Abstract
Excitatory signaling mediated by N-methyl-d-aspartate receptor (NMDAR) is critical for brain development and function, as well as for neurological diseases and disorders. Channel blockers of NMDARs are of medical interest owing to their potential for treating depression, Alzheimer’s disease, and epilepsy. However, precise mechanisms underlying binding and channel blockade have remained limited owing to challenges in obtaining high-resolution structures at the binding site within the transmembrane domains. Here, we monitor the binding of three clinically important channel blockers: phencyclidine, ketamine, and memantine in GluN1-2B NMDARs at local resolutions of 2.5–3.5 Å around the binding site using single-particle electron cryo-microscopy, molecular dynamics simulations, and electrophysiology. The channel blockers form different extents of interactions with the pore-lining residues, which control mostly off-speeds but not on-speeds. Our comparative analyses of the three unique NMDAR channel blockers provide a blueprint for developing therapeutic compounds with minimal side effects.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Cryo-EM density maps for agonist-, (S)-(+)-ketamine-, PCP-, and memantine-bound GluN1a-2B NMDARs have been deposited in the Electron Microscopy Data Bank (EMDB) under accession codes EMD-24946, EMD-24947, EMD-24948, and EMD-24949, respectively, and atomic coordinates have been deposited in the Protein Data Bank (PDB) accession codes 7SAA, 7SAB, 7SAC, and 7SAD, respectively. The MD simulations data have been deposited to Zenodo (https://zenodo.org/record/6380196#.YkNwjSUpCHs). Source data are provided with this paper.
References
Mayer, M. L., Westbrook, G. L. & Guthrie, P. B. Voltage-dependent block by Mg2+ of NMDA responses in spinal cord neurones. Nature 309, 261–263 (1984).
Nowak, L., Bregestovski, P., Ascher, P., Herbet, A. & Prochiantz, A. Magnesium gates glutamate-activated channels in mouse central neurones. Nature 307, 462–465 (1984).
Mayer, M. L., MacDermott, A. B., Westbrook, G. L., Smith, S. J. & Barker, J. L. Agonist- and voltage-gated calcium entry in cultured mouse spinal cord neurons under voltage clamp measured using arsenazo III. J. Neurosci. 7, 3230–3244 (1987).
Nicoll, R. A. & Roche, K. W. Long-term potentiation: peeling the onion. Neuropharmacology 74, 18–22 (2013).
Nabavi, S. et al. Engineering a memory with LTD and LTP. Nature 511, 348–352 (2014).
Hansen, K. B. et al. Structure, function, and allosteric modulation of NMDA receptors. J. Gen. Physiol. 150, 1081–1105 (2018).
Traynelis, S. F. et al. Glutamate receptor ion channels: structure, regulation, and function. Pharmacol. Rev. 62, 405–496 (2010).
Luby, E. D., Cohen, B. D., Rosenbaum, G., Gottlieb, J. S. & Kelley, R. Study of a new schizophrenomimetic drug—sernyl. AMA Arch. Neurol. Psychiatry 81, 363–369 (1959).
Kornhuber, J. & Weller, M. Psychotogenicity and N-methyl-d-aspartate receptor antagonism: implications for neuroprotective pharmacotherapy. Biol. Psychiatry 41, 135–144 (1997).
Anis, N. A., Berry, S. C., Burton, N. R. & Lodge, D. The dissociative anaesthetics, ketamine and phencyclidine, selectively reduce excitation of central mammalian neurones by N-methyl-aspartate. Br. J. Pharmacol. 79, 565–575 (1983).
MacDonald, J. F., Miljkovic, Z. & Pennefather, P. Use-dependent block of excitatory amino acid currents in cultured neurons by ketamine. J. Neurophysiol. 58, 251–266 (1987).
Wilkinson, D. A review of the effects of memantine on clinical progression in Alzheimer’s disease. Int. J. Geriatr. Psychiatry 27, 769–776 (2012).
Li, L. & Vlisides, P. E. Ketamine: 50 years of modulating the mind. Front. Hum. Neurosci. 10,https://doi.org/10.3389/fnhum.2016.00612 (2016).
Lodge, D. & Mercier, M. S. Ketamine and phencyclidine: the good, the bad and the unexpected. Br. J. Pharmacol. 172, 4254–4276 (2015).
