Abstract
Transient receptor potential vanilloid subfamily member 3 (TRPV3) is a temperature-sensitive cation channel. Previous cryo-EM analyses of TRPV3 in detergent micelles or amphipol proposed that the lower gate opens by α-to-π helical transitions of the nearby S6 helix. However, it remains unclear how physiological lipids are involved in the TRPV3 activation. Here we determined the apo state structure of mouse (Mus musculus) TRPV3 in a lipid nanodisc at 3.3 Å resolution. The structure revealed that lipids bound to the pore domain stabilize the selectivity filter in the narrow state, suggesting that the selectivity filter of TRPV3 affects cation permeation. When the lower gate is closed in nanodisc-reconstituted TRPV3, the S6 helix adopts the π-helical conformation without agonist- or heat-sensitization, potentially stabilized by putative intra-subunit hydrogen bonds and lipid binding. Our findings provide insights into the lipid-associated gating mechanism of TRPV3.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Cryo-EM density maps have been deposited in the Electron Microscopy Data Bank (EMDB), under accession number EMD-0882. Model coordinates have been deposited in the Protein Data Bank (wwPDB), under accession code PDB 6LGP. The associated electron microscopy data have been deposited in the Electron Microscopy Public Image Archive, under accession code EMPIAR-10400. Source data for Fig. 4e and Extended Data Fig. 1a are available with the paper online.
References
Ramsey, I. S., Delling, M. & Clapham, D. E. An introduction to Trp channels. Ann. Rev. Physiol. 68, 619–647 (2005).
Smith, G. D. et al. TRPV3 is a temperature-sensitive vanilloid receptor-like protein. Nature 418, 186–190 (2002).
Xu, H. et al. TRPV3 is a calcium-permeable temperature-sensitive cation channel. Nature 418, 181–186 (2002).
Sherkheli, M. A., Vogt-Eisele, A. K., Weber, K. & Hatt, H. Camphor modulates TRPV3 cation channels activity by interacting with critical pore-region cysteine residues. Pak. J. Pharm. Sci. 26, 431–438 (2013).
Chung, M.-K. 2-Aminoethoxydiphenyl borate activates and sensitizes the heat-gated ion channel TRPV3. J. Neurosci. 24, 5177–5182 (2004).
Vogt-Eisele, A. K. et al. Monoterpenoid agonists of TRPV3. Br. J. Pharmacol. 151, 530–540 (2007).
Qu, Y., Wang, G., Sun, X. & Wang, K. Inhibition of the warm-temperature activated Ca2+-permeable transient receptor potential vanilloid TRPV3 channel attenuates atopic dermatitis. Mol. Pharmacol. 96, 393–400 (2019).
Lin, Z. et al. Exome sequencing reveals mutations in TRPV3 as a cause of Olmsted syndrome. Am. J. Hum. Genet. 90, 558–564 (2012).
Atherton, D. J., Sutton, C. & Jones, B. M. Mutilating palmoplantar keratoderma with periorificial keratotic plaques (Olmsted’s syndrome). Br. J. Dermatol. 122, 245–252 (1990).
Asakawa, M. et al. Association of a mutation in TRPV3 with defective hair growth in rodents. J. Invest. Dermatol. 126, 2664–2672 (2006).
Klein, A. S., Tannert, A. & Schaefer, M. Cholesterol sensitises the transient receptor potential channel TRPV3 to lower temperatures and activator concentrations. Cell Calcium 55, 59–68 (2014).
Hu, H.-Z. et al. Potentiation of TRPV3 channel function by unsaturated fatty acids. J. Cell. Physiol. 208, 201–212 (2006).
Doerner, J. F., Hatt, H. & Ramsey, I. S. Voltage- and temperature-dependent activation of TRPV3 channels is potentiated by receptor-mediated PI(4,5)P2 hydrolysis. J. Gen. Physiol. 137, 271–288 (2011).
Gao, Y., Cao, E., Julius, D. & Cheng, Y. TRPV1 structures in nanodiscs reveal mechanisms of ligand and lipid action. Nature 534, 347–351 (2016).
