Abstract
The extent to which neurogenesis occurs in adult primates remains controversial. In this study, using an optimized single-cell RNA sequencing pipeline, we profiled 207,785 cells from the adult macaque hippocampus and identified 34 cell populations comprising all major hippocampal cell types. Analysis of their gene expression, specification trajectories and gene regulatory networks revealed the presence of all key neurogenic precursor cell populations, including a heterogeneous pool of radial glia-like cells (RGLs), intermediate progenitor cells (IPCs) and neuroblasts. We identified HMGB2 as a novel IPC marker. Comparison with mouse single-cell transcriptomic data revealed differences in neurogenic processes between species. We confirmed that neurogenesis is recapitulated in ex vivo neurosphere cultures from adult primates, further supporting the existence of neural precursor cells (NPCs) that are able to proliferate and differentiate. Our large-scale dataset provides a comprehensive adult neurogenesis atlas for primates.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The datasets generated in this paper are publicly available at ArrayExpress under accession codes E-MTAB-10225 and E-MTAB-10236. Source data are provided with this paper.
Code availability
The code for generating figures can be found at https://github.com/haozhaozhe/FM_hippo.
References
Gonçalves, J. T., Schafer, S. T. & Gage, F. H. Adult neurogenesis in the hippocampus: from stem cells to behavior. Cell 167, 897–914 (2016).
Kaplan, M. S. & Hinds, J. W. Neurogenesis in the adult rat: electron microscopic analysis of light radioautographs. Science 197, 1092–1094 (1977).
Doetsch, F., Caillé, I., Lim, D. A., García-Verdugo, J. M. & Alvarez-Buylla, A. Subventricular zone astrocytes are neural stem cells in the adult mammalian brain. Cell 97, 703–716 (1999).
Altman, J. & Das, G. D. Autoradiographic and histological evidence of postnatal hippocampal neurogenesis in rats. J. Comp. Neurol. 124, 319–335 (1965).
Yang, C.-H. et al. Circuit integration initiation of new hippocampal neurons in the adult brain. Cell Rep. 30, 959–968 (2020).
Song, J., Olsen, R. H. J., Sun, J., Ming, G.-L. & Song, H. Neuronal circuitry mechanisms regulating adult mammalian neurogenesis. Cold Spring Harb. Perspect. Biol. 8, a018937 (2016).
Jessberger, S. et al. Dentate gyrus-specific knockdown of adult neurogenesis impairs spatial and object recognition memory in adult rats. Learn. Mem. 16, 147–154 (2009).
Kempermann, G., Georg Kuhn, H. & Gage, F. H. More hippocampal neurons in adult mice living in an enriched environment. Nature 386, 493–495 (1997).
van Praag, H., Kempermann, G. & Gage, F. H. Running increases cell proliferation and neurogenesis in the adult mouse dentate gyrus. Nat. Neurosci. 2, 266–270 (1999).
Livneh, Y., Adam, Y. & Mizrahi, A. Odor processing by adult-born neurons. Neuron 81, 1097–1110 (2014).
Eriksson, P. S. et al. Neurogenesis in the adult human hippocampus. Nat. Med. 4, 1313–1317 (1998).
Spalding, K. L. et al. Dynamics of hippocampal neurogenesis in adult humans. Cell 153, 1219–1227 (2013).
Gould, E. et al. Hippocampal neurogenesis in adult Old World primates. Proc. Natl Acad. Sci. USA 96, 5263–5267 (1999).
Kornack, D. R. & Rakic, P. Continuation of neurogenesis in the hippocampus of the adult macaque monkey. Proc. Natl Acad. Sci. USA 96, 5768–5773 (1999).
Yuan, T.-F., Li, J., Ding, F. & Arias-Carrion, O. Evidence of adult neurogenesis in non-human primates and human. Cell Tissue Res. 358, 17–23 (2014).
