Abstract
The role of different cell types and their interactions in Alzheimer’s disease (AD) is a complex and open question. Here, we pursued this question by assembling a high-resolution cellular map of the aging frontal cortex using single-nucleus RNA sequencing of 24 individuals with a range of clinicopathologic characteristics. We used this map to infer the neocortical cellular architecture of 638 individuals profiled by bulk RNA sequencing, providing the sample size necessary for identifying statistically robust associations. We uncovered diverse cell populations associated with AD, including a somatostatin inhibitory neuronal subtype and oligodendroglial states. We further identified a network of multicellular communities, each composed of coordinated subpopulations of neuronal, glial and endothelial cells, and we found that two of these communities are altered in AD. Finally, we used mediation analyses to prioritize cellular changes that might contribute to cognitive decline. Thus, our deconstruction of the aging neocortex provides a roadmap for evaluating the cellular microenvironments underlying AD and dementia.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
snRNA-seq data were deposited in the AD Knowledge Portal (https://www.synapse.org/#!Synapse:syn16780177) and processed data can be browsed online at: https://vmenon.shinyapps.io/rosmap_snrnaseq24/. The bulk RNA-seq dataset can be accessed through the Synapse database (https://www.synapse.org/#!Synapse:syn3388564). The proteomic dataset can be accessed through the Synapse database (https://www.synapse.org/#!Synapse:syn17015098). All datasets are available for general research use according to the following requirements for data access and data attribution: https://adknowledgeportal.synapse.org/#/DataAccess/Instructions. Other ROSMAP resources can be requested at the RADC Resource Sharing Hub at https://www.radc.rush.edu. Source data are provided with this paper.
Code availability
The code for the CelMod deconvolution algorithm is available at: https://github.com/MenonLab/Celmod. Other code used within this study is available at: https://github.com/naomihabiblab/HumanDLPFC24.
References
Mostafavi, S. et al. A molecular network of the aging human brain provides insights into the pathology and cognitive decline of Alzheimer’s disease. Nat. Neurosci. 21, 811–819 (2018).
Wan, Y.-W. et al. Meta-analysis of the Alzheimer’s disease human brain transcriptome and functional dissection in mouse models. Cell Rep. 32, 107908 (2020).
Mathys, H. et al. Single-cell transcriptomic analysis of Alzheimer’s disease. Nature 570, 332–337 (2019).
Del-Aguila, J. L. et al. A single-nuclei RNA sequencing study of Mendelian and sporadic AD in the human brain. Alzheimers Res. Ther. 11, 71 (2019).
Grubman, A. et al. A single-cell atlas of entorhinal cortex from individuals with Alzheimer’s disease reveals cell-type-specific gene expression regulation. Nat. Neurosci. 22, 2087–2097 (2019).
Zhou, Y. et al. Human and mouse single-nucleus transcriptomics reveal TREM2-dependent and TREM2-independent cellular responses in Alzheimer’s disease. Nat. Med. 26, 131–142 (2020).
Olah, M. et al. Single cell RNA sequencing of human microglia uncovers a subset associated with Alzheimer’s disease. Nat. Commun. https://doi.org/10.1038/s41467-020-19737-2 (2020).
Bennett, D. A. et al. Religious Orders Study and Rush Memory and Aging Project. J. Alzheimers Dis. 64, S161–S189 (2018).
Bennett, D. A. et al. Overview and findings from the Rush Memory and Aging Project. Curr. Alzheimer Res. 9, 646–663 (2012).
Bennett, D. A., Schneider, J. A., Arvanitakis, Z. & Wilson, R. S. Overview and findings from the Religious Orders Study. Curr. Alzheimer Res. 9, 628–645 (2012).
De Jager, P. L. et al. A multi-omic atlas of the human frontal cortex for aging and Alzheimer’s disease research. Sci. Data 5, 180142 (2018).
Hodge, R. D. et al. Conserved cell types with divergent features in human versus mouse cortex. Nature 573, 61–68 (2019).