Glasgow, N. G. & Johnson, J. W. in Patch–Clamp Methods and Protocols (eds Martina, M. & Taverna, S.) 23-41 (Springer New York, 2014).
Karakas, E. & Furukawa, H. Crystal structure of a heterotetrameric NMDA receptor ion channel. Science 344, 992–997 (2014).
Lee, C. H. et al. NMDA receptor structures reveal subunit arrangement and pore architecture. Nature 511, 191–197 (2014).
Tajima, N. et al. Activation of NMDA receptors and the mechanism of inhibition by ifenprodil. Nature 534, 63–68 (2016).
Zhu, S. et al. Mechanism of NMDA receptor inhibition and activation. Cell 165, 704–714 (2016).
Regan, M. C. et al. Structural mechanism of functional modulation by gene splicing in NMDA receptors. Neuron 98, 521–529 (2018).
Chou, T. H., Tajima, N., Romero-Hernandez, A. & Furukawa, H. Structural basis of functional transitions in mammalian NMDA receptors. Cell 182, 357–371 (2020).
Jalali-Yazdi, F., Chowdhury, S., Yoshioka, C. & Gouaux, E. Mechanisms for zinc and proton inhibition of the GluN1/GluN2A NMDA receptor. Cell 175, 1520–1532.e1515 (2018).
Regan, M. C., Romero-Hernandez, A. & Furukawa, H. A structural biology perspective on NMDA receptor pharmacology and function. Curr. Opin. Struct. Biol. 33, 68–75 (2015).
Karakas, E., Regan, M. C. & Furukawa, H. Emerging structural insights into the function of ionotropic glutamate receptors. Trends Biochem. Sci. 40, 328–337 (2015).
Wang, J. X. & Furukawa, H. Dissecting diverse functions of NMDA receptors by structural biology. Curr. Opin. Struct. Biol. 54, 34–42 (2019).
Tajima, N. et al. Development and characterization of functional antibodies targeting NMDA receptors. Nat. Commun. 13, 923 (2022).
Song, X. et al. Mechanism of NMDA receptor channel block by MK-801 and memantine. Nature 556, 515–519 (2018).
Lu, W., Du, J., Goehring, A. & Gouaux, E. Cryo-EM structures of the triheteromeric NMDA receptor and its allosteric modulation. Science 355, eaal3729 (2017).
Zhang, Y. et al. Structural basis of ketamine action on human NMDA receptors. Nature 596, 301–305 (2021).
Furukawa, H., Simorowski, N. & Michalski, K. Effective production of oligomeric membrane proteins by EarlyBac-insect cell system. Methods Enzymol. 653, 3–19 (2021).
Jespersen, A., Tajima, N., Fernandez-Cuervo, G., Garnier-Amblard, E. C. & Furukawa, H. Structural insights into competitive antagonism in NMDA receptors. Neuron 81, 366–378 (2014).
Furukawa, H., Singh, S. K., Mancusso, R. & Gouaux, E. Subunit arrangement and function in NMDA receptors. Nature 438, 185–192 (2005).
Karakas, E., Simorowski, N. & Furukawa, H. Subunit arrangement and phenylethanolamine binding in GluN1/GluN2B NMDA receptors. Nature 475, 249–253 (2011).
Regan, M. C. et al. Structural elements of a pH-sensitive inhibitor binding site in NMDA receptors. Nat. Commun. 10, 321 (2019).
Jones, K. S., VanDongen, H. M. & VanDongen, A. M. The NMDA receptor M3 segment is a conserved transduction element coupling ligand binding to channel opening. J. Neurosci. 22, 2044–2053 (2002).
Parsons, C. G. & Gilling, K. Memantine as an example of a fast, voltage-dependent, open channel N-methyl-d-aspartate receptor blocker. Methods Mol. Biol. 403, 15–36 (2007).
Liu, K. & Kokubo, H. Exploring the stability of ligand binding modes to proteins by molecular dynamics simulations: a cross-docking study. J. Chem. Inf. Model. 57, 2514–2522 (2017).
Kashiwagi, K. et al. Channel blockers acting at N-methyl-d-aspartate receptors: differential effects of mutations in the vestibule and ion channel pore. Mol. Pharmacol. 61, 533–545 (2002).