Ngwenya, L. B., Heyworth, N. C., Shwe, Y., Moore, T. L. & Rosene, D. L. Age-related changes in dentate gyrus cell numbers, neurogenesis, and associations with cognitive impairments in the rhesus monkey. Front. Syst. Neurosci. 9, 102 (2015).
Sorrells, S. F. et al. Human hippocampal neurogenesis drops sharply in children to undetectable levels in adults. Nature 555, 377–381 (2018).
Moreno-Jiménez, E. P. et al. Adult hippocampal neurogenesis is abundant in neurologically healthy subjects and drops sharply in patients with Alzheimer’s disease. Nat. Med. 25, 554–560 (2019).
Kempermann, G. et al. Human adult neurogenesis: evidence and remaining questions. Cell Stem Cell 23, 25–30 (2018).
Tobin, M. K. et al. Human hippocampal neurogenesis persists in aged adults and Alzheimer’s disease patients. Cell Stem Cell 24, 974–982 (2019).
Boldrini, M. et al. Human hippocampal neurogenesis persists throughout aging. Cell Stem Cell 22, 589–599 (2018).
Terreros-Roncal, J. et al. Impact of neurodegenerative diseases on human adult hippocampal neurogenesis. Science 374, 1106–1113 (2021).
Aizawa, K., Ageyama, N., Terao, K. & Hisatsune, T. Primate-specific alterations in neural stem/progenitor cells in the aged hippocampus. Neurobiol. Aging 32, 140–150 (2011).
Seki, T. Understanding the real state of human adult hippocampal neurogenesis from studies of rodents and non-human primates. Front. Neurosci. 14, 839 (2020).
Curtis, M. A. et al. Human neuroblasts migrate to the olfactory bulb via a lateral ventricular extension. Science 315, 1243–1249 (2007).
Sanai, N. et al. Corridors of migrating neurons in the human brain and their decline during infancy. Nature 478, 382–386 (2011).
Snyder, J. S. Recalibrating the relevance of adult neurogenesis. Trends Neurosci. 42, 164–178 (2019).
Harris, L. et al. Coordinated changes in cellular behavior ensure the lifelong maintenance of the hippocampal stem cell population. Cell Stem Cell 28, 863-876 (2021).
Artegiani, B. et al. A single-cell RNA sequencing study reveals cellular and molecular dynamics of the hippocampal neurogenic niche. Cell Rep. 21, 3271–3284 (2017).
Hochgerner, H., Zeisel, A., Lönnerberg, P. & Linnarsson, S. Conserved properties of dentate gyrus neurogenesis across postnatal development revealed by single-cell RNA sequencing. Nat. Neurosci. 21, 290–299 (2018).
Huang, W. et al. Origins and proliferative states of human oligodendrocyte precursor cells. Cell 182, 594–608 (2020).
Kriegstein, A. & Alvarez-Buylla, A. The glial nature of embryonic and adult neural stem cells. Annu. Rev. Neurosci. 32, 149–184 (2009).
Shin, J. et al. Single-cell RNA-seq with waterfall reveals molecular cascades underlying adult neurogenesis. Cell Stem Cell 17, 360–372 (2015).
Suh, H. et al. In vivo fate analysis reveals the multipotent and self-renewal capacities of Sox2 neural stem cells in the adult hippocampus. Cell Stem Cell 1, 515–528 (2007).
Berg, D. A. et al. A common embryonic origin of stem cells drives developmental and adult neurogenesis. Cell 177, 654–668 (2019).
Llorens-Bobadilla, E. et al. Single-cell transcriptomics reveals a population of dormant neural stem cells that become activated upon brain injury. Cell Stem Cell 17, 329–340 (2015).
Joglekar, A. et al. A spatially resolved brain region- and cell type-specific isoform atlas of the postnatal mouse brain. Nat. Commun. 12, 463 (2021).
Zhong, S. et al. Decoding the development of the human hippocampus. Nature 577, 531–536 (2020).