Thrupp, N. et al. Single-nucleus RNA-seq is not suitable for detection of microglial activation genes in humans. Cell Rep. 32, 108189 (2020).
Bakken, T. E. et al. Single-nucleus and single-cell transcriptomes compared in matched cortical cell types. PLoS ONE 13, e0209648 (2018).
Habib, N. et al. Disease-associated astrocytes in Alzheimer’s disease and aging. Nat. Neurosci. 23, 701–706 (2020).
Girgrah, N. et al. Localization of the CD44 glycoprotein to fibrous astrocytes in normal white matter and to reactive astrocytes in active lesions in multiple sclerosis. J. Neuropathol. Exp. Neurol. 50, 779–792 (1991).
Leitão, R. A. et al. Aquaporin-4 as a new target against methamphetamine-induced brain alterations: focus on the neurogliovascular unit and motivational behavior. Mol. Neurobiol. 55, 2056–2069 (2018).
Yang, A. C. et al. A human brain vascular atlas reveals diverse mediators of Alzheimera’s risk. Nature 603, 885–892 (2022).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Blei, D. M., Ng, A. & Jordan, M. I. Latent Dirichlet allocation. J. Mach. Learn. Res. 3, 993–1022 (2003).
Dey, K. K., Hsiao, C. J. & Stephens, M. Visualizing the structure of RNA-seq expression data using grade of membership models. PLoS Genet. 13, e1006599 (2017).
Bielecki, P. et al. Skin-resident innate lymphoid cells converge on a pathogenic effector state. Nature 592, 128–132 (2021).
Walker, K. A. et al. Large-scale plasma proteomic analysis identifies proteins and pathways associated with dementia risk. Nat. Aging 1, 473–489 (2021).
Foster, E. M., Dangla-Valls, A., Lovestone, S., Ribe, E. M. & Buckley, N. J. Clusterin in Alzheimer’s disease: mechanisms, genetics, and lessons from other pathologies. Front. Neurosci. https://doi.org/10.3389/fnins.2019.00164 (2019).
Leng, K. et al. Molecular characterization of selectively vulnerable neurons in Alzheimer’s disease. Nat. Neurosci. 24, 276–287 (2021).
Altboum, Z. et al. Digital cell quantification identifies global immune cell dynamics during influenza infection. Mol. Syst. Biol. 10, 720 (2014).
Gong, T. & Szustakowski, J. D. DeconRNASeq: a statistical framework for deconvolution of heterogeneous tissue samples based on mRNA-Seq data. Bioinformatics 29, 1083–1085 (2013).
Hunt, G. J., Freytag, S., Bahlo, M. & Gagnon-Bartsch, J. A. dtangle: accurate and robust cell type deconvolution. Bioinformatics 35, 2093–2099 (2019).
Patrick, E. et al. Deconvolving the contributions of cell-type heterogeneity on cortical gene expression. PLoS Comput. Biol. 16, e1008120 (2020).
Johnson, E. C. B. et al. Large-scale proteomic analysis of Alzheimer’s disease brain and cerebrospinal fluid reveals early changes in energy metabolism associated with microglia and astrocyte activation. Nat. Med. 26, 769–780 (2020).
Edfors, F. et al. Gene-specific correlation of RNA and protein levels in human cells and tissues. Mol. Syst. Biol. 12, 883 (2016).
Farfel, J. M., Yu, L., De Jager, P. L., Schneider, J. A. & Bennett, D. A. Association of APOE with tau-tangle pathology with and without β-amyloid. Neurobiol. Aging 37, 19–25 (2016).
Bennett, D. A. et al. Amyloid mediates the association of apolipoprotein E e4 allele to cognitive function in older people. J. Neurol. Neurosurg. Psychiatry 76, 1194–1199 (2005).
Neff, R. A. et al. Molecular subtyping of Alzheimer’s disease using RNA sequencing data reveals novel mechanisms and targets. Sci. Adv. 7, eabb5398 (2021).
