Abstract
Repeat expansion in C9ORF72 is the most common genetic cause of amyotrophic lateral sclerosis (ALS) and frontotemporal dementia (FTD). Here we show that N6-methyladenosine (m6A), the most prevalent internal mRNA modification, is downregulated in C9ORF72-ALS/FTD patient-derived induced pluripotent stem cell (iPSC)-differentiated neurons and postmortem brain tissues. The global m6A hypomethylation leads to transcriptome-wide mRNA stabilization and upregulated gene expression, particularly for genes involved in synaptic activity and neuronal function. Moreover, the m6A modification in the C9ORF72 intron sequence upstream of the expanded repeats enhances RNA decay via the nuclear reader YTHDC1, and the antisense RNA repeats can also be regulated through m6A modification. The m6A reduction increases the accumulation of repeat RNAs and the encoded poly-dipeptides, contributing to disease pathogenesis. We further demonstrate that, by elevating m6A methylation, we could significantly reduce repeat RNA levels from both strands and the derived poly-dipeptides, rescue global mRNA homeostasis and improve survival of C9ORF72-ALS/FTD patient iPSC-derived neurons.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Requests for further information or resources and reagents should be directed to and will be filled by the lead author, Shuying Sun (shuying.sun@jhmi.edu). Plasmids generated in this study are available from the lead author upon reasonable request. The sequencing data (Supplementary Table 7) have been deposited at the National Center for Biotechnology Information under Gene Expression Omnibus accession number GSE203581. Source data are provided with this paper.
Code availability
This paper does not report original code.
References
Nussbacher, J. K., Tabet, R., Yeo, G. W. & Lagier-Tourenne, C. Disruption of RNA metabolism in neurological diseases and emerging therapeutic interventions. Neuron 102, 294–320 (2019).
Ferrari, R., Kapogiannis, D., Huey, E. D. & Momeni, P. FTD and ALS: a tale of two diseases. Curr. Alzheimer Res. 8, 273–294 (2011).
Ito, D., Hatano, M. & Suzuki, N. RNA binding proteins and the pathological cascade in ALS/FTD neurodegeneration. Sci. Transl. Med. 9, eaah5436 (2017).
Freibaum, B. D. & Taylor, J. P. The role of dipeptide repeats in C9ORF72-related ALS-FTD. Front. Mol. Neurosci. 10, 35 (2017).
Cheng, W. et al. CRISPR–Cas9 screens identify the RNA helicase DDX3X as a repressor of C9ORF72 (GGGGCC)n repeat-associated non-AUG translation. Neuron 104, 885–898 (2019).
Dominissini, D. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485, 201–206 (2012).
Meyer, K. D. et al. Comprehensive analysis of mRNA methylation reveals enrichment in 3′ UTRs and near stop codons. Cell 149, 1635–1646 (2012).
Liu, J. et al. A METTL3–METTL14 complex mediates mammalian nuclear RNA N6-adenosine methylation. Nat. Chem. Biol. 10, 93–95 (2014).
Roundtree, I. A., Evans, M. E., Pan, T. & He, C. Dynamic RNA modifications in gene expression regulation. Cell 169, 1187–1200 (2017).
Jia, G. et al. N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat. Chem. Biol. 7, 885–887 (2011).
Zheng, G. et al. ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol. Cell 49, 18–29 (2013).
Xiao, W. et al. Nuclear m6A reader YTHDC1 regulates mRNA splicing. Mol. Cell 61, 507–519 (2016).
Du, H. et al. YTHDF2 destabilizes m6A-containing RNA through direct recruitment of the CCR4–NOT deadenylase complex. Nat. Commun. 7, 12626 (2016).
Wang, X. et al. N6-methyladenosine modulates messenger RNA translation efficiency. Cell 161, 1388–1399 (2015).
Wang, X. et al. N6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505, 117–120 (2014).
Liu, J. et al. Landscape and regulation of m6A and m6Am methylome across human and mouse tissues. Mol. Cell 77, 426–440 (2020).