Li, M. et al. Integrative functional genomic analysis of human brain development and neuropsychiatric risks. Science 362, eaat7615 (2018).
Franjic, D. et al. Transcriptomic taxonomy and neurogenic trajectories of adult human, macaque, and pig hippocampal and entorhinal cells. Neuron 110, 452–469 (2022).
Eckenhoff, M. F. & Rakic, P. Nature and fate of proliferative cells in the hippocampal dentate gyrus during the life span of the rhesus monkey. J. Neurosci. 8, 2729–2747 (1988).
Bergen, V., Lange, M., Peidli, S., Wolf, F. A. & Theis, F. J. Generalizing RNA velocity to transient cell states through dynamical modeling. Nat. Biotechnol. 38, 1408–1414 (2020).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Pilz, G.-A. et al. Live imaging of neurogenesis in the adult mouse hippocampus. Science 359, 658–662 (2018).
Bonaguidi, M. A. et al. In vivo clonal analysis reveals self-renewing and multipotent adult neural stem cell characteristics. Cell 145, 1142–1155 (2011).
Bonaguidi, M. A., Song, J., Ming, G.-L. & Song, H. A unifying hypothesis on mammalian neural stem cell properties in the adult hippocampus. Curr. Opin. Neurobiol. 22, 754–761 (2012).
Miller, J. A. et al. Conserved molecular signatures of neurogenesis in the hippocampal subgranular zone of rodents and primates. Development 140, 4633–4644 (2013).
Denoth-Lippuner, A. & Jessberger, S. Formation and integration of new neurons in the adult hippocampus. Nat. Rev. Neurosci. 22, 223–236 (2021).
Oddi, S., Scipioni, L. & Maccarrone, M. Endocannabinoid system and adult neurogenesis: a focused review. Curr. Opin. Pharmacol. 50, 25–32 (2020).
Hodge, R. D. et al. Conserved cell types with divergent features in human versus mouse cortex. Nature 573, 61–68 (2019).
Gayoso, A. et al. A Python library for probabilistic analysis of single-cell omics data. Nat. Biotechnol. 40, 163–166 (2022).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Guo, Z. et al. RGCC balances self-renewal and neuronal differentiation of neural stem cells in the developing mammalian neocortex. EMBO Rep. 22, e51781 (2021).
Adusumilli, V. S. et al. ROS dynamics delineate functional states of hippocampal neural stem cells and link to their activity-dependent exit from quiescence. Cell Stem Cell 28, 300–314 (2021).
Sun, J., Sun, J., Ming, G.-L. & Song, H. Epigenetic regulation of neurogenesis in the adult mammalian brain. Eur. J. Neurosci. 33, 1087–1093 (2011).
Kimura, A., Matsuda, T., Sakai, A., Murao, N. & Nakashima, K. HMGB2 expression is associated with transition from a quiescent to an activated state of adult neural stem cells. Dev. Dyn. 247, 229–238 (2018).
Augusto-Oliveira, M., Arrifano, G. P. F., Malva, J. O. & Crespo-Lopez, M. E. Adult hippocampal neurogenesis in different taxonomic groups: possible functional similarities and striking controversies. Cells 8, 125 (2019).
Gould, E., Vail, N., Wagers, M. & Gross, C. G. Adult-generated hippocampal and neocortical neurons in macaques have a transient existence. Proc. Natl Acad. Sci. USA 98, 10910–10917 (2001).
Urbán, N. et al. Return to quiescence of mouse neural stem cells by degradation of a proactivation protein. Science 353, 292–295 (2016).
Bottes, S. et al. Long-term self-renewing stem cells in the adult mouse hippocampus identified by intravital imaging. Nat. Neurosci. 24, 225–233 (2021).
Jia, J., Cui, Y., Tan, Z., Liu, M. & Jiang, Y. Transcriptional factor FoxM1-activated microRNA-335-3p maintains the self-renewal of neural stem cells by inhibiting p53 signaling pathway via Fmr1. Stem Cell Res. Ther. 12, 169 (2021).
