Abstract
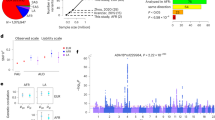
Problematic alcohol use (PAU) is a leading cause of death and disability worldwide. Although genome-wide association studies have identified PAU risk genes, the genetic architecture of this trait is not fully understood. We conducted a proxy-phenotype meta-analysis of PAU, combining alcohol use disorder and problematic drinking, in 435,563 European-ancestry individuals. We identified 29 independent risk variants, 19 of them novel. PAU was genetically correlated with 138 phenotypes, including substance use and psychiatric traits. Phenome-wide polygenic risk score analysis in an independent biobank sample (BioVU, n = 67,589) confirmed the genetic correlations between PAU and substance use and psychiatric disorders. Genetic heritability of PAU was enriched in brain and in conserved and regulatory genomic regions. Mendelian randomization suggested causal effects on liability to PAU of substance use, psychiatric status, risk-taking behavior and cognitive performance. In summary, this large PAU meta-analysis identified novel risk loci and revealed genetic relationships with numerous other traits.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The full summary-level association data from the meta-analysis are available through dbGaP at accession no. phs001672.v3.p1.
Code availability
Kinship analysis was performed using KING (http://people.virginia.edu/~wc9c/KING/). PCAs were performed using EIGENSOFT (https://data.broadinstitute.org/alkesgroup/EIGENSOFT/). Imputation was performed using EAGLE2 (https://data.broadinstitute.org/alkesgroup/Eagle/), Minimac3 (https://genome.sph.umich.edu/wiki/Minimac3), Sanger imputation server (https://imputation.sanger.ac.uk/) or RICOPILI (https://data.broadinstitute.org/mpg/ricopili/), the choice depending on the sample. GWAS was performed using PLINK (https://www.cog-genomics.org/plink2). Meta-analyses were performed using METAL (https://genome.sph.umich.edu/wiki/METAL_Documentation). Polygenic risk score analyses were performed using PRSice-2 (https://www.prsice.info/) or PRS-CS (https://github.com/getian107/PRScs). GCTA (https://cnsgenomics.com/software/gcta/#Overview) was used for identification of independent loci (GCTA-COJO), multi-trait conditional analysis (GCTA-mtCOJO) and MR (GCTA-GSMR). LDSC (https://github.com/bulik/ldsc) was used for heritability estimation, genetic correlation analysis (also using LD Hub (http://ldsc.broadinstitute.org/)) and heritability enrichment analyses. FUMA (https://fuma.ctglab.nl/) was used for gene association, functional enrichment and gene set enrichment analyses. Transcriptomic analyses were performed using S-PrediXcan and S-MultiXcan (https://github.com/hakyimlab/MetaXcan). PheWAS analyses were run using the PheWAS R package (https://github.com/PheWAS/PheWAS). The Mendelian Randomization R Package (https://cran.r-project.org/web/packages/MendelianRandomization/index.html) and MR–PRESSO (https://github.com/rondolab/MR-PRESSO) were used for MR analyses. MTAG (https://github.com/omeed-maghzian/mtag) was used for multiple trait analysis.
References
GBD 2016 Alcohol Collaborators. Alcohol use and burden for 195 countries and territories, 1990–2016: a systematic analysis for the Global Burden of Disease Study 2016. Lancet 392, 1015–1035 (2018).
Walters, R. K. et al. Transancestral GWAS of alcohol dependence reveals common genetic underpinnings with psychiatric disorders. Nat. Neurosci. 21, 1656–1669 (2018).
Kranzler, H. R. et al. Genome-wide association study of alcohol consumption and use disorder in 274,424 individuals from multiple populations. Nat. Commun. 10, 1499 (2019).
Sanchez-Roige, S. et al. Genome-wide association study meta-analysis of the alcohol use disorders identification test (AUDIT) in two population-based cohorts. Am. J. Psychiatry 176, 107–118 (2019).
Gelernter, J. et al. Genome-wide association study of maximum habitual alcohol intake in >140,000 US European and African American veterans yields novel risk loci. Biol. Psychiatry 86, 365–376 (2019).
Gaziano, J. M. et al. Million Veteran Program: a mega-biobank to study genetic influences on health and disease. J. Clin. Epidemiol. 70, 214–223 (2016).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Gelernter, J. et al. Genome-wide association study of alcohol dependence:significant findings in African- and European-Americans including novel risk loci. Mol. Psychiatry 19, 41–49 (2014).
Liu, M. et al. Association studies of up to 1.2 million individuals yield new insights into the genetic etiology of tobacco and alcohol use. Nat. Genet. 51, 237–244 (2019).