Jha, M. K., Jo, M., Kim, J.-H. & Suk, K. Microglia-astrocyte crosstalk: an intimate molecular conversation. Neuroscientist 25, 227–240 (2019).
Vainchtein, I. D. & Molofsky, A. V. Astrocytes and microglia: in sickness and in health. Trends Neurosci. 43, 144–154 (2020).
Qiao, W. et al. Intercellular network structure and regulatory motifs in the human hematopoietic system. Mol. Syst. Biol. 10, 741 (2014).
Ramilowski, J. A. et al. A draft network of ligand–receptor-mediated multicellular signalling in human. Nat. Commun. https://doi.org/10.1038/ncomms8866 (2015).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. CellPhoneDB: inferring cell–cell communication from combined expression of multi-subunit ligand–receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
West, D. C. et al. Interactions of multiple heparin binding growth factors with neuropilin-1 and potentiation of the activity of fibroblast growth factor-2. J. Biol. Chem. 280, 13457–13464 (2005).
Jack, C. R. Jr et al. Hypothetical model of dynamic biomarkers of the Alzheimer’s pathological cascade. Lancet Neurol. 9, 119–128 (2010).
Selkoe, D. J. & Hardy, J. The amyloid hypothesis of Alzheimer’s disease at 25 years. EMBO Mol. Med. 8, 595–608 (2016).
Hanseeuw, B. J. et al. Association of amyloid and tau with cognition in preclinical Alzheimer disease: a longitudinal study. JAMA Neurol. 76, 915–924 (2019).
Sperling, R. A. et al. The impact of amyloid-beta and tau on prospective cognitive decline in older individuals. Ann. Neurol. 85, 181–193 (2019).
Jennrich, R. I. An asymptotic Χ2 test for the equality of two correlation matrices. J. Am. Stat. Assoc. 65, 904 (1970).
Bennett, D. A. et al. Neuropathology of older persons without cognitive impairment from two community-based studies. Neurology https://doi.org/10.1212/01.wnl.0000219668.47116.e6 (2006).
Wilson, R. S. et al. Individual differences in rates of change in cognitive abilities of older persons. Psychol. Aging 17, 179–193 (2002).
Wilson, R. S., Barnes, L. L. & Bennett, D. A. Assessment of lifetime participation in cognitively stimulating activities. J. Clin. Exp. Neuropsychol. 25, 634–642 (2003).
Wilson, R. S. et al. Conscientiousness, dementia related pathology, and trajectories of cognitive aging. Psychol. Aging 30, 74–82 (2015).
De Jager, P. L. et al. A genome-wide scan for common variants affecting the rate of age-related cognitive decline. Neurobiol. Aging 33, 1017.e1–1017e.15 (2012).
McKhann, G. et al. Clinical diagnosis of Alzheimer’s disease: report of the NINCDS-ADRDA Work Group under the auspices of Department of Health and Human Services Task Force on Alzheimer’s Disease. Neurology 34, 939–944 (1984).
Schneider, J. A., Arvanitakis, Z., Bang, W. & Bennett, D. A. Mixed brain pathologies account for most dementia cases in community-dwelling older persons. Neurology 69, 2197–2204 (2007).
Bennett, D. A. et al. Natural history of mild cognitive impairment in older persons. Neurology 59, 198–205 (2002).
Bennett, D. A. et al. Decision rules guiding the clinical diagnosis of Alzheimer’s disease in two community-based cohort studies compared to standard practice in a clinic-based cohort study. Neuroepidemiology 27, 169–176 (2006).
Hyman, B. T. & Trojanowski, J. Q. Editorial on consensus recommendations for the postmortem diagnosis of Alzheimer disease from the National Institute on Aging and the Reagan Institute Working Group on Diagnostic Criteria for the Neuropathological Assessment of Alzheimer Disease. J. Neuropathol. Exp. Neurol. 56, 1095–1097 (1997).
Habib, N. et al. Massively parallel single-nucleus RNA-seq with DroNc-seq. Nat. Methods 14, 955–958 (2017).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
McInnes, L., Healy, J., Saul, N. & Großberger, L. UMAP: uniform manifold approximation and projection. J. Open Source Softw. 3, 861 (2018).