Blanpied, T. A., Clarke, R. J. & Johnson, J. W. Amantadine inhibits NMDA receptors by accelerating channel closure during channel block. J. Neurosci. 25, 3312–3322 (2005).
Glasgow, N. G., Povysheva, N. V., Azofeifa, A. M. & Johnson, J. W. Memantine and ketamine differentially alter NMDA receptor desensitization. J. Neurosci. 37, 9686–9704 (2017).
Tegunov, D. & Cramer, P. Real-time cryo-electron microscopy data preprocessing with Warp. Nat. Methods 16, 1146–1152 (2019).
Zivanov, J., Nakane, T. & Scheres, S. H. W. A Bayesian approach to beam-induced motion correction in cryo-EM single-particle analysis. IUCrJ 6, 5–17 (2019).
Grant, T., Rohou, A. & Grigorieff, N. cisTEM, user-friendly software for single-particle image processing. eLife 7, e35383 (2018).
Pettersen, E. F. et al. UCSF Chimera — a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr D. Biol. Crystallogr 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr D. Biol. Crystallogr 66, 213–221 (2010).
Webb, B. & Sali, A. Comparative protein structure modeling using MODELLER. Curr. Protoc. Bioinforma. 2016, 5.6.1–5.6.37 (2016).
Abraham, M. J. et al. GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1-2, 19–25 (2015).
Wang, J., Wolf, R. M., Caldwell, J. W., Kollman, P. A. & Case, D. A. Development and testing of a general Amber force field. J. Comput. Chem. 25, 1157–1174 (2004).
Wang, J., Wang, W., Kollman, P. A. & Case, D. A. Automatic atom type and bond type perception in molecular mechanical calculations. J. Mol. Graph. Modell. 25, 247–260 (2006).
Jakalian, A., Bush, B. L., Jack, D. B. & Bayly, C. I. Fast, efficient generation of high-quality atomic charges. AM1-BCC model: I. method. J. Comput. Chem. 21, 132–146 (2000).
Jämbeck, J. P. M. & Lyubartsev, A. P. Derivation and systematic validation of a refined all-atom force field for phosphatidylcholine lipids. J. Phys. Chem. B 116, 3164–3179 (2012).
Jorgensen, W. L., Maxwell, D. S. & Tirado-Rives, J. Development and testing of the OPLS all-atom force field on conformational energetics and properties of organic liquids. J. Am. Chem. Soc. 118, 11225–11236 (1996).
Lindorff-Larsen, K. et al. Improved side-chain torsion potentials for the Amber ff99SB protein force field. Proteins Struct. Funct. Bioinf. 78, 1950–1958 (2010).
Hess, B., Bekker, H., Berendsen, H. J. C. & Fraaije, J. G. E. M. LINCS: a linear constraint solver for molecular simulations. J. Comput. Chem. 18, 1463–1472 (1997).
Bussi, G., Donadio, D. & Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 126, 014101 (2007).
Evans, D. J. & Holian, B. L. The Nose–Hoover thermostat. J. Chem. Phys. 83, 4069–4074 (1985).
Parrinello, M. & Rahman, A. Polymorphic transitions in single crystals: a new molecular dynamics method. J. Appl. Phys. 52, 7182–7190 (1981).
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald: an N·log(N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089–10092 (1993).
Acknowledgements
We thank N. Simorowski for technical assistance. D. Thomas and M. Wang are thanked for managing the cryo-EM facility and the computing facility, respectively. T. Heywood is thanked for technical support in usage of HPC facility at Cold Spring Harbor Laboratory (CSHL). We thank M. Mayer for insightful comments on this study. This work was funded by the NIH (NS111745 and MH085926 to H.F.), Robertson funds at CSHL, Doug Fox Alzheimer’s fund, Austin’s purpose, Heartfelt Wing Alzheimer’s fund, and the Gertrude and Louis Feil Family Trust (all to H. F.); and NIH F32MH121061 (to K. M.). The computational work was performed with assistance from an NIH grant (S10OD028632-01). P.C.B thanks the BBSRC for support (BB/M006395/1).