DeCarolis, N. A. et al. In vivo contribution of nestin- and GLAST-lineage cells to adult hippocampal neurogenesis. Hippocampus 23, 708–719 (2013).
Toda, T., Parylak, S. L., Linker, S. B. & Gage, F. H. The role of adult hippocampal neurogenesis in brain health and disease. Mol. Psychiatry 24, 67–87 (2019).
Saleem, K. S. & Logothetis, N. K. A Combined MRI and Histology Atlas of the Rhesus Monkey Brain in Stereotaxic Coordinates (Academic Press, 2012).
Xiao, D. et al. Direct reprogramming of fibroblasts into neural stem cells by single non-neural progenitor transcription factor Ptf1a. Nat. Commun. 9, 2865 (2018).
Yates, A. D. et al. Ensembl 2020. Nucleic Acids Res. 48, D682–D688 (2019).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, 281–291 (2019).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Van de Sande, B. et al. A scalable SCENIC workflow for single-cell gene regulatory network analysis. Nat. Protoc. 15, 2247–2276 (2020).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44 (2018).
Blondel, V. D., Guillaume, J.-L., Lambiotte, R. & Lefebvre, E. Fast unfolding of communities in large networks. J. Stat. Mech. 2008, P10008 (2008).
Miao, Z. et al. Putative cell type discovery from single-cell gene expression data. Nat. Methods 17, 621–628 (2020).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Raudvere, U. et al. g:Profiler: a web server for functional enrichment analysis and conversions of gene lists (2019 update). Nucleic Acids Res. 47, W191–W198 (2019).
Spearman, C. The proof and measurement of association between two things. Am J Psychol 15, 72 (1904).
Glasser, G. J. & Winter, R. F. Critical values of the coefficient of rank correlation for testing the hypothesis of independence. Biometrika 48, 444–448 (1961).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
Finak, G. et al. MAST: a flexible statistical framework for assessing transcriptional changes and characterizing heterogeneity in single-cell RNA sequencing data. Genome Biol. 16, 278 (2015).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 25, 25–29 (2000).
Gene Ontology Consortium. The Gene Ontology resource: enriching a GOld mine. Nucleic Acids Res. 49, D325–D334 (2021).
Kanehisa, M. & Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 28, 27–30 (2000).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28, 1947–1951 (2019).
Kanehisa, M., Furumichi, M., Sato, Y., Ishiguro-Watanabe, M. & Tanabe, M. KEGG: integrating viruses and cellular organisms. Nucleic Acids Res. 49, D545–D551 (2021).
Edgar, R., Domrachev, M. & Lash, A. E. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 30, 207–210 (2002).
Barrett, T. et al. NCBI GEO: archive for functional genomics data sets—update. Nucleic Acids Res. 41, D991–D995 (2013).
Mann, H. B. & Whitney, D. R. On a test of whether one of two random variables is stochastically larger than the other. Ann. Math. Stat. 18, 50–60 (1947).