Lancichinetti, A. & Fortunato, S. Benchmarks for testing community detection algorithms on directed and weighted graphs with overlapping communities. Phys. Rev. E Stat. Nonlin. Soft Matter Phys. 80, 016118 (2009).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 8, 329–337.e4 (2019).
Liberzon, A. et al. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
The National Institute on Aging, and Reagan Institute Working Group on Diagnostic Criteria for the Neuropathological Assessment of Alzheimer’s Disease. Consensus recommendations for the postmortem diagnosis of Alzheimer’s disease. Neurobiol. Aging 18, S1–S2 (1997).
MacKinnon, D. P., Krull, J. L. & Lockwood, C. M. Equivalence of the mediation, confounding and suppression effect. Prev. Sci. 1, 173–181 (2000).
Tingley, D., Yamamoto, T., Hirose, K., Keele, L. & Imai, K. mediation: R package for causal mediation analysis. J. Stat. Softw. 59, 1–38 (2014).
Acknowledgements
We thank the individuals who have generously donated their brain to research through the RUSH University Alzheimer’s Disease Center. This work was supported by the National Institute of Aging (NIA), grant nos. AMP-AD U01AG046152, U01AG061356 and RF1AG036042 (P.L.D. and D.A.B); grant nos. R01AG066831 and R21AG075754 and SenNet (grant no. U54AG076040) (V.M.); grant nos. P30AG10161, P30AG72975, R01AG15819 and R01AG17917 (ROSMAP, P.L.D.); the Chan Zuckerberg Initiative (grant no. CS-02018-191971) (P.L.D. and V.M.); Israel Science Foundation (ISF) grant no. 1709/19 (N.H.); the European Research Council grant no. 853409 (N.H.); grant no. MOST/IL 3-15687 (N.H.); and the Klarman Cell Observatory (A.R.). N.H. is a Goren Khazzam senior lecturer in neuroscience and is support by the Myers Foundation. F.Z. and A.R. are investigators of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
P.L.D. and N.H. designed the study. C.M. and N.H. prepared the single-nucleus libraries and performed sequencing. The majority of the computational analysis was performed by A.C.; statistical and modeling analysis by H.-S.Y. and C.C.W.; and additional analysis by G.S.G., D.I.L. and P.G., with the guidance of N.H., V.M. and P.L.D. I.H. performed analysis of cellular interactions under the guidance of E.Y.-L. M.T. performed the spatial transcriptomics. P.L.D., N.H., V.M., A.C., D.A.B. and A.R. wrote the manuscript. O.R.-R., F.Z. and all coauthors edited the manuscript for critical comments. D.A.B. is the principal investigator of the parent ROS and MAP studies. P.L.D., D.A.B. and N.H. obtained funding for the study and deposited data in Synapse and the RADC Resource Sharing Hub.
Corresponding authors
Ethics declarations
Competing interests
A.R. is a founder and equity holder of Celsius Therapeutics, an equity holder in Immunitas Therapeutics and until 31 August 2020 was an SAB member of Syros Pharmaceuticals, Neogene Therapeutics, Asimov and ThermoFisher Scientific. From 1 August 2020, A.R. is an employee of Genentech, a member of the Roche Group. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Neuroscience thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 A cellular-molecular map of the human aging DLPFC: Quality controls.
(a) High quality nuclei libraries generated across n = 24 post-mortem independent samples of the DLPFC brain region of aging individuals. Nuclei (dots) colored by the doublet score (Methods). (b) Distribution of cell type frequencies across batches. The fraction of nuclei (y-axis) per cell type for each batch (x-axis, n = 3 batches). (c, d) Distribution of number of genes (c) and transcripts (d) across the 9 major cell types in the DLPFC. Violin plots showing the distribution per cell type (for n = 172,659 nuclei). (e, f) Distribution of cell type frequencies across sex (e, n = 24 independent samples, 12 per group) and archetypes of AD (f, n = 24, 6 per group). Boxplot showing the fraction of nuclei per cell type for males (blue, n = 12) and females (red, n = 12). For box plots, the bottom and upper borders show the first and third quartiles. The central line indicates the median. The whiskers are extended to the extrema values (without accounting for outliers). Dots show individual samples. Archetypes defined as in Fig. 1a, b.