Cao, E., Liao, M., Cheng, Y. & Julius, D. TRPV1 structures in distinct conformations reveal activation mechanisms. Nature 504, 113–118 (2013).
Zubcevic, L. et al. Cryo-electron microscopy structure of the TRPV2 ion channel. Nat. Struct. Mol. Biol. 23, 180–186 (2016).
Liao, M., Cao, E., Julius, D. & Cheng, Y. Structure of the TRPV1 ion channel determined by electron cryo-microscopy. Nature 504, 107–112 (2013).
Singh, A. K., McGoldrick, L. L. & Sobolevsky, A. I. Structure and gating mechanism of the transient receptor potential channel TRPV3. Nat. Struct. Mol. Biol. 25, 805–813 (2018).
Zubcevic, L. et al. Conformational ensemble of the human TRPV3 ion channel. Nat. Commun. 9, 4773 (2018).
Zubcevic, L., Borschel, W. F., Hsu, A. L., Borgnia, M. J. & Lee, S. Y. Regulatory switch at the cytoplasmic interface controls TRPV channel gating. Elife 8, e47746 (2019).
Liu, B., Yao, J., Zhu, M. X. & Qin, F. Hysteresis of gating underlines sensitization of TRPV3 channels. J. Gen. Physiol. 138, 509–520 (2011).
Singh, A. K. et al. Structural basis of temperature sensation by the TRP channel TRPV3. Nat. Struct. Mol. Biol. 26, 994–998 (2019).
Liu, B. & Qin, F. Single-residue molecular switch for high-temperature dependence of vanilloid receptor TRPV3. Proc. Natl Acad. Sci. USA 114, 1589–1594 (2017).
Grandl, J. et al. Pore region of TRPV3 ion channel is specifically required for heat activation. Nat. Neurosci. 11, 1007–1013 (2008).
Bayburt, T. H. & Sligar, S. G. Membrane protein assembly into nanodiscs. FEBS Lett. 584, 1721–1727 (2010).
Chen, Q. et al. Structure of mammalian endolysosomal TRPML1 channel in nanodiscs. Nature 550, 415–418 (2017).
Cheng, Y. et al. Structure of the human TRPM4 ion channel in a lipid nanodisc. Science 359, 228–232 (2017).
Grinkova, Y. V., Denisov, I. G. & Sligar, S. G. Engineering extended membrane scaffold proteins for self-assembly of soluble nanoscale lipid bilayers. Protein Eng. Des. Sel. 23, 843–848 (2010).
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. P. HOLE: a program for the analysis of the pore dimensions of ion channel structural models. J. Mol. Graph. 14, 354–360 (1996).
Deng, Z. et al. Cryo-EM and X-ray structures of TRPV4 reveal insight into ion permeation and gating mechanisms. Nat. Struct. Mol. Biol. 25, 252–260 (2018).
Tang, L. et al. Structural basis for Ca2+ selectivity of a voltage-gated calcium channel. Nature 505, 56–61 (2014).
Shannon, R. D. Revised effective ionic radii and systematic studies of interatomic distances in halides and chalcogenides. Acta Crystallogr. A Found. Adv. 32, 751–767 (1976).
Nightingale, E. R. Phenomenological theory of ion solvation. Effective radii of hydrated ions. J. Phys. Chem. 63, 1381–1387 (1959).
Cheng, W. et al. Heteromeric heat-sensitive transient receptor potential channels exhibit distinct temperature and chemical response. J. Biol. Chem. 287, 7279–7288 (2012).
Jara-Oseguera, A., Huffer, K. E. & Swartz, K. J. The ion selectivity filter is not an activation gate in TRPV1-3 channels. Elife 8, e51212 (2019).
Dukkipati, A., Park, H. H., Waghray, D., Fischer, S. & Garcia, K. C. BacMam system for high-level expression of recombinant soluble and membrane glycoproteins for structural studies. Protein Expr. Purif. 62, 160–170 (2008).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. Elife 7, e42166 (2018).
Zivanov, J., Nakane, T. & Scheres, S. H. W. A Bayesian approach to beam-induced motion correction in cryo-EM single-particle analysis. IUCrJ 6, 5–17 (2019).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D Struct. Biol. 74, 531–544 (2018).
