Abstract
Zika virus (ZIKV) is a flavivirus linked to multiple birth defects including microcephaly, known as congenital ZIKV syndrome. The identification of host factors involved in ZIKV replication may guide efficacious therapeutic interventions. In genome-wide transcriptional studies, we found that ZIKV infection triggers aryl hydrocarbon receptor (AHR) activation. Specifically, ZIKV infection induces kynurenine (Kyn) production, which activates AHR, limiting the production of type I interferons (IFN-I) involved in antiviral immunity. Moreover, ZIKV-triggered AHR activation suppresses intrinsic immunity driven by the promyelocytic leukemia (PML) protein, which limits ZIKV replication. AHR inhibition suppressed the replication of multiple ZIKV strains in vitro and also suppressed replication of the related flavivirus dengue. Finally, AHR inhibition with a nanoparticle-delivered AHR antagonist or an inhibitor developed for human use limited ZIKV replication and ameliorated newborn microcephaly in a murine model. In summary, we identified AHR as a host factor for ZIKV replication and PML protein as a driver of anti-ZIKV intrinsic immunity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The authors declare that data supporting the findings of this study are available as Supplementary Tables 1–4. Sequencing data have been deposited into the Gene Expression Omnibus (GEO) under the SuperSeries accession nos. GSE147093 and GSE147094. Data from ZIKV-infected patients were accessed at accession no. GSE139181. Data from ZIKV-infected brain organoids were accessed at accession no. GSE129180. Source data are provided with this paper.
Change history
10 August 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41593-020-0700-0
References
Baud, D., Gubler, D. J., Schaub, B., Lanteri, M. C. & Musso, D. An update on Zika virus infection. Lancet 390, 2099–2109 (2017).
Musso, D. & Gubler, D. J. Zika virus. Clin. Microbiol. Rev. 29, 487–524 (2016).
Faria, N. R. et al. Establishment and cryptic transmission of Zika virus in Brazil and the Americas. Nature 546, 406–410 (2017).
França, G. V. A. et al. Congenital Zika virus syndrome in Brazil: a case series of the first 1501 livebirths with complete investigation. Lancet 388, 891–897 (2016).
Cao-Lormeau, V. M. et al. Guillain–Barré syndrome outbreak associated with Zika virus infection in French Polynesia: a case–control study. Lancet 387, 1531–1539 (2016).
Boldescu, V., Behnam, M. A. M., Vasilakis, N. & Klein, C. D. Broad-spectrum agents for flaviviral infections: dengue, Zika and beyond. Nat. Rev. Drug Discov. 16, 565–586 (2017).
Yan, N. & Chen, Z. J. Intrinsic antiviral immunity. Nat. Immunol. 13, 214–222 (2012).
Perelygin, A. A. et al. Positional cloning of the murine flavivirus resistance gene. Proc. Natl Acad. Sci. USA 99, 9322–9327 (2002).
Giovannoni, F. et al. Dengue non-structural protein 5 polymerase complexes with promyelocytic leukemia protein (PML) isoforms III and IV to disrupt PML-nuclear bodies in infected cells. Front. Cell. Infect. Microbiol. 9, 284 (2019).
Macnamara, F. N. Zika virus: a report on three cases of human infection during an epidemic of jaundice in Nigeria. Trans. R. Soc. Trop. Med. Hyg. 48, 139–145 (1954).
Liang, Q. et al. Zika virus NS4A and NS4B proteins deregulate Akt-mTOR signaling in human fetal neural stem cells to inhibit neurogenesis and induce autophagy. Cell Stem Cell 19, 663–671 (2016).
Tang, H. et al. Zika virus infects human cortical neural progenitors and attenuates their growth. Cell Stem Cell 18, 587–590 (2016).
Liu, L. et al. Protection of ZIKV infection-induced neuropathy by abrogation of acute antiviral response in human neural progenitors. Cell Death Differ. 26, 2607–2621 (2019).
Lum, F. M. et al. Immunological observations and transcriptomic analysis of trimester-specific full-term placentas from three Zika virus-infected women. Clin. Transl. Immunol. 8, e01082 (2019).
Gutierrez-Vazquez, C. & Quintana, F. J. Regulation of the immune response by the aryl hydrocarbon receptor. Immunity 48, 19–33 (2018).
Opitz, C. A. et al. An endogenous tumour-promoting ligand of the human aryl hydrocarbon receptor. Nature 478, 197–203 (2011).