Extended Data Fig. 2 Quality controls of neuronal subtypes and their cortical layer specificity.
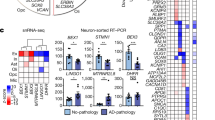
(a) Distinct expression of known and de novo marker genes in excitatory neuronal subtypes (top) and inhibitory neuronal subtypes (bottom) as assigned by our clustering analysis. Mean expression level in expressing cells (color) and percent of expressing cells (circle size) of selected markers in each neuronal subtype (rows) of marker genes. (b-c) Distribution of neuronal subtype frequencies across sex or batch for n = 24 independent sample. Boxplots showing the fraction of nuclei per neuronal subtype for sex (b, n = 12 per group) or batch (c, n = 8 per group): For box plots, the bottom and upper borders show the first and third quartiles. The central line indicates the median. The whiskers are extended to the extrema values (without accounting for outliers). Dots show outliers samples. (d) Marker genes of neuronal subtypes exhibit a spatial organization at distinct layers within DLPFC slices. Spatial transcriptomics across 6 slices from 3 individuals for five marker genes (RORB, TOX, CUX2, PVALB, SLC17A7). Note variable orientation of slices. Complementary images to Fig. 2c.
Extended Data Fig. 3 Quality controls of glial subsets.
(a) Distribution of number of genes across microglia, astrocytes and endothelial clusters (denoted as subsets). (b, c) Distribution of non-neuronal subsets frequencies across sex and batch for n = 24 independent samples. Boxplot showing the fraction of nuclei per neuronal subtype for sex (b, n = 12 per group) or batch (c, n = 8 per group): For box plots, the bottom and upper borders show the first and third quartiles. The central line indicates the median. The whiskers are extended to the extrema values (without accounting for outliers). Dots show outlier samples. (d) RIN, sex and batch do not affect the sub-clustering of astrocytes, microglia and endothelial cells. Heatmaps of Jaccard score comparing overlaps of cells (color scales) between assignment of de-novo clusters after regression of the confounding variables from the expression matrix (rows) compared to the clusters in this study without such correction (columns) (Methods). (e) Endothelial subsets express unique markers. Dot plot of the mean expression level in expressing cells (color) and percent of expressing cells (circle size) of selected marker genes across endothelial subsets.
Extended Data Fig. 4 Comparison of cell clusters to previous studies.
(a) Clusters of microglia nuclei from snRNA-seq match published live microglia cell clusters from scRNA-seq. The proportions (color scale, scaled per column) of nuclei per cluster (columns) mapped to each scRNA-seq cell cluster according to the best prediction (rows, Methods). (b) Mean expression level in expressing cells (color) and percent of expressing cells (circle size) across cell subsets (rows) of previously described up-regulated and down-regulated genes in AD brains compared to healthy individuals, as defined by Zhou et al.6 for astrocytes (left) and oligodendrocytes (right). (c) Nuclear-derived model is consistent with earlier, lower-resolution models across different cell types. Heatmaps (color scale) of assignment of nuclei from 24 individuals (rows) to published subsets (Methods) from 4 previous snRNA-seq derived cortical models3,5,6,25.
Extended Data Fig. 5 Spatial transcriptomics of glial markers.
Spatial transcriptomics of glial cell type and cell states markers, exhibit a spatial pattern across cortical layers matching DLPFC and white-matter anatomy. MBP (oligodendrocyte marker), GFAP (reactive astrocyte marker), ID3, CD44 (Ast.3 marker). Complementary to Fig. 3g.
Extended Data Fig. 6 CelMod evaluation and comparison to other methods and datasets.