Livneh, I., Moshitch-Moshkovitz, S., Amariglio, N., Rechavi, G. & Dominissini, D. The m6A epitranscriptome: transcriptome plasticity in brain development and function. Nat. Rev. Neurosci. 21, 36–51 (2020).
Deng, X. et al. Widespread occurrence of N6-methyladenosine in bacterial mRNA. Nucleic Acids Res. 43, 6557–6567 (2015).
Tian, R. et al. CRISPR interference-based platform for multimodal genetic screens in human iPSC-derived neurons. Neuron 104, 239–255 (2019).
Mackenzie, I. R. et al. Quantitative analysis and clinico-pathological correlations of different dipeptide repeat protein pathologies in C9ORF72 mutation carriers. Acta Neuropathol. 130, 845–861 (2015).
Mori, K. et al. The C9orf72 GGGGCC repeat is translated into aggregating dipeptide-repeat proteins in FTLD/ALS. Science 339, 1335–1338 (2013).
Cheng, W. et al. C9ORF72 GGGGCC repeat-associated non-AUG translation is upregulated by stress through eIF2α phosphorylation. Nat. Commun. 9, 51 (2018).
Wang, S. et al. Nuclear export and translation of circular repeat-containing intronic RNA in C9ORF72-ALS/FTD. Nat. Commun. 12, 4908 (2021).
Liu, J. et al. N6-methyladenosine of chromosome-associated regulatory RNA regulates chromatin state and transcription. Science 367, 580–586 (2020).
Lubas, M. et al. Interaction profiling identifies the human nuclear exosome targeting complex. Mol. Cell 43, 624–637 (2011).
Mori, K. et al. Bidirectional transcripts of the expanded C9orf72 hexanucleotide repeat are translated into aggregating dipeptide repeat proteins. Acta Neuropathol. 126, 881–893 (2013).
Lagier-Tourenne, C. et al. Targeted degradation of sense and antisense C9orf72 RNA foci as therapy for ALS and frontotemporal degeneration. Proc. Natl Acad. Sci. USA 110, E4530–E4539 (2013).
Gendron, T. F. et al. Antisense transcripts of the expanded C9ORF72 hexanucleotide repeat form nuclear RNA foci and undergo repeat-associated non-ATG translation in c9FTD/ALS. Acta Neuropathol. 126, 829–844 (2013).
Mizielinska, S. et al. C9orf72 frontotemporal lobar degeneration is characterised by frequent neuronal sense and antisense RNA foci. Acta Neuropathol. 126, 845–857 (2013).
Zu, T. et al. RAN proteins and RNA foci from antisense transcripts in C9ORF72 ALS and frontotemporal dementia. Proc. Natl Acad. Sci. USA 110, E4968–E4977 (2013).
Jiang, J. et al. Gain of toxicity from ALS/FTD-linked repeat expansions in C9ORF72 is alleviated by antisense oligonucleotides targeting GGGGCC-containing RNAs. Neuron 90, 535–550 (2016).
Donnelly, C. J. et al. RNA toxicity from the ALS/FTD C9ORF72 expansion is mitigated by antisense intervention. Neuron 80, 415–428 (2013).
Huang, Y. et al. Small-molecule targeting of oncogenic FTO demethylase in acute myeloid leukemia. Cancer Cell 35, 677–691.e10 (2019).
Su, R. et al. Targeting FTO suppresses cancer stem cell maintenance and immune evasion. Cancer Cell 38, 79–96 (2020).
Liu, Y. et al. Tumors exploit FTO-mediated regulation of glycolytic metabolism to evade immune surveillance. Cell Metab. 33, 1221–1233 (2021).
Jiang, X. et al. The role of m6A modification in the biological functions and diseases. Signal Transduct. Target Ther. 6, 74 (2021).
Lence, T. et al. m6A modulates neuronal functions and sex determination in Drosophila. Nature 540, 242–247 (2016).
