Abstract
Genetically encoded voltage indicators (GEVIs) hold immense potential for monitoring neuronal population activity. To date, best-in-class GEVIs rely on one-photon excitation. However, GEVI imaging of dense neuronal populations remains difficult because out-of-focus background fluorescence produces low contrast and excess noise when paired with conventional one-photon widefield imaging methods. To address this challenge, we developed an imaging system capable of efficient, high-contrast GEVI imaging at near-kHz rates and demonstrate it for in vivo and ex vivo imaging applications in the mouse neocortex. Our approach uses simultaneous multiplane imaging to monitor activity within contiguous tissue volumes with no penalty in speed or requirement for high excitation power. This approach, multi-Z imaging with confocal detection (MuZIC), permits high signal-to-noise ratio voltage imaging in densely labeled neuronal populations and is compatible with imaging through micro-optics. Moreover, it minimizes artifacts associated with concurrent imaging and optogenetic photostimulation for all-optical electrophysiology.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Data described in this study are available on figshare (https://doi.org/10.6084/m9.figshare.23542167). Information describing construction of our MuZIC system is available via GitHub (detector: https://github.com/tweber225/simple-sipm; assembly: https://github.com/tweber225/muzic).
Code availability
All code used for data analyses is provided on figshare (https://doi.org/10.6084/m9.figshare.23542167).
References
Chen, T.-W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Tian, L. et al. Imaging neural activity in worms, flies and mice with improved GCaMP calcium indicators. Nat. Methods 6, 875–881 (2009).
Zhang, Y. et al. Fast and sensitive GCaMP calcium indicators for imaging neural populations. Nature 615, 884–891 (2023).
Grienberger, C. & Konnerth, A. Imaging calcium in neurons. Neuron 73, 862–885 (2012).
Luo, L., Callaway, E. M. & Svoboda, K. Genetic dissection of neural circuits: a decade of progress. Neuron 98, 256–281 (2018).
Wachowiak, M. et al. Optical dissection of odor information processing in vivo using GCaMPs expressed in specified cell types of the olfactory bulb. J. Neurosci. 33, 5285–5300 (2013).
Wei, Z. et al. A comparison of neuronal population dynamics measured with calcium imaging and electrophysiology. PLOS Comput. Biol. 16, e1008198 (2020).
Abdelfattah, A. S. et al. Bright and photostable chemigenetic indicators for extended in vivo voltage imaging. Science 365, 699–704 (2019).
Abdelfattah, A. S. et al. Sensitivity optimization of a rhodopsin-based fluorescent voltage indicator. Neuron 111, 1547–1563 (2023).
Piatkevich, K. D. et al. Population imaging of neural activity in awake behaving mice. Nature 574, 413–417 (2019).
Adam, Y. et al. Voltage imaging and optogenetics reveal behaviour-dependent changes in hippocampal dynamics. Nature 569, 413–417 (2019).
Villette, V. et al. Ultrafast two-photon imaging of a high-gain voltage indicator in awake behaving mice. Cell 179, 1590–1608 (2019).
Hochbaum, D. R. et al. All-optical electrophysiology in mammalian neurons using engineered microbial rhodopsins. Nat. Methods 11, 825–833 (2014).
Knöpfel, T., Gallero-Salas, Y. & Song, C. Genetically encoded voltage indicators for large scale cortical imaging come of age. Curr. Opin. Chem. Biol. 27, 75–83 (2015).
Xiao, S. et al. Large-scale voltage imaging in behaving mice using targeted illumination. iScience 24, 103263 (2021).
Kazemipour, A. et al. Kilohertz frame-rate two-photon tomography. Nat. Methods 16, 778–786 (2019).
Wu, J. et al. Kilohertz two-photon fluorescence microscopy imaging of neural activity in vivo. Nat. Methods 17, 287–290 (2020).
Piatkevich, K. D. et al. A robotic multidimensional directed evolution approach applied to fluorescent voltage reporters. Nat. Chem. Biol. 14, 352–360 (2018).
Brinks, D., Klein, A. J. & Cohen, A. E. Two-photon lifetime imaging of voltage indicating proteins as a probe of absolute membrane voltage. Biophys. J. 109, 914–921 (2015).
Chamberland, S. et al. Fast two-photon imaging of subcellular voltage dynamics in neuronal tissue with genetically encoded indicators. eLife 6, e25690 (2017).
Badon, A. et al. Video-rate large-scale imaging with multi-Z confocal microscopy. Optica 6, 389–395 (2019).
Tsang, J.-M. et al. Fast, multiplane line-scan confocal microscopy using axially distributed slits. Biomed. Opt. Express 12, 1339–1350 (2021).
Lillis, K. P., Eng, A., White, J. A. & Mertz, J. Two-photon imaging of spatially extended neuronal network dynamics with high temporal resolution. J. Neurosci. Methods 172, 178–184 (2008).
Göbel, W., Kampa, B. M. & Helmchen, F. Imaging cellular network dynamics in three dimensions using fast 3D laser scanning. Nat. Methods 4, 73–79 (2007).