Cugola, F. R. et al. The Brazilian Zika virus strain causes birth defects in experimental models. Nature 534, 267–271 (2016).
Rothhammer, V. & Quintana, F. J. The aryl hydrocarbon receptor: an environmental sensor integrating immune responses in health and disease. Nat. Rev. Immunol. 19, 184–197 (2019).
Rothhammer, V. et al. Microglial control of astrocytes in response to microbial metabolites. Nature 557, 724–728 (2018).
Rothhammer, V. et al. Type I interferons and microbial metabolites of tryptophan modulate astrocyte activity and central nervous system inflammation via the aryl hydrocarbon receptor. Nat. Med 22, 586–597 (2016).
Yamaguchi, M. & Hankinson, O. 2,3,7,8-Tetrachlorodibenzopdioxin suppresses the growth of human liver cancer HepG2 cells in vitro: involvement of cell signaling factors. Int J. Oncol. 53, 1657–1666 (2018).
Choi, E. Y., Lee, H., Dingle, R. W., Kim, K. B. & Swanson, H. I. Development of novel CH223191-based antagonists of the aryl hydrocarbon receptor. Mol. Pharm. 81, 3–11 (2012).
Richardson, R. B. et al. A CRISPR screen identifies IFI6 as an ER-resident interferon effector that blocks flavivirus replication. Nat. Microbiol. 3, 1214–1223 (2018).
Elong Ngono, A. & Shresta, S. Immune response to dengue and zika. Annu. Rev. Immunol. 36, 279–308 (2018).
Pfeffer, L. M. et al. Role of nuclear factor-κB in the antiviral action of interferon and interferon-regulated gene expression. J. Biol. Chem. 279, 31304–31311 (2004).
Luecke, S., Wincent, E., Backlund, M., Rannug, U. & Rannug, A. Cytochrome P450 1A1 gene regulation by UVB involves crosstalk between the aryl hydrocarbon receptor and nuclear factor kappaB. Chem. Biol. Interact. 184, 466–473 (2010).
Yamada, T. et al. Constitutive aryl hydrocarbon receptor signaling constrains type I interferon–mediated antiviral innate defense. Nat. Immunol. 17, 687–694 (2016).
Dixit, E. et al. Peroxisomes are signaling platforms for antiviral innate immunity. Cell 141, 668–681 (2010).
Franz, K. M., Neidermyer, W. J., Tan, Y.-J., Whelan, S. P. J. & Kagan, J. C. STING-dependent translation inhibition restricts RNA virus replication. Proc. Natl Acad. Sci. USA 115, E2058–E2067 (2018).
Hubackova, S., Krejcikova, K., Bartek, J. & Hodny, Z. Interleukin 6 signaling regulates promyelocytic leukemia protein gene expression in human normal and cancer cells. J. Biol. Chem. 287, 26702–26714 (2012).
Stanford, E. A. et al. The role of the aryl hydrocarbon receptor in the development of cells with the molecular and functional characteristics of cancer stem-like cells. BMC Biol. 14, 20 (2016).
Taguwa, S. et al. Defining Hsp70 subnetworks in dengue virus replication reveals key vulnerability in flavivirus infection. Cell 163, 1108–1123 (2015).
Brass, A. L. et al. The IFITM proteins mediate cellular resistance to influenza A H1N1 virus, west nile virus, and dengue virus. Cell 139, 1243–1254 (2009).
Zhang, R. et al. A CRISPR screen defines a signal peptide processing pathway required by flaviviruses. Nature 535, 164–168 (2016).
Marceau, C. D. et al. Genetic dissection of Flaviviridae host factors through genome-scale CRISPR screens. Nature 535, 159–163 (2016).
Zhou, Q., Lavorgna, A., Bowman, M., Hiscott, J. & Harhaj, E. W. Aryl hydrocarbon receptor interacting protein targets IRF7 to suppress antiviral signaling and the induction of type I interferon. J. Biol. Chem. 290, 14729–14739 (2015).
Nganou-Makamdop, K. et al. Type I IFN signaling blockade by a PASylated antagonist during chronic SIV infection suppresses specific inflammatory pathways but does not alter T cell activation or virus replication. PLoS Pathog. 14, e1007246 (2018).
Gagliani, N. et al. Th17 cells transdifferentiate into regulatory T cells during resolution of inflammation. Nature 523, 221–225 (2015).