Wang, C. X. et al. METTL3-mediated m6A modification is required for cerebellar development. PLoS Biol. 16, e2004880 (2018).
Devlin, A. C. et al. Human iPSC-derived motoneurons harbouring TARDBP or C9ORF72 ALS mutations are dysfunctional despite maintaining viability. Nat. Commun. 6, 5999 (2015).
Wainger, B. J. et al. Intrinsic membrane hyperexcitability of amyotrophic lateral sclerosis patient-derived motor neurons. Cell Rep. 7, 1–11 (2014).
Koranda, J. L. et al. Mettl14 is essential for epitranscriptomic regulation of striatal function and learning. Neuron 99, 283–292 (2018).
Lall, D. et al. C9orf72 deficiency promotes microglial-mediated synaptic loss in aging and amyloid accumulation. Neuron 109, 2275–2291 (2021).
Zhu, Q. et al. Reduced C9ORF72 function exacerbates gain of toxicity from ALS/FTD-causing repeat expansion in C9orf72. Nat. Neurosci. 23, 615–624 (2020).
Schmitz, A., Pinheiro Marques, J., Oertig, I., Maharjan, N. & Saxena, S. Emerging perspectives on dipeptide repeat proteins in C9ORF72 ALS/FTD. Front. Cell. Neurosci. 15, 637548 (2021).
McEachin, Z. T., Parameswaran, J., Raj, N., Bassell, G. J. & Jiang, J. RNA-mediated toxicity in C9orf72 ALS and FTD. Neurobiol. Dis. 145, 105055 (2020).
Donnelly, C. J., Grima, J. C. & Sattler, R. Aberrant RNA homeostasis in amyotrophic lateral sclerosis: potential for new therapeutic targets? Neurodegener. Dis. Manag. 4, 417–437 (2014).
Conlon, E. G. et al. The C9ORF72 GGGGCC expansion forms RNA G-quadruplex inclusions and sequesters hnRNP H to disrupt splicing in ALS brains. eLife 5, e17820 (2016).
Lee, Y. B. et al. Hexanucleotide repeats in ALS/FTD form length-dependent RNA foci, sequester RNA binding proteins, and are neurotoxic. Cell Rep. 5, 1178–1186 (2013).
Sareen, D. et al. Targeting RNA foci in iPSC-derived motor neurons from ALS patients with a C9ORF72 repeat expansion. Sci. Transl. Med. 5, 208ra149 (2013).
Xu, Z. et al. Expanded GGGGCC repeat RNA associated with amyotrophic lateral sclerosis and frontotemporal dementia causes neurodegeneration. Proc. Natl Acad. Sci. USA 110, 7778–7783 (2013).
McMillan, M. et al. RNA methylation influences TDP43 binding and disease pathogenesis in models of amyotrophic lateral sclerosis and frontotemporal dementia. Mol. Cell. 83, 219–236 (2023).
Zhao, F. et al. METTL3-dependent RNA m6A dysregulation contributes to neurodegeneration in Alzheimer’s disease through aberrant cell cycle events. Mol. Neurodegener. 16, 70 (2021).
Alzheimer’s Association. 2016 Alzheimer’s disease facts and figures. Alzheimers Dement. 12, 459–509 (2016).
Shafik, A. M. et al. N6-methyladenosine dynamics in neurodevelopment and aging, and its potential role in Alzheimer’s disease. Genome Biol. 22, 17 (2021).
Chen, X. et al. Down-regulation of m6A mRNA methylation is involved in dopaminergic neuronal death. ACS Chem. Neurosci. 10, 2355–2363 (2019).
Ababneh, N. A. et al. Correction of amyotrophic lateral sclerosis related phenotypes in induced pluripotent stem cell-derived motor neurons carrying a hexanucleotide expansion mutation in C9orf72 by CRISPR/Cas9 genome editing using homology-directed repair. Hum. Mol. Genet. 29, 2200–2217 (2020).