Author information
Authors and Affiliations
Contributions
T.-H. C. and E. F. conducted cyro-EM experiments. M. E. conducted MD simulations. K. M. conducted electrophysiology. T.-H. C., M. E., K. M., P. C. B., and H. F. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural and Molecular Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Florian Ullrich was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Single-particle analysis on agonist-bound GluN1a-2B NMDARs.
a, A representative EM micrograph, 2D classes, and the 3D classification/refinement workflow. The scale bar represents 34.9 nm. b, The masked (Blue) and unmasked (Red) Fourier shell correlation (FSC) curves of two half maps (Left) and map vs model (Right). c, Zoom-in views of the gate and the pore-forming region fitted with the molecular model. d, The local resolution estimation calculated by ResMap.
Extended Data Fig. 2 Structural analysis of agonist-bound GluN1a-2B NMDARs.
a, Overall cryo-EM density of the agonist-bound rat GluN1a-2B NMDAR (GluN1a and GluN2B in magenta and forest, respectively). b, EM density of the bound glycine (left) and glutamate (right) at LBD of GluN1a and GluN2B, respectively. c, Zoom-in view of the channel blocker binding cavity at the TMD where there is no density (left). The binding site has three layers, Thr ring (GluN1a-Thr648 GluN2B-Thr647) at the channel gate, the hydrophobic ring (GluN1a-Val644 and GluN1a-Ala645 and GluN2B-Leu643 and GluN2B-Ala644), and the Asn ring (GluN1a-Asn616 and GluN2B-Asn615). d, Domain organization and functional state. The agonist-bound GluN1a-2B NMDAR here has similarity to the non-active1 conformation of GluN1b-2B NMDAR (gray, PDBID: 6WHS) with the similar ATD and LBD orientations and extent of separation between the LBD-TMD linkers of GluN2B (spheres = GluN2B-Gln662).
Extended Data Fig. 3 Single-particle analysis on PCP-bound GluN1a-2B NMDARs.
a, A representative EM micrograph, 2D classes, and the 3D classification/refinement workflow. The scale bar represents 34.9 nm. b, The masked (Blue) and unmasked (Red) Fourier shell correlation (FSC) curves of two half maps (Left) and map vs model (Right). c, Zoom-in views of the gate and the pore-forming region fitted with the molecular model. d, The local resolution estimation calculated by ResMap.
Extended Data Fig. 4 Comparison of channel blocker binding sites.
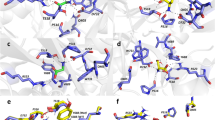
a-d, Cryo-EM density and fitted models of structures in the absence of a channel blocker (a, apo), PCP (b, orange), S(+)-ketamine (c, cyan), and memantine (d, grey). The residues that are forming the binding site are highlighted as sticks and the densities of the channel blockers are highlighted in green. e-g, Local structural comparison of binding residues between apo and PCP (e), S-(+)-ketamine (f), and memantine (g).
Extended Data Fig. 5 Single-particle analysis on S-(+)-ketamine-bound GluN1a-2B NMDARs.
a, A representative EM micrograph, 2D classes, and the 3D classification/refinement workflow. The scale bar represents 34.9 nm. b, The masked (Blue) and unmasked (Red) Fourier shell correlation (FSC) curves of two half maps (Left) and map vs model (Right). c, Zoom-in views of the gate and the pore-forming region fitted with the molecular model. d, The local resolution estimation calculated by ResMap.
Extended Data Fig. 6 Cryo-EM density at the (S)-(+)-ketamine binding site.
a, Zoom-in views of the cryo-EM density (mesh) fitted with the (S)-(+)-ketamine molecule (cyan sticks) in Pose-3. b, Cryo-EM density of human GluN1a-2A NMDARs (PDBID:7EU7 and EMD:31308, upper panel) complexed to (S)-(+)-ketamine (salmon sticks) and human GluN1a-GluN2B NMDARs (PDBID:7EU8 and EMD:31309, lower panel) complexed with (S)-(+)-ketamine (orange sticks).
