Abstract
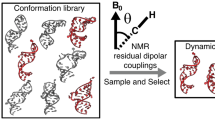
Dynamic ensembles hold great promise in advancing RNA-targeted drug discovery. Here we subjected the transactivation response element (TAR) RNA from human immunodeficiency virus type-1 to experimental high-throughput screening against ~100,000 drug-like small molecules. Results were augmented with 170 known TAR-binding molecules and used to generate sublibraries optimized for evaluating enrichment when virtually screening a dynamic ensemble of TAR determined by combining NMR spectroscopy data and molecular dynamics simulations. Ensemble-based virtual screening scores molecules with an area under the receiver operator characteristic curve of ~0.85–0.94 and with ~40–75% of all hits falling within the top 2% of scored molecules. The enrichment decreased significantly for ensembles generated from the same molecular dynamics simulations without input NMR data and for other control ensembles. The results demonstrate that experimentally determined RNA ensembles can significantly enrich libraries with true hits and that the degree of enrichment is dependent on the accuracy of the ensemble.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
06 May 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Howe, J. A. et al. Selective small-molecule inhibition of an RNA structural element. Nature 526, 672–677 (2015).
Connelly, C. M., Moon, M. H. & Schneekloth, J. S. Jr. The emerging role of RNA as a therapeutic target for small molecules. Cell Chem. Biol. 23, 1077–1090 (2016).
Shortridge, M. D. & Varani, G. Structure based approaches for targeting non-coding RNAs with small molecules. Curr. Opin. Struct. Biol. 30, 79–88 (2015).
Hermann, T. Small molecules targeting viral RNA. Wiley Interdiscip. Rev. RNA 7, 726–743 (2016).
Matsui, M. & Corey, D. R. Non-coding RNAs as drug targets. Nat. Rev. Drug Discov. 16, 167–179 (2017).
Disney, M. D., Yildirim, I. & Childs-Disney, J. L. Methods to enable the design of bioactive small molecules targeting RNA. Org. Biomol. Chem. 12, 1029–1039 (2014).
Feixas, F., Lindert, S., Sinko, W. & McCammon, J. A. Exploring the role of receptor flexibility in structure-based drug discovery. Biophys. Chem. 186, 31–45 (2014).
Amaro, R. E. & Li, W. W. Emerging methods for ensemble-based virtual screening. Curr. Top. Med. Chem. 10, 3–13 (2010).
Stelzer, A. C. et al. Discovery of selective bioactive small molecules by targeting an RNA dynamic ensemble. Nat. Chem. Biol. 7, 553–559 (2011).
Ferrari, A. M., Wei, B. Q., Costantino, L. & Shoichet, B. K. Soft docking and multiple receptor conformations in virtual screening. J. Med. Chem. 47, 5076–5084 (2004).
Cerqueira, N. M. F. S. A., Bras, N. F., Fernandes, P. A. & Ramos, M. J. MADAMM: a multistaged docking with an automated molecular modeling protocol. Proteins 74, 192–206 (2009).
Sherman, W., Day, T., Jacobson, M. P., Friesner, R. A. & Farid, R. Novel procedure for modeling ligand/receptor induced fit effects. J. Med. Chem. 49, 534–553 (2006).
Knegtel, R. M. A., Kuntz, I. D. & Oshiro, C. M. Molecular docking to ensembles of protein structures. J. Mol. Biol. 266, 424–440 (1997).
Carlson, H. A. et al. Developing a dynamic pharmacophore model for HIV-1 integrase. J. Med. Chem. 43, 2100–2114 (2000).
Lin, J.-H., Perryman, A. L., Schames, J. R. & McCammon, J. A. Computational drug design accommodating receptor flexibility: the relaxed complex scheme. J. Am. Chem. Soc. 124, 5632–5633 (2002).
Yang, S., Salmon, L. & Al-Hashimi, H. M. Measuring similarity between dynamic ensembles of biomolecules. Nat. Methods 11, 552–554 (2014).