Katona, G. et al. Fast two-photon in vivo imaging with three-dimensional random-access scanning in large tissue volumes. Nat. Methods 9, 201–208 (2012).
Grewe, B. F., Langer, D., Kasper, H., Kampa, B. M. & Helmchen, F. High-speed in vivo calcium imaging reveals neuronal network activity with near-millisecond precision. Nat. Methods 7, 399–405 (2010).
Nadella, K. M. N. S. et al. Random-access scanning microscopy for 3D imaging in awake behaving animals. Nat. Methods 13, 1001–1004 (2016).
Lakowicz, J. R. Principles of Fluorescence Spectroscopy (Springer, 2006).
Andermann, M. L. et al. Chronic cellular imaging of entire cortical columns in awake mice using microprisms. Neuron 80, 900–913 (2013).
Rickgauer, J. P., Deisseroth, K. & Tank, D. W. Simultaneous cellular-resolution optical perturbation and imaging of place cell firing fields. Nat. Neurosci. 17, 1816–1824 (2014).
Packer, A. M., Russell, L. E., Dalgleish, H. W. P. & Häusser, M. Simultaneous all-optical manipulation and recording of neural circuit activity with cellular resolution in vivo. Nat. Methods 12, 140–146 (2015).
Altkorn, R. & Zare, R. N. Effects of saturation on laser-induced fluorescence measurements of population and polarization. Annu. Rev. Phys. Chem. 35, 265–289 (1984).
Zhang, L. et al. Miniscope GRIN lens system for calcium imaging of neuronal activity from deep brain structures in behaving animals. Curr. Protoc. Neurosci. 86, e56 (2019).
Murayama, M. & Larkum, M. E. In vivo dendritic calcium imaging with a fiberoptic periscope system. Nat. Protoc. 4, 1551–1559 (2009).
Pologruto, T. A., Sabatini, B. L. & Svoboda, K. ScanImage: flexible software for operating laser scanning microscopes. Biomed. Eng. Online 2, 13 (2003).
Savarese, M. et al. Fluorescence lifetimes and quantum yields of rhodamine derivatives: new insights from theory and experiment. J. Phys. Chem. A 116, 7491–7497 (2012).
Kim, J.-Y., Grunke, S. D., Levites, Y., Golde, T. E. & Jankowsky, J. L. Intracerebroventricular viral injection of the neonatal mouse brain for persistent and widespread neuronal transduction. J. Vis. Exp. 91, e51863 (2014).
Ting, J. T., Daigle, T. L., Chen, Q. & Feng, G. in Patch-Clamp Methods and Protocols (eds Martina, M. & Taverna, S.) 221–242 (Springer, 2014).
Grimm, J. B. et al. A general method to fine-tune fluorophores for live-cell and in vivo imaging. Nat. Methods 14, 987–994 (2017).
Kılıç, K. et al. Chronic cranial windows for long term multimodal neurovascular imaging in mice. Front. Physiol. 11, 612678 (2021).
Peron, S. P., Freeman, J., Iyer, V., Guo, C. & Svoboda, K. A cellular resolution map of barrel cortex activity during tactile behavior. Neuron 86, 783–799 (2015).
Acknowledgements
We thank A. Abdelfattah, A. Singh, G. Murphy and K. Svoboda for technical advice and discussions. We thank E. Schreiter and I. Kolb for sharing Voltron2-ST reagents prepublication. We thank M. Giacomelli for advice on silicon photomultipliers. We thank T. Wang and D. Kleinfeld for comments on the paper. This work was supported by research grants from the National Institutes of Health: grant no. R01EB029171 (J.M.), grant no. RF1MH126882 (M.N.E.) and grant no. F32MH129149 (M.V.M.). The funder had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
T.D.W., M.V.M., J.M. and M.N.E. conceived of the project. M.N.E. and J.M. supervised research. T.D.W., J.M. and M.N.E. designed the MuZIC system. M.V.M., K.K. and M.N.E. prepared samples. T.D.W., M.V.M., K.K. and M.N.E. performed experiments. T.D.W., M.V.M. and M.N.E. analyzed data. T.D.W. performed simulations. T.D.W., M.V.M., J.M. and M.N.E. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
J.M., T.D.W. and Boston University have a US patent filed (no. 11,042,016) that relates to multiplane confocal imaging. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Neuroscience thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Detailed system schematic.
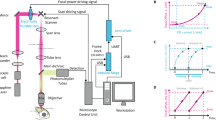
The light source is a continuous-wave laser diode (‘561 nm’). Blue dashed box (‘Photo-stim.’) denotes components of the full-field photo-stimulation module. Green dashed box (‘Epi-illum.’) denotes components of the epi-illumination widefield fluorescence microscopy unit. M: mirror, Ir: iris, L: lens, DM: dichroic mirror, PS: polygonal scanner, GS: galvanometer scanner, SM: switchable mirror, OL: objective lens, FPs: focal planes, PH: pinhole, D: detector, C: camera. All part numbers are for the manufacturer Thorlabs, except the objective lens (Nikon).