Extended Data Fig. 7 Long time-scale MD simulations of (S)-(+)-ketamine-bound GluN1a-2B NMDAR starting from Pose-1.
a, RMSD population density distributions (80 bins per distribution) for three individual Pose-1 1 µs long simulations with runs 1, 2 and 3 each colored blue, orange and green respectively. b, Time series analysis of the (S)-(+)-ketamine Z1 distance expressed as a rolling average (100 frames with a total of 10,000 frames per run) for runs 1 (blue), 2 (orange) and 3 (green). c, Probability density distribution (100 bins) for each independent run of their Z1 (red) and Z2 (green) distances. Dashed lines represent the starting Z1 and Z2 distances of Pose-1.
Extended Data Fig. 8 Single-particle analysis on memantine-bound GluN1a-2B NMDARs.
a, A representative EM micrograph, 2D classes, and the 3D classification/refinement workflow. The scale bar represents 34.9 nm. b, The masked (Blue) and unmasked (Red) Fourier shell correlation (FSC) curves of two half maps (Left) and map vs model (Right). c, Zoom-in views of the gate and the pore-forming region fitted with the molecular model. d, The local resolution estimation calculated by ResMap.
Extended Data Fig. 9 Potency measurement of channel blockers on GluN1a-2B NMDAR.
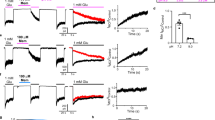
a-d, Dose-responses of PCP (panel a), S-(+)-ketamine (panel b), memantine (panel c), and Mg2+ (panel d) for wildtype (black circle), GluN1a-V644A (red circle), GluN2B-N615Q (blue triangle), GluN2B-L645A (blue square), and GluN2B-T646S (blue circle) channels recorded on cRNA injected Xenopus oocytes by TEVC. Data points represent the means ± s.d. e, A list of IC50 values, Hill coefficients (nH) calculated based on the dose-response curves. The statistical analysis was done by two-tail Student t-test (*** p < 0.001, ** 0.001 < p < 0.01, * 0.01 < p < 0.05, n.s. not significant. The IC50 values were estimated from independent dose-response recordings from at least four independent oocytes (n).
Extended Data Fig. 10 Hydrogen-bond network in the Asn ring and extent of hydrophobic interactions.
a,b, Probability density distributions (100 bins) of h-bond distances between all feasible pairs of hydrogen bond doner and acceptor moieties in the Asn ring for the agonist-bound (panel a) and agonist-memantine-bound (panel b) structures. Density distributions for GluN2B-Asn615 (chain B and D) are in green and red. Those for GluN1a-Asn616 (chain A)/GluN2B-Asn615 (chain B and D) and GluN1a-Asn616 (chain C)/GluN2B-Asn615 (chain B and D) are in yellow and blue, respectively. Asterisks denote the interaction pairs for which h-bonds are the main interaction, as suggested by the highest peak at ~2.5 Å. The structures are representative snapshots of collective variables that maximize the total number of possible simultaneous h-bonding interactions. The Asn rings are viewed from the extracellular space where representative h-bonds are depicted as dashed yellow lines. The nitrogen atom h-bond doner variable is represented as the COG between both N+-H atoms. Note that the h-bond network patterns for (S)-(+)-ketamine and PCP are similar to the agonist-bound structure. c, Comparison of binding of PCP pose-1 (orange sticks), memantine (gray sticks), and S-(+)-ketamine pose-2, 3, and 4 (cyan sticks) showing inter-carbon (C-C) interactions within 5 Å (yellow dots) and a hydrogen bond (green dots, arrow).
Supplementary information
Supplementary Information
Supplementary Table 1
Source Data
MD simulation coordinates for (S)-(+)-ketamine (ket_p1.pdb, ket_p2.pdb, ket_p3.pdb, and ket_p4.pdb), PCP (pcp_p1.pdb, and pcp_p2.pdb), and memantine (mematine.pdb)
Source data
Rights and permissions
About this article
Cite this article
Chou, TH., Epstein, M., Michalski, K. et al. Structural insights into binding of therapeutic channel blockers in NMDA receptors. Nat Struct Mol Biol 29, 507–518 (2022). https://doi.org/10.1038/s41594-022-00772-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-022-00772-0
This article is cited by
-
Therapeutic potential of N-methyl-D-aspartate receptor modulators in psychiatry
Neuropsychopharmacology (2024)
-
De novo GRIN variants in M3 helix associated with neurological disorders control channel gating of NMDA receptor
Cellular and Molecular Life Sciences (2024)