Turley, P. et al. Multi-trait analysis of genome-wide association summary statistics using MTAG. Nat. Genet. 50, 229–237 (2018).
Cotto, K. C. et al. DGIdb 3.0: a redesign and expansion of the drug-gene interaction database. Nucleic Acids Res. 46, D1068–D1073 (2018).
Bulik-Sullivan, B. K. et al. LD score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Finucane, H. K. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015).
Finucane, H. K. et al. Heritability enrichment of specifically expressed genes identifies disease-relevant tissues and cell types. Nat. Genet. 50, 621–629 (2018).
Roadmap Epigenomics Consortium, et al.Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
GTEx Consortium. Genetic effects on gene expression across human tissues. Nature 550, 204–213 (2017).
Watanabe, K. et al. Functional mapping and annotation of genetic associations with FUMA. Nat. Commun. 8, 1826 (2017).
de Leeuw, C. A. et al. MAGMA: generalized gene-set analysis of GWAS data. PLoS Comput. Biol. 11, e1004219 (2015).
Marees, A.T. et al. Potential influence of socioeconomic status on genetic correlations between alcohol consumption measures and mental health. Psychol. Med. 50, 484–498 (2020).
Grant, B. F. et al. Epidemiology of DSM-5 alcohol use disorder: results from the National Epidemiologic Survey on Alcohol and Related Conditions III. JAMA Psychiatry 72, 757–766 (2015).
Andersen, A. M. et al. Polygenic scores for major depressive disorder and risk of alcohol dependence. JAMA Psychiatry 74, 1153–1160 (2017).
Zhou, H. et al. Genetic risk variants associated with comorbid alcohol dependence and major depression. JAMA Psychiatry 74, 1234–1241 (2017).
Barbeira, A. N. et al. Exploring the phenotypic consequences of tissue-specific gene expression variation inferred from GWAS summary statistics. Nat. Commun. 9, 1825 (2018).
Battle, A. et al. Characterizing the genetic basis of transcriptome diversity through RNA-sequencing of 922 individuals. Genome Res. 24, 14–24 (2014).
Barbeira, A. N. et al. Integrating predicted transcriptome from multiple tissues improves association detection. PLoS Genet. 15, e1007889 (2019).
Evangelou, E. et al. New alcohol-related genes suggest shared genetic mechanisms with neuropsychiatric disorders. Nat. Hum. Behav. 3, 950–961 (2019).
Bowden, J. et al. A framework for the investigation of pleiotropy in two-sample summary data Mendelian randomization. Stat. Med. 36, 1783–1802 (2017).
Bowden, J. et al. Consistent estimation in Mendelian randomization with some invalid instruments using a weighted median estimator. Genet. Epidemiol. 40, 304–314 (2016).
Bowden, J., Davey Smith, G. & Burgess, S. Mendelian randomization with invalid instruments: effect estimation and bias detection through Egger regression. Int. J. Epidemiol. 44, 512–525 (2015).
Verbanck, M. et al. Detection of widespread horizontal pleiotropy in causal relationships inferred from Mendelian randomization between complex traits and diseases. Nat. Genet. 50, 693–698 (2018).
Zhu, Z. et al. Causal associations between risk factors and common diseases inferred from GWAS summary data. Nat. Commun. 9, 224 (2018).
Howard, D. M. et al. Genome-wide meta-analysis of depression identifies 102 independent variants and highlights the importance of the prefrontal brain regions. Nat. Neurosci. 22, 343–352 (2019).
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Stahl, E. A. et al. Genome-wide association study identifies 30 loci associated with bipolar disorder. Nat. Genet. 51, 793–803 (2019).
Nagel, M. et al. Meta-analysis of genome-wide association studies for neuroticism in 449,484 individuals identifies novel genetic loci and pathways. Nat. Genet. 50, 920–927 (2018).
Karlsson Linner, R. et al. Genome-wide association analyses of risk tolerance and risky behaviors in over 1 million individuals identify hundreds of loci and shared genetic influences. Nat. Genet. 51, 245–257 (2019).
Jansen, P. R. et al. Genome-wide analysis of insomnia in 1,331,010 individuals identifies new risk loci and functional pathways. Nat. Genet. 51, 394–403 (2019).
Lee, J. J. et al. Gene discovery and polygenic prediction from a genome-wide association study of educational attainment in 1.1 million individuals. Nat. Genet. 50, 1112–1121 (2018).
Pasman, J. A. et al. GWAS of lifetime cannabis use reveals new risk loci, genetic overlap with psychiatric traits, and a causal influence of schizophrenia. Nat. Neurosci. 21, 1161–1170 (2018).
Okbay, A. et al. Genetic variants associated with subjective well-being, depressive symptoms, and neuroticism identified through genome-wide analyses. Nat. Genet. 48, 624–633 (2016).