Cleary, E. M. et al. Improved PCR based methods for detecting C9orf72 hexanucleotide repeat expansions. Mol. Cell Probes 30, 218–224 (2016).
Zaepfel, B. L. et al. UPF1 reduces C9orf72 HRE-induced neurotoxicity in the absence of nonsense-mediated decay dysfunction. Cell Rep. 34, 108925 (2021).
Coyne, A. N. et al. Nuclear accumulation of CHMP7 initiates nuclear pore complex injury and subsequent TDP-43 dysfunction in sporadic and familial ALS. Sci. Transl. Med. 13, eabe1923 (2021).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, giab008 (2021).
Zhang, Y. et al. Model-based analysis of ChIP-seq (MACS). Genome Biol. 9, R137 (2008).
Cui, X. et al. MeTDiff: a novel differential RNA methylation analysis for MeRIP-seq data. IEEE/ACM Trans. Comput. Biol. Bioinform. 15, 526–534 (2018).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Dennis, G. et al. DAVID: database for annotation, visualization, and integrated discovery. Genome Biol. 4, P3 (2003).
Sherman, B. T. et al. DAVID: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res. 50, W216–W221 (2022).
Huang, D. W. et al. DAVID bioinformatics resources: expanded annotation database and novel algorithms to better extract biology from large gene lists. Nucleic Acids Res. 35, W169–W175 (2007).
Acknowledgements
We thank Target ALS Postmortem Tissue Core and Johns Hopkins ALS Postmortem Tissue Core for providing postmortem human brain tissue samples. We thank T. Thompson from Answer ALS consortium for providing help and detailed data information. We thank M. Ward and his laboratory at the National Institutes of Health (NIH) for sharing the i3Neuron iPSC line and the i3Neuron differentiation protocols. We thank K. Talbot at the University of Oxford for providing the isogenic pair of patient iPSC lines. We thank L. Xue for help with the GP ELISA. We thank J. Rothstein and his laboratory for sharing control and patient iPSC lines and protocols. We thank Sun laboratory members for helpful discussions. This work is supported by the Johns Hopkins University Catalyst Award (S.S.); NIH grants R01NS107347 (S.S.), RF1NS113820 (S.S.), R21AG072078 (S.S.), RF1NS127925 (S.S.), HG008935 (C.H.), ES030546 (C.H.), K08NS104273 (L.H.) and R01NS123538 (L.H.); and the Packard Center for ALS Research (S.S.). Z.Z. was a recipient of the Milton Safenowitz Post-Doctoral Fellowship from the ALS Association, the Toffler Scholar Award and the Postdoc Development Grant from the Muscular Dystrophy Association. The Mass Spectrometry Facility of the University of Chicago is funded by the National Science Foundation (CHE-1048528). C.H. is an investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
Y.L., X.D., C.H. and S.S. contributed to the overall design of the study and wrote the paper. Y.L. performed most molecular and cellular biology experiments, with help from Y.X., J.L., Z.Z., R.W., X.F. and Y.Y., under the mentorship of S.S. and C.H. X.D. and Y.L. performed bioinformatic analysis of the high-throughput sequencing data. B.Y. performed blinded analysis of glutamate excitotoxicity. L.H. performed GP ELISA assay and analysis. L.W.O. advised on patient samples and provided key reagents.
Corresponding authors
Ethics declarations
Competing interests
C.H. is a scientific founder, a member of the scientific advisory board and an equity holder of Aferna Bio, Inc. and AccuaDX Inc.; a scientific co-founder and equity holder of Accent Therapeutics, Inc.; and a member of the scientific advisory board of Rona Therapeutics. All other authors declare no competing interests.