Acknowledgements
This research is supported by research grants from the National Key R&D Program of China (2018YFA0108300), the Natural Science Foundation of China (81870682 and 81961128021), the Guangdong Provincial Key R&D Programs (2018B030335001 and 2018B030337001) and the Science and Technology Program of Guangzhou (202007030011, 202007030010 and 202007030001) to S.L.; the Natural Science Foundation of China (81721003) and the Local Innovative and Research Teams Project of Guangdong Pearl River Talents Program (2017BT01S138), CAMS Innovation Fund for Medical Sciences (2019-I2M-5-005) to Y.L.; the Natural Science Foundation of China (81970794), the Science and Technology Program of Guangzhou (201904020036) and the Technology Innovation 2030-Major Project on Brain Science and Brain-Like Computing of the Ministry of Science and Technology of China (2021ZD0202603) to M.X.; the Wellcome BioResource for a ‘Single Cell Gene Expression Atlasʼ (WT 108437/Z/15/Z) and the Open Targets grant (OTAR2067) to Z.M.; and the China Postdoctoral Science Foundation (2019M663256) to Z.-Z.H. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
S.L., Z.M., Y.L. and M.X. conceived and supervised the project. Z.-Z.H., Z.M., L.T., M.H., Y.S. and X.L. analyzed the data. J.-R.W., Z.-Z.H., R.L. and W.H. collected the tissues and performed the scRNA-seq experiment. D.X. performed the neurosphere culture experiment. J.-R.W., N.X., C.X. and Z.-Z.H. performed the immunofluorescence staining. S.L., Z.M., Y.L., M.X. and Z.-Z.H. wrote the manuscript, with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Neuroscience thanks Gerd Kempermann, Orly Lazarov and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Quality control of the dataset.
a) Distribution of the UMI counts detected in each cell in the whole dataset. b) Distribution of the unique genes detected in each cell in the whole dataset. c) UMAP colored by the 10x version and the animal ID. d) Violin plots of the UMI counts for each sample. The 10x version used for each sample is indicated by the colors. e) Violin plots of the unique genes detected for each sample. The 10x version used for each sample is indicated by the colors. f) The distribution of sample sizes for each animal. g) Proportion of different cell types present in individual animals. Cell types were colored using the same color scheme in Fig. 1d, and labeled at the bottom of panel.
Extended Data Fig. 2 The enriched gene regulatory networks (GRNs, termed regulons) in the hippocampal cell population.
a) tSNE plot derived from the regulon score, showing the neurogenic-related populations, colored by the cell types or the Gene Regulatory Network enrichment score derived from SCENIC AUcell. b) The dot plot shows the Gene Regulatory Network enrichment score derived from SCENIC AUcell. The color shows the normalized mean enrichment score, while the size of the dots shows the fraction of cells in each group that express the genes. The x-axis shows the name of the enriched regulons (Gene Regulatory Network), while the y-axis shows the cell populations in the dataset. c-f) TF regulatory networks predicted by SCENIC for the RGLs, IPCs and NBs.
Extended Data Fig. 3 The age-related shift in population distribution and gene expression.
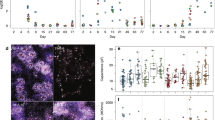
a) Sample distribution grouped by age. *, p < 0.05; using two-tailed Mann-Whitney U test for each category between the young adults (age 4–6 year-old, n = 17 biologically independent samples) and the middle-aged adults (age 13–15-year-old, n = 8 biologically independent samples). Error bars, standard error of mean. The p-value for the comparison between young and middle-aged astrocytes is 0.039; for NB, 0.0039. All other comparisons are not significantly different. b) 1,435 DEGs between the Y4–6 and Y13–15 neuroblasts. These genes are visualized using the volcano plot. Genes that are significantly enriched (blue dots, determined by a two-part generalized linear model implemented by MAST) and exhibit > 0.5 log2 fold-change are marked in red. Selective genes are marked with text. c) A strip-chart showing the logarithmic-fold change (Log2 FC) of all genes between young adult (Y4–6) and middle-aged adult (Y13–15) for neurogenic-related cell populations. The dashed line at the top and bottom of the plot indicate the 2-fold change threshold. The dashed line in the middle indicates no differential expression. Genes that are upregulated in young adults have positive FCs. Genes that are differentially expressed (Wilcoxon rank-sum test with FDR correction for multi-comparison) and >30% max FC are colored using the same color scheme in Fig. 1. d) Pair-wise comparison of the neurogenic-related gene (n = 2,081 genes) enrichment between young and middle-aged adults for selective cell populations. Solid line, equal expression; dashed lines: 2-fold enrichment. Gray dots, the mean expression of each neurogenic-related gene of all cells of each population. Blue dots, genes that show >0.5 log2 fold-change enrichment in young adults. Red dots, genes that show >0.5 log2 fold-change enrichment in middle-aged adults. All correlation with a rank-sum based Spearman’s correlation coefficient (r)>0.95 with p < 0.0001. e) Heatmaps showing the selective genes that are upregulated in young adults (upper panel) or middle-aged adults (lower panel) exclusively for selective cell populations. Nomalized FC, normalized fold-change.