(a, b) CelMod estimated proportions match snRNA-seq measured proportions. Scatter plots of CelMod proportions (Y-axis) compared to snRNA-seq proportions (X-axis), for 6 major cell classes (a) and all cell subsets and topic models (b, colored by subset). Line: linear regression. Each point is a sample (n = 24 independent samples). (c) Validation of CelMod in an independent dataset. CelMod estimated proportions of cell subsets (Y-axis) compared to snRNA-seq proportions (X-axis) measured across n = 48 independent samples of the prefrontal cortex Brodmann area 10 (Mathys et al.3). Each point is a single individual. Cell subset annotations are based on our cell atlas (Methods). R=Correlation. (d) Protein expression of selected markers reflect cell subsets abundance. Scaled protein expression levels (X-axis) compared to CelMod estimated proportions of the related cell subset, colored by the bulk RNA-seq (proteomics30 in n = 196 independent individuals with matching bulk RNA profiles). Each point is a single individual. Line: linear regression fit with confidence interval (grey). (e) CelMod outperforms previous methods. Spearman correlation scores (color scale) of the snRNA-seq measured proportions of each cell type (left) and cell subset (right), compared to the estimated proportions by CelMod and three previous models. (f) Correlations between AD traits within our data. Pairwise correlations (color scale) of the three traits across n = 638 independent individuals. (g) Measured proportions of cellular subsets from snRNA-seq in n = 24 individuals correlate with AD pathology and cognitive decline. Correlation (color scale) of the proportions of each cell subset (columns) to AD-traits (rows). The proportions are calculated over the total number of nuclei per individual (n = 24) within each cell. (h) Cellular proportions associations to AD-ranking in the independent MSBB32 cohort matches the ROSMAP1 cohort. Association scores −log(FDR) × sign(β), by multivariable linear regression) of proportions of cell subsets to two measures of AD ranking: BRAAK stage (tangles load) and the CDR (level of cognitive decline), in n = 106 independent samples32 (Methods). Cell subsets colored by the statistical significance of associations to the cognitive decline rate in the ROSMAP cohort1 (n = 638).
Extended Data Fig. 7 Evaluation of multi-cellular communities.
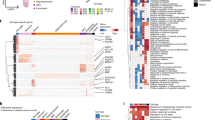
(a) Similar structure of coordinated changes between proportions of cell subsets and neuronal subtypes across individuals found in two independent cohorts. Pairwise Spearman correlation coefficients of the proportions of cell states and subtypes across individuals estimated by CelMod in bulk RNA-seq data of ROSMAP1 (left) and MSBB32 (right) cohorts. Bottom: Correlation between the pairwise association pattern of the ROSMAP and the MSBB cohorts per cell subset. (b) A network of cellular subsets reveals coordinated variation across individuals in multiple cell types. Network of coordinated and anti-coordinated cell subsets (nodes). Edges between pairs of subsets with statistically significant correlated proportions across individuals (r > 0.4, p-value threshold=0.05, solid red line) or anti-correlated (r < −0.4, dashed blue line) based on snRNA-seq proportions (n = 24 independent samples, Spearman correlation, two sided, not adjusted for multiple comparison). Celmod based network in n = 638 individuals in Fig. 6b. Nodes are colored by the cell type and numbered by the subset as in Fig. 2a and Fig. 3a, d, h). (c) Coordinated changes in proportions of cell states and subtypes across individuals is independent of the cortical layer, except for excitatory neurons. Pairwise Spearman correlation coefficient of the CelMod proportions of all cell subsets and neuronal subtypes across individuals with low levels of Exc.1 (n = 371 individuals, left) or high levels of Exc.1 (n = 267 individuals, middle). Right: The differences in pairwise correlations between the Exc.1-high and Exc.1-low groups of individuals. Showing the partition mainly affects excitatory neurons (red). (d) Shared pathways within the cognitive non-impaired community. Enriched pathways (hypergeometric test, FDR q-value < 0.05, blue) in up-regulated genes for each cell subset within the cognitively non-impaired community (End.1, Oli.1, Ast.2, and Inh.3). Displaying shared enriched pathways between at least three subsets.