Peer review
Peer review information
Nature Neuroscience thanks Adrian Isaacs, Yongchao Ma and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 m6A pathway-related gene expression in C9ORF72-ALS/FTD iPSNs.
a, Proteomic heatmap of the m6A pathway components in control (n = 26) and C9ORF72-ALS/FTD (n = 22) iPSNs. Data were normalized to the batch controls. b, Boxplots of the proteomic data in (a). Box plots indicate the interquartile range with the central line representing the median, and the vertical lines extend to the extreme values in the group. c, Western blotting (left) and quantification (right) of METTL3 and METTL14 expression in control and C9ORF72 iPSNs. Data are mean ± s.e.m. ***P = 0.0008 and 0.0003, respectively, by two-tailed t test. d, Relative RNA expression of neuronal marker genes (MAP2, PSD95 and TUBB3) was measured by RT-qPCR in control and C9 iPSNs, showing comparable neuron maturity between control and C9. Points represent individual control or patient lines. n = 6 in each group. Data are mean ± s.e.m. MAP2, P = 0.304; PSD95, P = 0.86; TUBB3, P = 0.716, by two-tailed t test.
Extended Data Fig. 2 METTL3 and METTL14 m6A methyltransferases were reduced in C9ORF72-ALS/FTD postmortem brain.
a, b, Immunohistochemistry (IHC) staining of METTL3 (a), METTL14 (b) at temporal cortex section of control and C9ORF72-ALS/FTD samples. Scale bar = 50 μm. c, Western blotting (top) and quantification (bottom) of METTL3, METTL14 using postmortem motor cortex tissue samples. *Non-specific bands. Points represent individual control or patient samples. n = 3 in each group. Data are mean ± s.e.m. *P = 0.0171, **P = 0.0011, by two-tailed t test.
Extended Data Fig. 3 Expression changes of the m6A ‘writer’ core components in sporadic ALS iPSNs.
a, Proteomic heatmap of the core components of m6A ‘writer’ complex in control (n = 19) and sporadic ALS (n = 93) iPSNs. Data were normalized to the batch controls. b, Boxplots of the proteomic data in (a). Box plots indicate the interquartile range with the central line representing the median, and the vertical lines extend to the extreme values in the group. c, UHPLC-QQQ-MS/MS quantification of the m6A/A ratio in poly-A mRNAs from control (n = 5) and sporadic ALS patient (n = 7) postmortem frontal cortex. Data are mean ± s.e.m. Points represent individual samples. *P = 0.0247 by two-tailed t test.
Extended Data Fig. 4 The repeat RNA induces downregulation of METTL3 and METTL14 expression.
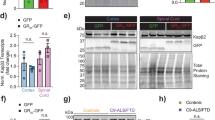
a, Timeline for doxycycline-induced neuron differentiation, lentiviral transduction of transgenes to express repeats or DPRs, and functional analysis of i3Neurons. b, Relative RNA expression of METTL3 and METTL14 in i3Neurons expressing GFP control, modified poly-GA, modified poly-GR (with randomized codons), or (GGGGCC)160 repeats. METTL3: mGA, P = 0.1154; mGR, P = 0.9755; GC160, ****P < 0.0001. METTL14: mGA, P = 0.3892; mGR, P = 0.9853; GC160, **P = 0.0073. P values were calculated by one-way analysis of variance (ANOVA) with Tukey’s multiple comparisons c, Detection of GFP, modified poly-GA and modified poly-GR post lentiviral transduction in i3Neurons by Nano-Glo HiBiT bioluminescence measurement. The luciferase signal was normalized to the total protein level. i3Neurons without lentiviral infection served as the negative control. *P = 0.0148, **P = 0.0062, ***P = 0.0004, by two-tailed t test. d, Immunofluorescent staining of GFP, modified poly-GA or modified poly-GR in transduced i3Neurons. Neurons were stained using FLAG antibody against the C-terminal FLAG epitope tag in each construct. (b,c) Points represent three biological replicates. Data are mean ± s.e.m.