Extended Data Fig. 2 System resolution.
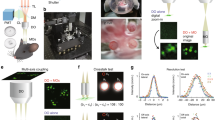
Experimental measurements of the combined excitation and emission point spread functions (PSFs)corresponding to the four simultaneously acquired image planes. Confocal pinhole diameters were selected so that PSFs corresponding to adjacent detectors overlap, eliminating gaps between the four imaging planes, which together cover ~40 μm in the axial direction. Lateral resolution was somewhat anisotropic on account of the slower detector fall time. (X: 2.6 μm full-width half-max; Y: 1.9 μm).
Extended Data Fig. 3 Photobleaching.
a. Left, Example of raw, unprocessed fluorescence from all cells identified from the in vitro imaging experiment depicted in Fig. 6a,b. Right, Exponential fits to raw fluorescence normalized to initial brightness. These data were collected with the highest excitation power of all in vitro experiments (561 nm; 23.6 mW/mm2). b. Raw, unprocessed recorded from all cells identified in the in vivo imaging experiment depicted in Fig. 3b (top) and exponential fits to in vivo fluorescence, as in (a). The same excitation power was used for all in vivo imaging experiments (561 nm; 300 mW/mm2).
Extended Data Fig. 4 Multi-Z imaging minimizes axial motion artifacts.
a. During periods of brain motion, fluorescence is redistributed across planes, with cells becoming brighter in some planes and dimmer in others. Top, average change in fluorescence during a period of movement indicated by arrow below in adjacent planes. Bottom, average fluorescence time series corresponding to the cell indicated (yellow arrowhead). b. Top, mean fluorescence (left), difference image prior to (middle) and following (right)lateral motion correction. Sum of 4 image planes shown. During periods of brain motion (indicated by black arrow, bottom), lateral motion correction compensates for brain motion. Bottom, fluorescence time series corresponding to the cell indicated (yellow arrowhead, top). Time series extracted from a single plane(plane 2) before lateral motion correction (light blue, top), after lateral motion correction (dark blue, middle), and from a region of interest (ROI) spanning all four image planes following lateral motion correction (gray, bottom). Artifacts introduced by brain motion are suppressed when extracting time series from multiplane ROIs. Data are representative of axial motion across 34 fields, 5 animals.
Extended Data Fig. 5 Confocal collection efficiency.
Simulated confocal collection efficiency for fluorescence emitted from point sources (left) and 15-μm shells (right) located at different axial positions. Axial resolution is degraded as a result of spherical aberration induced by imaging through 0.5 mm cover glass (bottom; RI = 1.52). Lateral resolution is not degraded as much as axial resolution, meaning that the total fluorescence together collected by all detectors is only modestly reduced. See Methods and Supplementary Note 4 for more details.
Extended Data Fig. 6 Comparison of MuZIC and widefield fluorescence microscopy (WFM).
a. Signal-to-noise ratio of MuZIC relative to WFM increases with increasing WFM background-to-signal ratio (BSR). SNR of MuZIC is mostly insensitive to out-of-focus background fluorescence, while SNR of WFM decreases with increasing background. When BSR is greater than about 5, SNR of MuZIC is higher than SNR of WFM (assuming contrast of MuZIC is 0.6 and detection efficiency of MuZIC is 25% relative to WFM; see Supplementary Note 2). b. WFM image of Voltron2-labeled neurons in vivo. Representative BSRs associated with different neurons are indicated for reference. Individual neurons were difficult to resolve in regions of higher labeling density (representative of 4 animals). c. Example in-vivo widefield images of neurons without (top) and with (bottom) targeted illumination delimited by dashed circles. Dashed circles span regions of interest encompassing a single neuron (top left: signal + background) and background only (top right). Corresponding images when targeted illumination was applied (bottom) are shown recentered. In this extreme example where only a single neuron is targeted, the BSR is found to be reduced by a factorγ = 0.2(see Supplementary Note 3, representative of 4 animals). d. Signal-to-noise ratio of MuZIC relative to WFM without (solid) and with (dashed) the addition of targeted illumination of a single neuron (that isγ= 0.2). In general, when multiple neurons are targeted, the reduction in BSR is not as significant and the relative SNR gain provided by MuZIC lies within the shaded region. Targeted illumination has the effect of shifting the break-even point where MuZIC becomes advantageous over WFM to a higher BSR value, up to about 25 in the very extreme case of single-neuron targeted illumination.
Supplementary information
Supplementary Information
Supplementary Notes 1–5 and Tables 1 and 2.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Weber, T.D., Moya, M.V., Kılıç, K. et al. High-speed multiplane confocal microscopy for voltage imaging in densely labeled neuronal populations. Nat Neurosci 26, 1642–1650 (2023). https://doi.org/10.1038/s41593-023-01408-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-023-01408-2