Extended Data Fig. 8 Signaling within and between multi-cellular communities and causal modeling.
(a) Cell subsets positively associated with cognitive decline have an increased expression of ligand-receptor pairs compared to the negatively associated subsets. For each pair of cell subsets showing the number (color scale) of ligand-receptor pairs (LRP, row: ligand, column: receptor) where both the ligand and the receptor are differentially expressed in the relevant subsets. Top and side bar marking: subsets positively (purple) or negatively (turquoise) associated with cognitive decline, and the total number of ligands and receptors expressed (color scale). (b) Expression of the HLA-A - APLP2 ligand-receptor pair across subtypes of different cell types. Dot plot of the mean expression level in expressing cells (color) and percent of expressing cells (circle size) of HLA-A and of APLP2 across subsets of selected cell types. (c) Mediation analysis results showing tangle pathology burden (tau) is predicted to be upstream of changes in proportion of Inh.3, Oli.1, Oli.2, Ast.4, and End.2, but not Inh.2. (d) Mediation analysis results showing partial effect of changes in proportion of Inh.3, Olig.1, Ast.4, and End.2 on cognitive decline independent of tau pathology burden.
Supplementary information
Supplementary Table 1
Supplementary Tables 1–5.
Source data
Source Data Fig. 1
Tab 1. Metadata of participants in the study and summary of metadata. Tab 2. Coordinates of single nuclei in 2D embedding. Tab 3. Cell type annotations per nuclei.
Source Data Fig. 2
Tab 1. Cell subtype annotations per nuclei. Tab 2. Raw output of classifier predicting cortical layer per nuclei of excitatory neurons. Tab 3. Raw output of classifier predicting cortical layer per nuclei of inhibitory neurons.
Source Data Fig. 3
Tab 1. Cell subset annotations per nuclei and topic modeling score for oligodendrocyte nuclei. Tab 2. Raw output of pathway enrichment within differential genes of cell subsets.
Source Data Fig. 4
Tab 1. CelMod output of estimated proportions and snRNA-seq measured proportions per cell subset for 24 individuals. Tab 2. CelMod output of proportions for 48 individuals from the Mathys et al. dataset. Tab 3. Output of regression analysis associated cell subset proportions to AD traits. Tab 4. Expression values of marker proteins per individual.
Source Data Fig. 5
CelMod output of estimated proportions per cell subset and per individual across 638 individuals.
Source Data Fig. 6
Tab 1. CelMod output of estimated proportions per cell subset and per individual across 638 individuals. Tab 2. snRNA-seq measured proportions of cell subset per individual. Tab 3. Output of regression analysis associated cell subset proportions to AD traits.
Source Data Fig. 7
Tab 1. Raw output of pathway enrichment within differential genes of cell subsets. Tab 2. Output of CellPhoneDB predicting statistically significant ligand–receptor pairs (LRPs). Tab 3. Output of mediation analysis for key cell subsets and AD traits.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Cain, A., Taga, M., McCabe, C. et al. Multicellular communities are perturbed in the aging human brain and Alzheimer’s disease. Nat Neurosci 26, 1267–1280 (2023). https://doi.org/10.1038/s41593-023-01356-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-023-01356-x
This article is cited by
-
An integrated toolkit for human microglia functional genomics
Stem Cell Research & Therapy (2024)
-
The concept of resilience to Alzheimer’s Disease: current definitions and cellular and molecular mechanisms
Molecular Neurodegeneration (2024)
-
Molecular and cellular mechanisms of selective vulnerability in neurodegenerative diseases
Nature Reviews Neuroscience (2024)
-
CHRNA5 links chandelier cells to severity of amyloid pathology in aging and Alzheimer’s disease
Translational Psychiatry (2024)
-
Transcriptomics of Human Brain Tissue in Parkinson’s Disease: a Comparison of Bulk and Single-cell RNA Sequencing
Molecular Neurobiology (2024)