides in GR-vel-SGK1-PGC1α, GR-FF-SGK1-PGC1α, GR-dex-SGK1-PGC1α, GR-vel, GR-FF and GR-dex complexes and protein coverage of GR and GR-SGK1-PGC1α in HDX-MS.
a, peptide 425–436. b, peptide 536–544. c, peptide 628–636. Data points are mean +/− SD of 3 independent replicates. d, Peptides used for HDX-MS analysis of GR. A coverage of 89.2% of the sequence was achieved. e, Peptides used for HDX-MS analysis of GR-SGK-PGC1α complexes. A coverage of 61.8% of the sequence was achieved.
Extended Data Fig. 10 HDX difference plots showing deuterium uptake difference between (complex A) – (complex B) for all peptides.
Negative value indicate that peptides are protected in A relative to B and vice versa.
Supplementary information
Supplementary Information
Supplementary Tables 1–6 and Validation of Cos7 cells including mycoplasma report and barcoding form ECACC
Source data
Source Data Fig. 1
Data table
Source Data Fig. 5
Data table
Source Data Fig. 6
Data table
Source Data Fig. 6
Unprocessed western blot images
Source Data Extended Data Fig. 1
Data tables
Source Data Extended Data Fig. 9
Data tables
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Postel, S., Wissler, L., Johansson, C.A. et al. Quaternary glucocorticoid receptor structure highlights allosteric interdomain communication. Nat Struct Mol Biol 30, 286–295 (2023). https://doi.org/10.1038/s41594-022-00914-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-022-00914-4