Extended Data Fig. 5 MeRIP-seq in control and C9ORF72-ALS/FTD iPSNs and postmortem motor cortex.
a, Venn diagram depicting the total number and overlaps of m6A peaks detected in control and C9ORF72-ALS/FTD groups of iPSN (top) and motor cortex (bottom). b, The most enriched consensus m6A motifs in all conditions. c, Metagene profiles of m6A peak density along transcripts with three non-overlapping segments (5’ UTR, CDS, and 3’ UTR) for control and C9ORF72-ALS/FTD groups. d, Principal component analysis (PCA) of control and C9ORF72-ALS/FTD samples. e, Gene Ontology (GO) enrichment analysis of hypomethylated genes that were identified in both iPSNs and motor cortex. P value was calculated by one-sided Fisher’s Exact test.
Extended Data Fig. 6 The impact of m6A hypomethylation on mRNAs.
a, b, Volcano plot of differentially expressed genes in C9ORF72-ALS/FTD compared to the control, in iPSNs (a) and postmortem motor cortex (b). Genes with significant changes are represented by orange (upregulated in C9) and blue (downregulated in C9) dots. iPSNs: n = 4 individual lines per group, 1212 upregulated vs 1168 downregulated. Motor cortex: n = 3 individual tissue samples per group, 1892 upregulated versus 1239 downregulated. P < 0.05, two-tailed Wald test. c, YTHDF2-RIP qPCR of selected YTHDF2 mRNA targets identified from the non-YTHDF2 bound RNAs. HPRT1 and SMARCA1 mRNAs have no m6A modification according to the MeRIP-seq results. Points represent individual iPSN lines. n = 3 in each group. Data are mean ± s.e.m. P = 0.7047 and 0.4001, respectively, by two-tailed t test. d, Cumulative distribution demonstrating the half-life of YTHDF2 targets between control and C9ORF72-ALS/FTD iPSNs. P value was calculated by a two-tailed non-parametric Wilcoxon-Matt-Whitney test. e, Gene Ontology (GO) enrichment analysis of hypomethylated YTHDF2 binding targets which also showed significantly increased half-life. P value was calculated by one-sided Fisher’s Exact test.
Extended Data Fig. 7 m6A regulates C9ORF72 repeat-containing RNA.
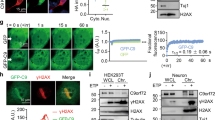
a, IGV profile showing the m6A peaks (generated by comparing MeRIP against input) in control and C9ORF72-ALS/FTD iPSNs. Red arrow head represents the repeat expansion. b, Western blotting of METTL14 in control or METTL14 knockdown i3Neurons. *non-specific band. c, The relative basal expression of NLuc RNA in the HeLa Flp-In cells expressing the reporters as described in Fig. 4b. *P = 0.015 by two-tailed t test. d, The relative expression level of C9R-NLuc RNA upon knockdown of METTL14 in i3Neurons, measured by RT-qPCR and normalized to non-targeting shRNA control. ****P < 0.0001 by two-tailed t test. (c,d) Points represent three biological replicates. Data are mean ± s.e.m.
Extended Data Fig. 8 The turnover of C9ORF72 repeat-containing RNA is regulated by m6A in i3Neurons.
a, RT-qPCR of C9 repeat RNA expression from the C9R-NLuc HeLa reporter upon knockdown of YTHDF2. P = 0.5418 by two-tailed t test. b, Relative DPR level was measured by luciferase assay upon YTHDF2 knockdown in the C9R-NLuc HeLa reporter. c, Western blotting of YTHDF2 in control or YTHDF2 knockdown HeLa Flp-In cells. P = 0.9456, 0.8801 and 0.9731, respectively, by one-way analysis of variance (ANOVA) with Tukey’s multiple comparisons. d, e, The relative expression level of YTHDC1 RNA (d) or C9R-NLuc RNA (e) upon knockdown of YTHDC1 by shRNA in i3Neurons. **P = 0.0046, ***P = 0.0006 by two-tailed t test. f, g, Relative DPR levels were measured by luciferase assay upon YTHDC1 knockdown (f) or ZCCHC8 knockdown (g) in the i3Neurons expressing the reporters in Fig. 4b. NLuc signal was normalized to the total protein level in each sample and the relative expression was compared to non-targeting shRNA control. ****P < 0.0001 by one-way analysis of variance (ANOVA) with Tukey’s multiple comparisons. h, Western blotting of ZCCHC8 in control or ZCCHC8 knockdown i3Neurons. Arrow points the expected size of ZCCHC8. (a, b, d-g) Data are mean ± s.e.m. Points represent three biological replicates.