Extended Data Fig.4 The sex-related shift in population distribution and gene expression.
a) Sample distribution grouped by sex. *, p < 0.05; using two-tailed Mann-Whitney U test for each category between the females (n = 7 biologically independent samples) and the males (n = 18 biologically independent samples). Error bars, standard error of mean. The p-value for the comparison between female and male IPC is 0.0015; for pre-OPC, 0.049; for PVM, 0.008. All other comparisons are not significantly different. b) Strip-chart showing the logarithmic-fold change (Log2FC) for all genes between females and males for neurogenic-related cell populations. The dashed line at the top and bottom of the plot indicate the 2-fold change threshold. The dashed line in the middle indicates not differentially expressed. Genes that are upregulated in females have potived FCs. Genes that are significantly expressed (Wilcoxon rank-sum test with FDR correction for multi-comparison) and >30% max FC are colored using the same color scheme in Fig. 1. c) Pair-wise comparison of the neurogenic-related gene (n = 2,081 genes) enrichment between females and males for selected cell populations. Solid line, equal expression; dashed lines: 2-fold enrichment. Gray dots, mean expression of each gene of all cells of each population. Blue dots, genes that show >0.5-fold enrichment in females. Red dots, genes that show >0.5-fold enrichment in males. All correlation with a rank-sum based Spearman’s correlation coefficient (r)>0.95 with p < 0.0001. d) Heatmaps showing the selective genes that are upregulated in female adults (upper panel) or male adults (lower panel) exclusively for selective cell populations. Normalized FC, normalized fold-change.
Extended Data Fig. 5 Immunostaining of canonical neural markers validated the existence of neural progenitor cells in SGZ.
a) NESTIN + cells in SGZ with apical processes (white arrowheads) touch the capillaries (yellow arrows), Scale bar, 10 μm. n = 4 brains. b) NESTIN + MKI67 + cells in SGZ with apical processes crossing the GCL into the ML, the processes touch capillaries of tubular morphology (yellow arrows). Scale bar, 10 μm. n = 4 brains. c) HOPX protein (gray), progenitor marker SOX2 (green), and neural stem cell marker GFAP (red) are colocalized in RGLs of macaques. Scale bar, 10 μm. n = 4 brains. Dashed lines indicate the outline of the cell. d) Immunolabeling for ASCL1, SOX2 and GFAP in the subgranular zone (SGZ) of the dentate gyrus (DG) of macaque. White arrowheads indicate ASCL1 + SOX2 + GFAP- IPC; yellow arrowheads indicate ASCL1 + SOX2 + GFAP + RGL. Scale bars, 25 μm (low magnification), 10 μm (high magnification). n = 4 brains.
Extended Data Fig. 6 Macaque adult hippocampal transcriptomic cell types are aligned with well-established mouse datasets.