Salmon, L. et al. Modulating RNA alignment using directional dynamic kinks: application in determining an atomic-resolution ensemble for a hairpin using NMR residual dipolar couplings. J. Am. Chem. Soc. 137, 12954–12965 (2015).
Barril, X. & Morley, S. D. Unveiling the full potential of flexible receptor docking using multiple crystallographic structures. J. Med. Chem. 48, 4432–4443 (2005).
Craig, I. R., Essex, J. W. & Spiegel, K. Ensemble docking into multiple crystallographically derived protein structures: an evaluation based on the statistical analysis of enrichments. J. Chem. Inf. Model. 50, 511–524 (2010).
Tian, S. et al. Assessing an ensemble docking-based virtual screening strategy for kinase targets by considering protein flexibility. J. Chem. Inf. Model. 54, 2664–2679 (2014).
Treiber, D. K. & Williamson, J. R. Beyond kinetic traps in RNA folding. Curr. Opin. Struct. Biol. 11, 309–314 (2001).
Blackledge, M. Recent progress in the study of biomolecular structure and dynamics in solution from residual dipolar couplings. Prog. Nucl. Magn. Reson. Spectrosc. 46, 23–61 (2005).
Frank, A. T., Stelzer, A. C., Al-Hashimi, H. M. & Andricioaei, I. Constructing RNA dynamical ensembles by combining MD and motionally decoupled NMR RDCs: new insights into RNA dynamics and adaptive ligand recognition. Nucleic Acids Res. 37, 3670–3679 (2009).
Salmon, L., Bascom, G., Andricioaei, I. & Al-Hashimi, H. M. A general method for constructing atomic-resolution RNA ensembles using NMR residual dipolar couplings: the basis for interhelical motions revealed. J. Am. Chem. Soc. 135, 5457–5466 (2013).
Lange, O. F. et al. Recognition dynamics up to microseconds revealed from an RDC-derived ubiquitin ensemble in solution. Science 320, 1471–1475 (2008).
Salmon, L., Yang, S. & Al-Hashimi, H. M. Advances in the determination of nucleic acid conformational ensembles. Annu. Rev. Phys. Chem. 65, 293–316 (2014).
Fischer, M., Coleman, R. G., Fraser, J. S. & Shoichet, B. K. Incorporation of protein flexibility and conformational energy penalties in docking screens to improve ligand discovery. Nat. Chem. 6, 575–583 (2014).
Tóth, G. et al. Targeting the intrinsically disordered structural ensemble of α-synuclein by small molecules as a potential therapeutic strategy for Parkinson’s disease. PLoS One 9, e87133 (2014).
Tolman, J. R., Flanagan, J. M., Kennedy, M. A. & Prestegard, J. H. Nuclear magnetic dipole interactions in field-oriented proteins: information for structure determination in solution. Proc. Natl Acad. Sci. USA 92, 9279–9283 (1995).
Tjandra, N. & Bax, A. Direct measurement of distances and angles in biomolecules by NMR in a dilute liquid crystalline medium. Science 278, 1111–1114 (1997).
Mysinger, M. M., Carchia, M., Irwin, J. J. & Shoichet, B. K. Directory of useful decoys, enhanced (DUD-E): better ligands and decoys for better benchmarking. J. Med. Chem. 55, 6582–6594 (2012).
Chen, L., Calin, G. A. & Zhang, S. Novel insights of structure-based modeling for RNA-targeted drug discovery. J. Chem. Inf. Model. 52, 2741–2753 (2012).
Li, Y. et al. Accuracy assessment of protein-based docking programs against RNA targets. J. Chem. Inf. Model. 50, 1134–1146 (2010).
Morley, S. D. & Afshar, M. Validation of an empirical RNA-ligand scoring function for fast flexible docking using Ribodock. J. Comput. Aided Mol. Des. 18, 189–208 (2004).
Aboul-ela, F. Strategies for the design of RNA-binding small molecules. Future Med. Chem. 2, 93–119 (2010).