Acknowledgements
We thank T. Nakane for computational support and assistance with the single-particle analysis and model building, K. Yamashita for computational support and model building and Y. Lee for fruitful discussions and encouragement. We also thank the staff scientists at the University of Tokyo’s cryo-EM facility, especially K. Kobayashi, H. Yanagisawa, A. Tsutsumi, M. Kikkawa and R. Danev. This work was supported by a MEXT Grant-in-Aid for Specially Promoted Research (grant no. 16H06294) to O.N. and by MEXT KAKENHI Grant-in-Aid for Scientific Research(C) (18K60156). This research was supported by the Platform Project for Supporting Drug Discovery and Life Science Research (Basis for Supporting Innovative Drug Discovery and Life Science Research) from AMED under grant no. JP19am01011115 (support number 1110).
Author information
Authors and Affiliations
Contributions
H.S., T.K., T.H. and O.N. designed the project. T.K. investigated the conditions for TRPV3 expression. H.S. performed protein expression, purification and cryo-EM sample preparation, with assistance from T.K. H.S., T.K. and T.N. performed data collection. H.S. processed and analyzed the cryo-EM data with assistance from T.K. T.N., H.S. and T.K. built the models. T.H.D.N. and M.T. performed the electrophysiology, protein biotinylation, western blotting and statistical analysis. H.S., T.K., T.H.D.N., T.N., T.H., M.T. and O.N. wrote the manuscript. O.N. supervised the research.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Katarzyna Marcinkiewicz was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Functional characterization and nanodisc reconstitution of truncated mutant TRPV3.
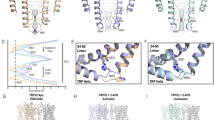
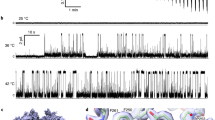
a, Temperature thresholds of mTRPV3WT (light blue) and mTRPV-ΔN (light orange), stimulated by heat, 10 mM camphor, or 300 mM 2-APB. b, c, Electrophysiological comparison between mouse TRPV3 WT (mTRPV3-WT) (b) and N-terminal truncated mutant (mTRPV3ΔN) (c), by whole-cell patch-clamp recordings in HEK293 cells. mTRPV3-ΔN has the same electrophysiological profile as mTRPV3-WT. Graphs show mean and s.e.m. for indicated number of recordings. The source data are available online. d, e, Size exclusion chromatogram (d) and SDS–PAGE (e) of mTRPV3-ΔN in a nanodisc.
Extended Data Fig. 2 Overview of single-particle cryo-EM for mTRPV3-ΔN in a nanodisc.
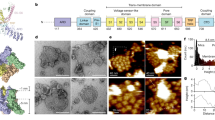
a, Cryo-EM micrograph of mTRPV3-ΔN in a nanodisc, with example particles circled in red. b, Reference-free 2D class images of mTRPV3-ΔN in a nanodisc. c, 3D class averages of mTRPV3-ΔN in a nanodisc. d, Refined density, at 3.31 Å resolution (FSC = 0.143). e, FSC curves of the map of TRPV3 in a nanodisc. The horizontal dashed line indicated FSC = 0.143. f, Euler angular distribution for particles used in the final map of TRPV3 in a nanodisc.
Extended Data Fig. 3 The local resolution mapped on the density of mTRPV3-ΔN in a nanodisc.
The map was obtained using CueMol2 at a 0.010 threshold level (http://www.cuemol.org/). The local resolution is higher in the Transmembrane core than the Intracellular region.
Extended Data Fig. 4 Cryo-EM density of mTRPV3-ΔN in a nanodisc.
a, Cryo-EM density (blue mesh) at 2.4σ for a single TRPV3 subunit; the structure, shown as a ribbon, is colored according to the domains. b–i, Fragments of the TRPV3 transmembrane region with the corresponding cryo-EM densities shown as a blue mesh at 2.4σ. j, Cross-validation FSC curves for the refined model versus unfiltered half maps (only half map 1 was used for refinement with PHENIX42) and the summed map.