Takenaka, M. C. et al. Control of tumor-associated macrophages and T cells in glioblastoma via AHR and CD39. Nat. Neurosci. 22, 729–740 (2019).
Schiering, C. et al. Feedback control of AHR signalling regulates intestinal immunity. Nature 542, 242–245 (2017).
Yockey, L. J. et al. Type I interferons instigate fetal demise after Zika virus infection. Sci. Immunol. 3, eaao1680 (2018).
Hernandez-Ochoa, I., Karman, B. N. & Flaws, J. A. The role of the aryl hydrocarbon receptor in the female reproductive system. Biochem. Pharmacol. 77, 547–559 (2009).
Lawrence, B. P. & Vorderstrasse, B. A. New insights into the aryl hydrocarbon receptor as a modulator of host responses to infection. Semin. Immunopathol. 35, 615–626 (2013).
Wheeler, J. L. H., Martin, K. C. & Lawrence, B. P. Novel cellular targets of AHR underlie alterations in neutrophilic inflammation and inducible nitric oxide synthase expression during influenza virus infection. J. Immunol. 190, 659–668 (2012).
Winans, B. et al. Linking the aryl hydrocarbon receptor with altered DNA methylation patterns and developmentally induced aberrant antiviral CD8+ T cell responses. J. Immunol. 194, 4446–4457 (2015).
Safe, S., Cheng, Y. & Jin, U. H. The aryl hydrocarbon receptor (AhR) as a drug target for cancer chemotherapy. Curr. Opin. Toxicol. 2, 24–29 (2017).
Geoffroy, M.-C. & Chelbi-Alix, M. K. Role of promyelocytic leukemia protein in host antiviral defense. J. Interferon Cytokine Res. 31, 145–158 (2011).
Franchini, A. M. & Lawrence, B. P. Environmental exposures are hidden modifiers of anti-viral immunity. Curr. Opin. Toxicol. 10, 54–59 (2018).
Thackray, L. B. et al. Oral antibiotic treatment of mice exacerbates the disease severity of multiple flavivirus infections. Cell Rep. 22, 3440–3453.e3446 (2018).
Manfredo Vieira, S. et al. Translocation of a gut pathobiont drives autoimmunity in mice and humans. Science 359, 1156–1161 (2018).
Judd, N. P. et al. ERK1/2 regulation of CD44 modulates oral cancer aggressiveness. Cancer Res. 72, 365–374 (2012).
Bosch, I. et al. Rapid antigen tests for dengue virus serotypes and Zika virus in patient serum. Sci. Transl. Med. 9, ii: eaan1589 (2017).
Chan, J. F. et al. Improved detection of Zika virus RNA in human and animal specimens by a novel, highly sensitive and specific real-time RT-PCR assay targeting the 5′-untranslated region of Zika virus. Trop. Med. Int. Health 22, 594–603 (2017).
Lanciotti, R. S. et al. Genetic and serologic properties of Zika virus associated with an epidemic, Yap State, Micronesia, 2007. Emerg. Infect. Dis. 14, 1232–1239 (2008).
Soumillon, M., Cacchiarelli, D., Semrau, S., van Oudenaarden, A. & Mikkelsen, T. S. Characterization of Directed Differentiation by High-throughput Single-cell RNA-Seq (Cold Spring Harbor Laboratory, 2014).
Acknowledgements
This work was supported by grant nos. NS102807, NS087867, ES02530, AI126880, AI093903 and AI100190 from the National Institutes of Health, RSG-14-198-01-LIB from the American Cancer Society, and RG4111A1 and JF2161-A-5 from the National Multiple Sclerosis Society to F.J.Q., who also received support from the International Progressive MS Alliance. C.C.G. was supported by Universidad de Buenos Aires (grant no. 20020160100091BA), Consejo Nacional de Investigaciones Científicas y Tecnológicas (CONICET, grant no. PIP11220170100171CO) and Agencia de Promoción Científica y Tecnológica (ANPCyT) (grant no. BID-PICT 3080). C.C.G. is member of the Research Career CONICET. F.G. was supported by a Du Pre grant from the International MS Foundation and a postdoctoral fellowship from CONICET. J.P.S.P. was supported by FAPESP (grant nos. 2017/26170-0 and 2017/22504-1). C.M.P. and N.G.Z. received an FAPESP fellowship (grant nos. 2017/11828-0 and 2016/07371-2). We thank all members of the Garcia and Quintana laboratories for helpful advice and discussions.