Verdonk, M. L. et al. Virtual screening using protein-ligand docking: avoiding artificial enrichment. J. Chem. Inf. Comput. Sci. 44, 793–806 (2004).
Abagyan, R., Totrov, M. & Kuznetsov, D. ICM - a new method for protein modeling and design: applications to docking and structure prediction from the distorted native conformation. J. Comput. Chem. 15, 488–506 (1994).
Neves, M. A. C., Totrov, M. & Abagyan, R. Docking and scoring with ICM: the benchmarking results and strategies for improvement. J. Comput. Aided Mol. Des. 26, 675–686 (2012).
Brozell, S. R. et al. Evaluation of DOCK 6 as a pose generation and database enrichment tool. J. Comput. Aided Mol. Des. 26, 749–773 (2012).
Gaudreault, F. & Najmanovich, R. J. FlexAID: revisiting docking on non-native-complex structures. J. Chem. Inf. Model. 55, 1323–1336 (2015).
Aboul-ela, F., Karn, J. & Varani, G. Structure of HIV-1 TAR RNA in the absence of ligands reveals a novel conformation of the trinucleotide bulge. Nucleic Acids Res. 24, 3974–3981 (1996).
Shi, H. & Moore, P. B. The crystal structure of yeast phenylalanine tRNA at 1.93 A resolution: a classic structure revisited. RNA 6, 1091–1105 (2000).
Yang, S. & Al-Hashimi, H. M. Unveiling inherent degeneracies in determining population- weighted ensembles of interdomain orientational distributions using NMR residual dipolar couplings: application to RNA helix junction helix motifs. J. Phys. Chem. B 119, 9614–9626 (2015).
Du, Z., Lind, K. E. & James, T. L. Structure of TAR RNA complexed with a Tat-TAR interaction nanomolar inhibitor that was identified by computational screening. Chem. Biol. 9, 707–712 (2002).
Murchie, A. I. H. et al. Structure-based drug design targeting an inactive RNA conformation: exploiting the flexibility of HIV-1 TAR RNA. J. Mol. Biol. 336, 625–638 (2004).
Davis, B. et al. Rational design of inhibitors of HIV-1 TAR RNA through the stabilisation of electrostatic “hot spots”. J. Mol. Biol. 336, 343–356 (2004).
Faber, C., Sticht, H., Schweimer, K. & Rösch, P. Structural rearrangements of HIV-1 Tat-responsive RNA upon binding of neomycin B. J. Biol. Chem. 275, 20660–20666 (2000).
Aboul-ela, F., Karn, J. & Varani, G. The structure of the human immunodeficiency virus type-1 TAR RNA reveals principles of RNA recognition by Tat protein. J. Mol. Biol. 253, 313–332 (1995).
Bailor, M. H., Sun, X. & Al-Hashimi, H. M. Topology links RNA secondary structure with global conformation, dynamics, and adaptation. Science 327, 202–206 (2010).
Pitt, S. W., Majumdar, A., Serganov, A., Patel, D. J. & Al-Hashimi, H. M. Argininamide binding arrests global motions in HIV-1 TAR RNA: comparison with Mg2+-induced conformational stabilization. J. Mol. Biol. 338, 7–16 (2004).
Pitt, S. W., Zhang, Q., Patel, D. J. & Al-Hashimi, H. M. Evidence that electrostatic interactions dictate the ligand-induced arrest of RNA global flexibility. Angew. Chem. Int. Edn. Engl. 44, 3412–3415 (2005).
Lang, P. T. et al. DOCK 6: combining techniques to model RNA-small molecule complexes. RNA 15, 1219–1230 (2009).
Rueda, M., Bottegoni, G. & Abagyan, R. Recipes for the selection of experimental protein conformations for virtual screening. J. Chem. Inf. Model. 50, 186–193 (2010).
Nichols, S. E., Baron, R., Ivetac, A. & McCammon, J. A. Predictive power of molecular dynamics receptor structures in virtual screening. J. Chem. Inf. Model. 51, 1439–1446 (2011).