Cross-Disorder Group of the Psychiatric Genomics Consortium. Identification of risk loci with shared effects on five major psychiatric disorders: a genome-wide analysis. Lancet 381, 1371–1379 (2013).
Demontis, D. et al. Discovery of the first genome-wide significant risk loci for attention deficit/hyperactivity disorder. Nat. Genet. 51, 63–75 (2019).
Pilling, L. C. et al. Human longevity is influenced by many genetic variants: evidence from 75,000 UK Biobank participants. Aging (Albany NY) 8, 547–560 (2016).
Loh, P. R. et al. Reference-based phasing using the Haplotype Reference Consortium panel. Nat. Genet. 48, 1443–1448 (2016).
Das, S. et al. Next-generation genotype imputation service and methods. Nat. Genet. 48, 1284–1287 (2016).
1000 Genomes Project Consortium. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Manichaikul, A. et al. Robust relationship inference in genome-wide association studies. Bioinformatics 26, 2867–2873 (2010).
Galinsky, K. J. et al. Fast principal-component analysis reveals convergent evolution of ADH1B in Europe and East Asia. Am. J. Hum. Genet. 98, 456–472 (2016).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Euesden, J., Lewis, C. M. & O’Reilly, P. F. PRSice: Polygenic Risk Score software. Bioinformatics 31, 1466–1468 (2015).
Yang, J. et al. Conditional and joint multiple-SNP analysis of GWAS summary statistics identifies additional variants influencing complex traits. Nat. Genet. 44, 369–375 (2012).
Zhu, Z. et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat. Genet. 48, 481–487 (2016).
InternationalHapMap Consortium, et al.Integrating common and rare genetic variation in diverse human populations. Nature 467, 52–58 (2010).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Zheng, J. et al. LD Hub: a centralized database and web interface to perform LD score regression that maximizes the potential of summary level GWAS data for SNP heritability and genetic correlation analysis. Bioinformatics 33, 272–279 (2017).
Savage, J. E. et al. Genome-wide association meta-analysis in 269,867 individuals identifies new genetic and functional links to intelligence. Nat. Genet. 50, 912–919 (2018).
Jansen, I. E. et al. Genome-wide meta-analysis identifies new loci and functional pathways influencing Alzheimer’s disease risk. Nat. Genet. 51, 404–413 (2019).
Gamazon, E. R. et al. A gene-based association method for mapping traits using reference transcriptome data. Nat. Genet. 47, 1091–1098 (2015).
Ge, T. et al. Polygenic prediction via Bayesian regression and continuous shrinkage priors. Nat. Commun. 10, 1776 (2019).
Price, A. L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–904 (2006).
McCarthy, S. et al. A reference panel of 64,976 haplotypes for genotype imputation. Nat. Genet. 48, 1279–1283 (2016).
Denny, J. C. et al. Systematic comparison of phenome-wide association study of electronic medical record data and genome-wide association study data. Nat. Biotechnol. 31, 1102–1110 (2013).
Carroll, R. J., Bastarache, L. & Denny, J. C. R PheWAS: data analysis and plotting tools for phenome-wide association studies in the R environment. Bioinformatics 30, 2375–2376 (2014).
Pedersen, C. B. et al. The iPSYCH2012 case-cohort sample: new directions for unravelling genetic and environmental architectures of severe mental disorders. Mol. Psychiatry 23, 6–14 (2018).
Lam, M. et al. RICOPILI: Rapid Imputation for COnsortias PIpeLIne. Bioinformatics 36, 930–933 (2020).
McCarthy, S. et al. A reference panel of 64,976 haplotypes for genotype imputation. Nat. Genet. 48, 1279–1283 (2016).
Yavorska, O. O. & Burgess, S. MendelianRandomization: an R package for performing Mendelian randomization analyses using summarized data. Int. J. Epidemiol. 46, 1734–1739 (2017).