Author information
Authors and Affiliations
Contributions
F.G., I.B., C.M.P., M.F.T., L.R., M.S.R.V., V.R., A.B., L.M.S., M.C.T., S.M., C.G.V., M.A.W., N.G.Z., N.B.S., C.D.M., Z.W. and J.P.S.P. performed in vitro and in vivo experiments. F.G., Z.L. and E.C.T. performed bioinformatics analysis. I.B, L.G., E.B.D., D.S., J.P.S.P., C.C.G. and F.J.Q. provided unique reagents and discussed and/or interpreted findings. F.G., C.C.G. and F.J.Q. wrote the manuscript. J.P.S.P, C.C.G. and F.J.Q. designed and supervised the study and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
F.J.Q. is a member of the Scientific Advisory Board of Kyn Therapeutics. D.S. is a co-founder of and holds equity in Hercules Pharmaceuticals, The Netherlands. The other authors declare no competing interests.
Additional information
Peer review information Nature Neuroscience thanks Charlotte Esser and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 AHR signaling is activated by ZIKV infection in human brain organoids and in patients.
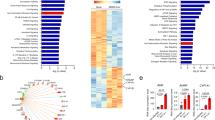
a, IPA analysis of RNA-seq data (GEO accession GSE129180) of ZIKV-infected human brain organoids. Pathways enriched in ZIKV-infected NPCs compared to control cells are shown (n = 11 per condition). Dashed red line indicates P = 0.05. P values were determined using Fisher’s exact test b, IPA analysis identified AHR as an upstream regulator. P value was determined using Fisher’s exact test c, IPA analysis of RNA-seq data (GEO accession GSE139181) of placental tissue from ZIKV-infected pregnant women compared to healthy controls (n = 3 patients per condition). Pathways enriched in ZIKV-infection are shown. Dashed red line indicates P = 0.05. P values were determined using Fisher’s exact test.
Extended Data Fig. 2 ZIKV infects NPCs in vitro.
a, NPCs were infected with ZIKV (MOI = 1) and 48 h postinfection, cells were fixed for immunofluorescence against ZIKV NS5 (green). Nuclei were counterstained with DAPI (blue). Images were acquired using an Olympus FV1000 confocal laser scanning microscope. Scale bar = 50 µm. Images are representative of 3 independent experiments. b, AHR modulation in NPCs. NPCs were treated with I3S or CH223191. 24 h after treatment, NPCs were harvested for qPCR analysis of AHR, CYP1A1, IDO1 and TDO2. Data represent the mean ± SD (n = 3 independent experiments). P values were determined by a two-sided Student’s t-test.
Extended Data Fig. 3 Effect of I3S and CH223191 on HepG2 cells viability and proliferation.
a, HepG2 cells viability after treatment for 24 h with different concentrations of I3S and CH223191 was evaluated by an MTT assay. Drug concentrations used for experiments are highlighted in color. Data represent the mean ± SD (n = 3 independent experiments). b, HepG2 cells were incubated for 24 h or 48 h with the indicated concentrations of CH223191 and the number of cells was quantified using a hemocytometer. Data represent the mean ± SD (n = 3 independent experiments). P values were determined by a one-way ANOVA followed by Dunnet’s post-hoc test.
Extended Data Fig. 4 Vero cells show functional AHR signaling.
Vero cells were treated with I3S or CH223191. 48 h after treatment, cells were harvested for qPCR analysis of AHR, CYP1A1, and CYP1B1. Data represent the mean ± SD (n = 3 independent experiments for AHR, CYP1A1; n = 4 for CYP1B1). P values were determined by a two-sided Student’s t-test.
Extended Data Fig. 5 NaF and TSA inhibit the IFN-I pathway.
Schematic representation of the IFN-I pathway. TSA inhibits the assembly of STAT1/STAT2/IRF9 complex and NaF impairs its translocation to the nucleus.