Korb, O. et al. Potential and limitations of ensemble docking. J. Chem. Inf. Model. 52, 1262–1274 (2012).
Lind, K. E., Du, Z., Fujinaga, K., Peterlin, B. M. & James, T. L. Structure-based computational database screening, in vitro assay, and NMR assessment of compounds that target TAR RNA. Chem. Biol. 9, 185–193 (2002).
Yoon, S. & Welsh, W. J. Identification of a minimal subset of receptor conformations for improved multiple conformation docking and two-step scoring. J. Chem. Inf. Comput. Sci. 44, 88–96 (2004).
Puglisi, J. D., Tan, R., Calnan, B. J., Frankel, A. D. & Williamson, J. R. Conformation of the TAR RNA-arginine complex by NMR spectroscopy. Science 257, 76–80 (1992).
Bailor, M. H., Mustoe, A. M., Brooks, C. L. III & Al-Hashimi, H. M. 3D maps of RNA interhelical junctions. Nat. Protoc. 6, 1536–1545 (2011).
Matsumoto, C., Hamasaki, K., Mihara, H. & Ueno, A. A high-throughput screening utilizing intramolecular fluorescence resonance energy transfer for the discovery of the molecules that bind HIV-1 TAR RNA specifically. Bioorg. Med. Chem. Lett. 10, 1857–1861 (2000).
Calnan, B. J., Biancalana, S., Hudson, D. & Frankel, A. D. Analysis of arginine-rich peptides from the HIV Tat protein reveals unusual features of RNA-protein recognition. Genes Dev. 5, 201–210 (1991).
Loret, E. P., Georgel, P., Johnson, W. C. Jr. & Ho, P. S. Circular dichroism and molecular modeling yield a structure for the complex of human immunodeficiency virus type 1 trans-activation response RNA and the binding region of Tat, the trans-acting transcriptional activator. Proc. Natl Acad. Sci. USA 89, 9734–9738 (1992).
Shojania, S. & O’Neil, J. D. HIV-1 Tat is a natively unfolded protein: the solution conformation and dynamics of reduced HIV-1 Tat-(1-72) by NMR spectroscopy. J. Biol. Chem. 281, 8347–8356 (2006).
Zhang, J.-H., Chung, T. D. Y. & Oldenburg, K. R. A simple statistical parameter for use in evaluation and validation of high throughput screening assays. J. Biomol. Screen. 4, 67–73 (1999).
Sathyamoorthy, B., Lee, J., Kimsey, I., Ganser, L. R. & Al-Hashimi, H. Development and application of aromatic [(13)C, (1)H] SOFAST-HMQC NMR experiment for nucleic acids. J. Biomol. NMR 60, 77–83 (2014).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Goddard, T. D. & Kneller, D. G. SPARKY 3 https://www.cgl.ucsf.edu/home/sparky/ (University of California, San Francisco, 2008).
Sutter, P. & Weis, C. D. Ring opening reactions of 6H-anthra[1,9-cd]isoxazol-6-ones and related compounds. J. Heterocycl. Chem. 19, 997–1011 (1982).
Walter, F., Vicens, Q. & Westhof, E. Aminoglycoside-RNA interactions. Curr. Opin. Chem. Biol. 3, 694–704 (1999).
Zweckstetter, M. & Bax, A. Prediction of sterically induced alignment in a dilute liquid crystalline phase: Aid to protein structure determination by NMR. J. Am. Chem. Soc. 122, 3791–3792 (2000).
Zweckstetter, M. NMR: prediction of molecular alignment from structure using the PALES software. Nat. Protoc. 3, 679–690 (2008).
Hansen, A. L. & Al-Hashimi, H. M. Insight into the CSA tensors of nucleobase carbons in RNA polynucleotides from solution measurements of residual CSA: towards new long-range orientational constraints. J. Magn. Reson. 179, 299–307 (2006).