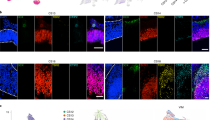
a) UMAP visualization of macaque hippocampal transcriptomic cell types aligned with the Hochgerner_2 dataset. UMAP visualization of macaque cells (n = 15,878) and mouse cells (n = 3,132), stained with the species. b) UMAP visualization of macaque cell annotations using the same UMAP coordinates in panel a, stained according to the color scheme as in Fig. 1d. Astro, Astro_1–4; Astro_im, Astro_im1, Astro_im2; GC, GC_1–3; Microglia, Microglia_1 and 2; OPC, OPC_1 and 2; Pyr, Pyr_1 and 2; RGL, RGL_1, RGL_2. c) UMAP visualization of mouse cell annotations using the same UMAP coordinates in panel a, stained according to the color scheme as in Fig. 1d. d) Cell-type homologous heatmap between macaque (in rows) and mouse (in columns) for the Hochgerner_2 dataset. Gray shades correspond to the minimum proportion of co-clustering between mouse and macaque cells. Rows show macaque populations and columns show mouse populations. Colorbar indicates the alignment score. e) Macaque hippocampal transcriptomic cell types aligned with the Joglekar dataset, visualization of macaque cells (blue, n = 15,878) and mouse cells (orange, n = 5,305) f) UMAP visualization of macaque cells, stained according to the color scheme in Fig. 1d using the same coordinates as in panel d. g) UMAP visualization of mouse cells, stained according to the color scheme in Fig. 1d using the same coordinates as in panel d. h) Cell-type homologous heatmap between macaque (in rows) and mouse (in columns) for the Joglekar dataset. Gray shades correspond to the minimum proportion of co-clustering between mouse and macaque cells. Rows show macaque populations and columns show mouse populations. Colorbar indicates the alignment score. i) Macaque hippocampal transcriptomic cell types aligned with the Shin dataset, visualization of macaque cells (blue, n = 800) and mouse cells (orange, n = 132). j) UMAP visualization of macaque cells, stained according to the color scheme in Fig. 1d using the same coordinates as in panel i. k) UMAP visualization of mouse cells, stained according to the color scheme in Fig. 1d using the same coordinates as in panel i. Gray dots are macaque cells to facilitate the visualization of the coordinates.
Extended Data Fig. 7 Macaque adult hippocampal transcriptomic cell types are aligned with well-established human datasets.
a) UMAP visualization of macaque hippocampal transcriptomic cell types aligned with the Li dataset. UMAP visualization of macaque cells (n = 2,400) and human embryonic/fetal cells (n = 323), stained with the species. b) UMAP visualization of macaque cell annotations using the same UMAP coordinates in panel a, stained according to the color scheme as in Fig. 1d. Astro, Astro_1–4; GC, GC_1–3; MG, Microglia_1 and 2; OPC, OPC_1 and 2; Pyr, Pyr_1 and 2; RGL, RGL_1 and 2. c) UMAP visualization of human cell annotations using the same UMAP coordinates in panel a, stained according to the color scheme as in Fig. 1d. d) Macaque hippocampal transcriptomic cell types aligned with the Zhong dataset, visualization of macaque cells (blue, n = 15,878) and human cells (orange, n = 6,383) e) UMAP visualization of macaque cells, stained according to the color scheme in Fig. 1d using the same coordinates as in panel d. f) UMAP visualization of human cells, stained according to the color scheme in Fig. 1d using the same coordinates as in panel d. g) Cell-type homologous heatmap between macaque (in rows) and human (in columns) for the Zhong et al. dataset. The color of each cell represents the alignment score between human and macaque cells. The larger value (darker in the heatmap) indicates a better alignment. Rows show macaque populations and columns show human populations. Colorbar indicates the alignment score.
Extended Data Fig. 8 The lineage of the neurogenic-related populations.
a) The PCA visualization of the neurogenic-related population stained by cell populations, following the same color scheme as in Fig. 1. b) The PCA visualization of the neurogenic populations stained by canonical neurogenic marker genes. Colorbar indicates log-normalized gene-expression. c) the PCA visualization with the RNA velocities visualized as arrows. d) The heatmap shows the expression of the pseudotime related genes as Viridis (light yellow as a high expression), while the colorbar on top shows the cells in the neurogenic stages using the same color scheme in a. Selective genes are labeled to the left of the heatmap. Colorbar indicates gene expression level. e) The neurogenic lineage inferred by Slingshot. f) Pseudotime analysis derived from Slingshot suggests trajectories from RGL, via IPC to NB. Colorbar indicates the derived pseudotime.