Extended Data Fig. 9 m6A hypomethylation of the endogenous C9 repeat-containing intron and YTHDC1 expression in the postmortem brain tissues.
a, MeRIP coupled with RT-qPCR of endogenous C9 repeat-containing intronic RNA in control and C9ORF72-ALS/FTD iPSNs. Points represent individual cell lines. n = 6 per group. *P = 0.0427. b, Representative IHC staining of YTHDC1 at temporal cortex section of control and C9ORF72-ALS/FTD samples. Scale bar = 50 μm. c, Western blotting (left) and quantification (right) of YTHDC1 using postmortem motor cortex tissue samples. n = 3 individuals per group. *Non-specific bands. *P = 0.0246. (a, c) Data are mean ± s.e.m. Points represent individual control or patient samples. P values were calculated by two-tailed t test.
Extended Data Fig. 10 m6A restoration reduces glutamate-induced excitotoxicity in C9ORF72-ALS/FTD iPSNs.
a, RT-qPCR of METTL14 in C9ORF72-ALS/FTD iPSNs expressing exogenous METTL14 or GFP control. Data are mean ± s.e.m. Points represent individual cell lines. n = 6 in each group. ***P = 0.0006 by paired two-tailed t test. b, Western blotting of FTO in C9ORF72-ALS/FTD iPSNs upon knockdown of FTO by shRNA. *non-specific band. c, d, Representative images of Hoechst and propidium iodide (PI) staining of iPSNs in the glutamate-induced excitotoxicity assay. C9ORF72-ALS/FTD iPSNs were infected with lentivirus overexpressing METTL14 or GFP as negative control (c), or expressing FTO shRNA or non-targeting control shRNA (d).
Supplementary information
41593_2023_1374_MOESM2_ESM.xlsx
Supplementary Table 1: Demographic information for lymphoblast lines. Supplementary Table 2: Demographic information for iPSC lines. Supplementary Table 3: Demographic information for frozen postmortem tissues. Supplementary Table 4: Demographic information for formalin-fixed, paraffin embedded (FFPE) tissue slides of temporal cortex. Supplementary Table 5: Sequences of qPCR primers and RT primer for antisense C9. Supplementary Table 6: Demographic information for Answer ALS and NeuroLINCS samples. Supplementary Table 7: High-throughput sequencing sample information.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 5
Unprocessed western blots.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 1
Unprocessed western blots.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 2 and Fig. 9
Unprocessed western blots.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 7
Unprocessed western blots.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 8
Unprocessed western blots.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Source Data Extended Data Fig. 10
Unprocessed western blots.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, Y., Dou, X., Liu, J. et al. Globally reduced N6-methyladenosine (m6A) in C9ORF72-ALS/FTD dysregulates RNA metabolism and contributes to neurodegeneration. Nat Neurosci 26, 1328–1338 (2023). https://doi.org/10.1038/s41593-023-01374-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-023-01374-9
This article is cited by
-
Molecular hallmarks of ageing in amyotrophic lateral sclerosis
Cellular and Molecular Life Sciences (2024)
-
Dysregulated N6-methyladenosine modification in peripheral immune cells contributes to the pathogenesis of amyotrophic lateral sclerosis
Frontiers of Medicine (2024)
-
Single-molecule imaging reveals distinct elongation and frameshifting dynamics between frames of expanded RNA repeats in C9ORF72-ALS/FTD
Nature Communications (2023)