Acknowledgements
We thank M. Larsen and S. Vander Roest (University of Michigan Center of Chemical Genomics) for their help and input in carrying out high-throughput screening. We also thank the Duke Magnetic Resonance Spectroscopy Center for NMR resources and assistance in carrying out experiments and thank the Duke Compute Cluster for computational resources and support. This work was supported by the US National Institutes of Health (P50 GM103297, R01 AI066975 and P01 GM0066275 to H.M.A.-H.; T32 GM08487 and F31 GM119306 to L.R.G.).
Author information
Authors and Affiliations
Contributions
Experiments were designed by L.R.G., J.L., and H.M.A.-H.; performed by L.R.G., J.L., A.R., D.K.M., M.L.K., A.D.K., and B.S.; and analyzed by L.R.G. and H.M.A.-H. L.R.G. and H.M.A.-H. wrote the manuscript with input from all other authors.
Corresponding author
Ethics declarations
Competing interests
H.M.A.-H. is an advisor to and holds an ownership interest in Nymirum Inc., an RNA-based drug discovery company. Some of the technology used in this paper has been licensed to Nymirum.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 HTS identified seven hit molecules that bind TAR and displace Tat peptide.
a. Tat-displacement assays for the hit molecules in the absence (black) and presence (gray) of 100-fold excess tRNA. Points represent the mean and error bars represent the s.d. from n = 3 independent experiments. b. Overlay of SOFAST-[1H-13C] HMQC NMR spectra of 50 μM TAR free (black) and in the presence of 3X hit molecule (red, purple or blue). Spectra are overlaid for hits that induced similar chemical shift perturbations. (*) denotes folded peaks.
Supplementary Figure 2 NMR identified false positive molecules from HTS.
a. Examples of molecules from HTS that show activity in the Tat-displacement assay, but that do not bind TAR by NMR. b. Tat-displacement assay (black) and Tat-only control assay (gray). Points represent the mean and error bars represent the s.d. from n = 3 independent experiments. c. Overlay of SOFAST-[1H-13C] HMQC NMR spectra of 50 μM TAR both free (black) and in the presence of 6X CCG-111926 (red), 4X CCG-106134 (red), or 3X CCG-160257 (red). (*) denotes folded peaks.
Supplementary Figure 3 Testing for false positives in HTS by assaying compounds that are chemically similar to hit molecules.
a. Examples of molecules with chemical similarity to hit molecules that do not bind TAR. b. Tat-displacement assays (points saturating the fluorescence reader are removed). Points represent the mean and error bars represent the s.d. from n = 3 independent experiments. c. Overlay of SOFAST-[1H-13C] HMQC NMR spectra of 50 μM TAR both free (black) and in the presence of 3X small molecule (red). (*) denotes folded peaks.
Supplementary Figure 4 Testing for false positives in HTS based on EBVS scores.
a. TAR-binding molecules identified by testing a set of ten molecules from the top 5% of docking scores b. Tat-displacement assays in the absence (black) and presence (gray) of 100-fold excess tRNA (points saturating the fluorescence reader are removed). Points represent the mean and error bars represent the s.d. from n = 3 independent experiments. c. Overlay of SOFAST-[1H-13C] HMQC NMR spectra of 50 μM TAR both free (black) and in the presence of 3X molecule (red). (*) denotes folded peaks.
Supplementary Figure 5 Hit molecules of the Filtered and Optimized libraries identified in the literature.
Hit number corresponds to Supplementary Table 1 and (*) denotes molecules also in the Optimized library.
Supplementary Figure 6 ROC plot AUC values are robust across varied methods of defining hits and non-hits.
Variations in the similarity cutoff and number of non-hits selected per hit results in minor changes in the a. chemical property distributions and b. ROC plots for the Optimized library. c. ROC plots for all libraries (Full, Filtered and Optimized libraries) when using the Boltzmann-weighted average score, arithmetic average score, and best score for all hits before (purple) and after (blue) clustering by Bemis-Murcko atomic framework as well as cell-active hits before (red) and after (orange) clustering. ROC plots were generated from one run of docking all molecules to all receptors.