Extended Data Fig. 5 Lipid-like densities observed in mTRPV3-ΔN in a nanodisc.
a, The seven lipid-like densities (1-7) observed in mTRPV3-ΔN in a nanodisc map. Five densities (1-5) are sufficient to be modeled as phosphatidylcholine. The density is shown at 3.5σ. b, The site occupied by Lipid 1, known as the Vanilloid Binding Pocket, in TRPV1 structures in nanodiscs (PDB: 5IRZ). c, d, The lipid-like densities ascribed to Lipid 2 (c) and Lipid 6 (d) are observed in the TRPV3 structure in digitonin micelles (PDB: 6DVW).
Extended Data Fig. 6 Model validation map of selectivity filter.
a, Ribbon model around the selectivity filter, in stereo. The oxygens of the Gly638 and Gly640 mainchains, and the side chains of Leu639 and Asp641 are shown in stick models. EM density map around Gly638 – Asp641 shown in a gray mesh. b, Superimposition of the pore domain of TRPV3 in a nanodisc (pink) and the TRPV3 open state (light blue) (PDB: 6DVZ), according to the Cα atoms of the S6 helices (Pro651–Ser685), viewed from the extracellular side (same perspective as Fig. 3a). c, Comparison of the phospholipid-binding site in Fig. 3c between TRPV3 in a nanodisc (pink) and the TRPV3 open state (light blue) (same perspective as Fig. 3d). Both models are superimposed on the Cα atoms of the S6 helix (Pro651–Ser685) as a guide. The phospholipid observed in a nanodisc and the residues shown in Fig. 3c in the open state are indicated by stick and CPK models, respectively, revealing that the residues clashed with Lipid 3. d-g, Known TRPV3 structural models fitted to each EM density map (shown in blue mesh). TRPV3 in the closed state in digitonin micelles (purple) (d), closed state in amphipol (yellow) (e), sensitized state (green) (f), and open state (light blue) (g).
Extended Data Fig. 7 Close-up views of Lipid 4, Lipid 5, and Lipid 7.
a, Locations of the densities observed in Lipid 4, Lipid 5, and Lipid 7, indicated as a blue mesh at 3.0σ. The modeled phosphatidylcholine (Lipid 4, Lipid 5), the partially modeled acyl chain (Lipid 7), and the residues near Lipid 4 and Lipid 7 are shown as sticks. b, Interaction between Lipid 4 and the extracellular part of the S5 helix, viewed from the extracellular side and the residues nearby Lipid 4 are shown as sticks. c, Interactions among the intracellular region of the S5 helix, Lipid 5, and Lipid 7, viewed parallel to the membrane, and the residues nearby Lipid 7 are shown as sticks. d, e, Location of the density observed for Lipid 7 in previously reported structures: TRPV3-sensitized state (PDB: 6MHS) (d), TRPV3 open state (PDB: 6DVZ) (e), and the residues nearby the densities are shown as sticks.
Supplementary information
Supplementary Information
Supplementary Fig. 1.
Source data
Source Data Fig. 4
Statistical source data of Fig. 4e
Source Data Extended Data Fig. 1
Statistical source data of Extended data Fig. 1a
Rights and permissions
About this article
Cite this article
Shimada, H., Kusakizako, T., Dung Nguyen, T.H. et al. The structure of lipid nanodisc-reconstituted TRPV3 reveals the gating mechanism. Nat Struct Mol Biol 27, 645–652 (2020). https://doi.org/10.1038/s41594-020-0439-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-020-0439-z
This article is cited by
-
Structural mechanisms of TRPV2 modulation by endogenous and exogenous ligands
Nature Chemical Biology (2023)
-
Thermoring basis for the TRPV3 bio-thermometer
Scientific Reports (2023)
-
TRP (transient receptor potential) ion channel family: structures, biological functions and therapeutic interventions for diseases
Signal Transduction and Targeted Therapy (2023)
-
Structural basis of TRPV3 inhibition by an antagonist
Nature Chemical Biology (2023)
-
Cannabinoid non-cannabidiol site modulation of TRPV2 structure and function
Nature Communications (2022)