Extended Data Fig. 6 HP163 is a second generation AHR antagonist.
a, SUM149 cells were transfected with the AHR-driven pGudLuc reporter construct and treated with DMSO (vehicle) or 20 μM HP163. 24 h later, cells were assayed for pGudLuc (luciferase) activity and normalized to the CMV-green fluorescence signal. Data represent the mean ± SD (n = 3 independent experiments). P values were determined by a one-way ANOVA followed by Tukey’s post-hoc test b, Murine MOC1 oral cancer cells were treated with DMSO (vehicle) or 20 μM HP163 for 30 min and then treated with DMSO, 100 μM Kyn or 1 nM TCDD. Nuclear and cytoplasmic extracts were prepared and AHR protein quantified by Western immunoblotting. White spaces between gel pieces indicates cropping of the original image. Data are representative of three independent experiments. c, Quantification of Western immunoblotting bands from the three experiments described in (b). Data are presented as mean ± SD (n = 3 independent experiments). P values were determined by a two-sided Student’s t-test.
Extended Data Fig. 7 AHR activation boosts DENV replication in vitro.
a, A549 cells were treated with I3S (50 μM) or CH223191 (2 μM) for 24 h. Then, cells were harvested for qPCR analysis of CYP1A1. Data represent the mean ± SD (n = 3 independent experiments). P values were determined by a two-sided Student’s t-test. b, AHR activation boosts DENV replication. A549 cells were pretreated with the indicated concentrations of I3S for 24 h and infected with DENV-2 (MOI = 0.1). 48 h postinfection supernatants were harvested for plaque assay. Data represent the mean ± SD (n = 3 independent experiments). P values were determined by a one-way ANOVA followed by Dunnet’s post-hoc test.
Extended Data Fig. 8 ZIKV-driven AHR activation suppresses IFN-I dependent and IFN-I independent mechanisms that limit viral replication.
IDO1/TDO2 upregulation induced by ZIKV infection boosts the generation of Kyn, which activates AHR to inhibit IFN-I dependent and IFN-I independent mechanisms that limit ZIKV replication as follows: 1) AHR activation limits IFN-I. 2) AHR also limits NF-kB activation, suppressing PML expression. AHR signaling may also operate in a similar manner to promote DENV replication.
Supplementary information
Supplementary Tables
Supplementary Table 1: List of differentially expressed genes in ZIKV-infected HepG2 cells compared with mock-infected HepG2 cells. Supplementary Table 2: IPA analysis of ZIKV-infected HepG2 cells. Supplementary Table 3: IPA analysis of RNA-seq data from fetal mouse CNS samples. Supplementary Table 4: IPA analysis of PCR array data from fetal mouse CNS samples.
Source data
Source Data Fig. 1
Statistical source data for Fig. 1.
Source Data Fig. 2
Statistical source data for Fig. 2.
Source Data Fig. 3
Statistical source data for Fig. 3.
Source Data Fig. 3
Western blotting (Fig. 3a in manuscript).
Source Data Fig. 4
Statistical source data for Fig. 4.
Source Data Fig. 6
Statistical source data for Fig. 6.
Source Data Fig. 7
Statistical source data for Fig. 7.
Source Data Extended Data Fig. 2
Statistical source data for Extended Data Fig. 2.
Source Data Extended Data Fig. 3
Statistical source data for Extended Data Fig. 3.
Source Data Extended Data Fig. 4
Statistical source data for Extended Data Fig. 4.
Source Data Extended Data Fig. 6
Statistical source data for Extended Data Fig. 6.
Source Data Extended Data Fig. 6
Western blotting (Extended Data Fig. 6c).
Source Data Extended Data Fig. 7
Statistical source data for Extended Data Fig. 7.
Rights and permissions
About this article
Cite this article
Giovannoni, F., Bosch, I., Polonio, C.M. et al. AHR is a Zika virus host factor and a candidate target for antiviral therapy. Nat Neurosci 23, 939–951 (2020). https://doi.org/10.1038/s41593-020-0664-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-020-0664-0
This article is cited by
-
Bioluminescence imaging of Cyp1a1-luciferase reporter mice demonstrates prolonged activation of the aryl hydrocarbon receptor in the lung
Communications Biology (2024)
-
When AHR signaling pathways meet viral infections
Cell Communication and Signaling (2023)
-
Monoterpenoid aryl hydrocarbon receptor allosteric antagonists protect against ultraviolet skin damage in female mice
Nature Communications (2023)
-
Immune checkpoint modulators in cancer immunotherapy: recent advances and emerging concepts
Journal of Hematology & Oncology (2022)
-
Gain-of-function genetic screening identifies the antiviral function of TMEM120A via STING activation
Nature Communications (2022)