Supplementary Figure 7 Enrichment scores generally decrease when EBVS is applied to less accurate TAR ensembles.
a. ROC AUC and ROC(2%) scores for docking against individual conformers of the E0,4rdc ensemble, a randomly selected MD ensemble (E0,ran), and the lowest energy NOE-based structures for apo-TAR (PDB 1ANR) and tRNA (PDB 1EHZ) for the Full and Optimized libraries. Dashed lines indicate the values for the full ensemble. b. Dependence of the ROC AUC and ROC(2%) scores on ensemble size for the Full and Optimized libraries. c. Dependence of the ROC AUC and ROC(2%) scores on the RDC RMSD for the Full and Optimized libraries. d. Dependence of the ROC AUC and ROC(2%) scores on the RDC RMSD for the other 20-member ensembles of TAR. ROC plots were generated from one run of docking all molecules to all receptors. For b. c. and d. the mean and s.d. values over all possible sub-ensembles of each ensemble size are plotted.
Supplementary Figure 8 Each conformer of the ensemble contributes differentially to the small molecule scores and some hyper-enriching sub-ensembles outperform the N = 20 ensemble.
a. The Boltzmann-weighted average population of each conformer averaged across the Full library. b. The difference in the Boltzmann-weighted population of each conformer for each subset of molecules relative to the Full library. c. The percent of hyper-enriching sub-ensembles that contain each conformer for all libraries. (*) denotes conformers that most resemble ligand bound TAR conformations and (2) denotes conformers with two binding sites.
Supplementary Figure 9 Dependence of enrichment on TAR ensemble accuracy.
a. Dependence of the ROC AUC and ROC(2%) scores on the accuracy (RDC RMSD) of the TAR ensembles for the Full and Optimized libraries for all hits (blue) and cell-active hits (orange). All ROC values were generated from one run of docking all molecules to all receptors. b. Mean and s.d. of EBVS scores for hits (blue) and non-hits (gray) for all ensembles for the Full and Optimized libraries. Dashed lines represent the values for the E0,4rdc ensemble. c. Binding pocket volume (Å) and buriedness (ranging between 0.5–1.0 for fully open to fully occluded pockets, respectively) defined by ICM for each conformer of all TAR ensembles.
Supplementary Figure 10 RDC evaluation of ligand bound NMR structures and comparison of inter-helical angles for ligand bound NMR structures to EBVS predicted structures for E0,ran.
a. Agreement between NOE-based NMR structures and previously published RDCs using the best-fit order tensor determined using RAMAH. Red points denote bulge residues (U23-U25). The number of RDCs used in each correlation plot is n = 46, 30 and 40 for argininamide, acetylpromazine and Neomycin B, respectively. b. Comparison of inter-helical angles for the ligand-bound NMR structures (black, mean and s.d. values over all deposited structures) with conformers in the E0,ran ensemble (open squares) and the Boltzmann-weighted EBVS-predicted structures (red, mean and s.d. values over n = 20 independent docking runs).
Supplementary information
Supplementary Tables and Notes
Supplementary Notes 1 and 2, and Supplementary Table 2.
Supplementary Table 1
TAR binders augmented with hits reported in the literature. Measured CD50 values represent the mean and s.d. from n = 3 independent experiments.
Supplementary Table 3
Table of PDB structures used in the ligand bound RNA docking benchmark.
Rights and permissions
About this article
Cite this article
Ganser, L.R., Lee, J., Rangadurai, A. et al. High-performance virtual screening by targeting a high-resolution RNA dynamic ensemble. Nat Struct Mol Biol 25, 425–434 (2018). https://doi.org/10.1038/s41594-018-0062-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41594-018-0062-4
This article is cited by
-
Small molecule approaches to targeting RNA
Nature Reviews Chemistry (2024)
-
An RNA excited conformational state at atomic resolution
Nature Communications (2023)
-
Targeting RNA structures with small molecules
Nature Reviews Drug Discovery (2022)
-
The roles of structural dynamics in the cellular functions of RNAs
Nature Reviews Molecular Cell Biology (2019)