Extended Data Fig. 9 Comparison of the gene enrichment between macaques and mice.
a) Neurogenic markers that are conserved between mice and macaques. b) Species-specific marker genes that are only enriched in macaques or in mice. c) Comparison of the gene enrichment between macaques and mice for the RGLs, IPCs, neuroblasts, and GCs, together with astrocytes and mature oligodendrocytes. Solid line, equal expression; dashed lines: 2-fold enrichment. Blue dots, high-diversity genes that are >2-fold enrichment in mice. Top five genes labeled by text. Gray dots, mean expression of each neurogenic-related gene (n = 2,081 genes) of all cells of each population. Red dots, high-diversity genes that are >2-fold enrichment in macaques with the top five genes labeled by text. The “r” value, the Spearman’s correlation coefficient for each comparison. All comparisons are significant with p < 0.001. d) Neurogenic-related cell composition for macaques (n = 39,555) and mice (n = 3,711) of the Hochgerner datasets.
Extended Data Fig. 10 Process of neurosphere formation and gene expression of cultured hippocampal neurospheres.
a-d) In vitro recording process of a cultured neurosphere clone from the adult macaque hippocampi. n = 4 clones. Scale bars, 40 μm. D, day. e) UMAP visualization of the cultured sphere cells (n = 6,988 cells), stained by the expression of selective marker genes. VIM, RGL and astrocyte marker; NNAT, novel neuroblast marker; SOX4, neuroblast marker; TUJ1, neuronal marker. Colorbar indicates log-normalized gene expression level. f-h) The cultured neurosphere cells from macaque hippocampus were immunoreactive for RGL and astrocyte marker VIMENTIN in f, neuroblast marker NNAT and SOX4 in g and h. n = 4 clones. Scale bars, 40 μm. i) Immunostaining analysis of EdU long-term labeling of the cultured neurospheres from the adult macaque hippocampus showed a small number of cells in the neurosphere were labeled by fluorescence for both TUJ1 and EdU in i (indicated by arrows). Scale bars, 40 μm. n = 4 clones for each staining. j) UMAP visualization of transcriptomic cell types projected from the Hochgerner_1 dataset. Red dots represent the distribution of the predicted cell type. IPC, intermediate progenitor cell; NB, neuroblast; GC, granule cell; Astro, astrocyte.
Supplementary information
Supplementary Information
Supplementary Discussion, Supplementary Figs. 1 and 2 and Supplementary References
Supplementary Tables 1
Supplementary Tables 1–11
Source data
Source Data Fig. 2
Statistical Source Data
Source Data Extended Data Fig. 1
Statistical Source Data
Source Data Extended Data Fig. 3
Statistical Source Data
Source Data Extended Data Fig. 4
Statistical Source Data
Source Data Extended Data Fig. 6
Statistical Source Data
Source Data Extended Data Fig. 7
Statistical Source Data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hao, ZZ., Wei, JR., Xiao, D. et al. Single-cell transcriptomics of adult macaque hippocampus reveals neural precursor cell populations. Nat Neurosci 25, 805–817 (2022). https://doi.org/10.1038/s41593-022-01073-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-022-01073-x
This article is cited by
-
Reconstruction of macroglia and adult neurogenesis evolution through cross-species single-cell transcriptomic analyses
Nature Communications (2024)
-
Deciphering cell types by integrating scATAC-seq data with genome sequences
Nature Computational Science (2024)
-
Drug targeting in psychiatric disorders — how to overcome the loss in translation?
Nature Reviews Drug Discovery (2024)
-
Multimodal Nature of the Single-cell Primate Brain Atlas: Morphology, Transcriptome, Electrophysiology, and Connectivity
Neuroscience Bulletin (2024)
-
Patch-seq: Advances and Biological Applications
Cellular and Molecular Neurobiology (2024)