Acknowledgements
This research used data from MVP, and was supported by funding from the Department of Veterans Affairs Office of Research and Development, Million Veteran Program Grant nos. I01BX003341 and I01CX001849; and the VA Cooperative Studies Program study, no. 575B. This publication does not represent the views of the Department of Veterans Affairs or the United States Government. A list of members and affiliations of MVP appears in the Supplementary Information. Supported also by NIH (NIAAA) no. P50 AA12870 (to J.G.), a NARSAD Young Investigator Grant from the Brain & Behavior Research Foundation (to H.Z.) and NIH grants nos. 5T32GM080178 (to J.M.S.) and K02DA32573 (to A.A.), and the National Institute for Healthcare Research (NIHR) Imperial Biomedical Research Centre (BRC; to S.R.A. and M.R.T.). This research also used summary data from the PGC Substance Use Disorders (SUD) working group. PGC–SUD is supported by funds from NIDA and NIMH to MH109532 and, previously, had analyst support from NIAAA to U01AA008401 (COGA). PGC–SUD gratefully acknowledges its contributing studies and the participants in those studies, without whom this effort would not be possible. This research also used individual-level/summary data from UKB, a population-based sample of participants whose contributions we gratefully acknowledge; project ID 41910. We thank the iPSYCH-Broad Consortium for access to data on the iPSYCH cohort. The iPSYCH project is funded by the Lundbeck Foundation (nos. R102-A9118 and R155-2014-1724) and the universities and university hospitals of Aarhus and Copenhagen. Genotyping of iPSYCH samples was supported by grants from the Lundbeck Foundation and the Stanley Foundation, and The Danish National Biobank resource was supported by the Novo Nordisk Foundation. Data handling and analysis on the GenomeDK HPC facility was supported by NIMH (no. 1U01MH109514-01 to A.D.B.). High-performance computer capacity for handling and statistical analysis of iPSYCH data on the GenomeDK HPC facility was provided by the Centre for Integrative Sequencing, Aarhus University, Denmark (grant to A.D.B.). The UCL and STOPAH case samples were genotyped with funding from the NIHR BRC. UCL cases and controls were collected with UK Medical Research Council project grants nos. G9623693N, G0500791, G0701007 and G1000708, and with support from the NIHR. Genotyping of the UCL control sample was supported by grants from the Stanley Foundation. The UK Household Longitudinal Study (Understanding Society) is led by the Institute for Social and Economic Research at the University of Essex and funded by the Economic and Social Research Council. The survey was conducted by NatCen, and the genome-wide scan data were analyzed and deposited by the Wellcome Trust Sanger Institute. Information on how to access the data can be found on the Understanding Society website: https://www.understandingsociety.ac.uk/. A.M. is supported by the University College London Hospitals NHS Foundation Trust NIHR BRC.
Author information
Authors and Affiliations
Contributions
H.Z., J.G., H.R.K. and A.A.P. conceived the analyses. H.Z. and J.G. wrote the first draft and prepared all drafts for submission. J.G. supervised and H.Z. accomplished primary analyses. J.M.S., S.S.-R., T.-K.C., D.F.L., Z.C., B.L. and A.M. conducted additional analyses. J.G., H.R.K., A.A.P., L.K.D., H.J.E. and A.A. supervised additional analyses. J.M.S., S.S.-R., T.-K.C., A.A.P., A.M. and L.K.D. prepared individual datasets and provided summary statistics or results. R.P., R.L.K., R.V.S., J.H.T., M.Y.M., S.R.A., M.R.T., M.N., M.M., A.D.B., E.C.J., A.C.J., A.M., L.K.D. and H.R.K. provided critical support regarding phenotypes and data in individual datasets. J.G., A.C.J. and H.R.K. provided resource support. All authors reviewed the manuscript and approved it for submission.
Corresponding author
Ethics declarations
Competing interests
H.R.K. is a member of the American Society of Clinical Psychopharmacology’s Alcohol Clinical Trials Initiative, which was supported over the past 3 years by AbbVie, Alkermes, Ethypharm, Indivior, Lilly, Lundbeck, Otsuka, Pfizer, Arbor, and Amygdala Neurosciences. H.R.K. and J.G. are named as inventors on PCT patent application no. 15/878,640, entitled Genotype-guided dosing of opioid agonists, filed 24 January 2018.
Additional information
Peer review information Nature Neuroscience thanks Gerome Breen, Eske Derks, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
A list of members and affiliations in MVP, and Supplementary Figs. 1–6.
Rights and permissions
About this article
Cite this article
Zhou, H., Sealock, J.M., Sanchez-Roige, S. et al. Genome-wide meta-analysis of problematic alcohol use in 435,563 individuals yields insights into biology and relationships with other traits. Nat Neurosci 23, 809–818 (2020). https://doi.org/10.1038/s41593-020-0643-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-020-0643-5
This article is cited by
-
Genetic factors associated with suicidal behaviors and alcohol use disorders in an American Indian population
Molecular Psychiatry (2024)
-
Medical and genetic correlates of long-term buprenorphine treatment in the electronic health records
Translational Psychiatry (2024)
-
Pleiotropy and genetically inferred causality linking multisite chronic pain to substance use disorders
Molecular Psychiatry (2024)
-
Genetic architecture of the structural connectome
Nature Communications (2024)
-
Unsupervised deep representation learning enables phenotype discovery for genetic association studies of brain imaging
Communications Biology (2024)