Abstract
To make appropriate decisions, animals need to accumulate sensory evidence. Simple integrator models can explain many aspects of such behavior, but how the underlying computations are mechanistically implemented in the brain remains poorly understood. Here we approach this problem by adapting the random-dot motion discrimination paradigm, classically used in primate studies, to larval zebrafish. Using their innate optomotor response as a measure of decision making, we find that larval zebrafish accumulate and remember motion evidence over many seconds and that the behavior is in close agreement with a bounded leaky integrator model. Through the use of brain-wide functional imaging, we identify three neuronal clusters in the anterior hindbrain that are well suited to execute the underlying computations. By relating the dynamics within these structures to individual behavioral choices, we propose a biophysically plausible circuit arrangement in which an evidence integrator competes against a dynamic decision threshold to activate a downstream motor command.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon request.
Code availability
All analysis and hardware control code will be made available by the corresponding author upon request.
References
Shadlen, M. N. & Kiani, R. Decision making as a window on cognition. Neuron 80, 791–806 (2013).
Ratcliff, R., Smith, P. L., Brown, S. D. & McKoon, G. Diffusion decision model: current issues and history. Trends Cogn. Sci. 20, 260–281 (2016).
Groschner, L. N., Chan Wah Hak, L., Bogacz, R., DasGupta, S. & Miesenböck, G. Dendritic integration of sensory evidence in perceptual decision-making. Cell 173, 894–905 (2018).
Borst, A. & Bahde, S. Visual information processing in the fly’s landing system. J. Comp. Physiol. A 163, 167–173 (1988).
DasGupta, S., Ferreira, C. H. & Miesenböck, G. FoxP influences the speed and accuracy of a perceptual decision in Drosophila. Science 344, 901–904 (2014).
Evans, D. A. et al. A synaptic threshold mechanism for computing escape decisions. Nature 558, 590–594 (2018).
Odoemene, O., Pisupati, S., Nguyen, H. & Churchland, A. K. Visual evidence accumulation guides decision-making in unrestrained mice. J. Neurosci. 38, 10143–10155 (2018).
Hanks, T. D. et al. Distinct relationships of parietal and prefrontal cortices to evidence accumulation. Nature 520, 220–223 (2015).
Hanks, T. D., Ditterich, J. & Shadlen, M. N. Microstimulation of macaque area LIP affects decision-making in a motion discrimination task. Nat. Neurosci. 9, 682–689 (2006).
Huk, A. C., Katz, L. N. & Yates, J. L. The role of the lateral intraparietal area in (the study of) decision making. Annu. Rev. Neurosci. 40, 349–372 (2017).
Shadlen, M. N. & Newsome, W. T. Neural basis of a perceptual decision in the parietal cortex (area LIP) of the rhesus monkey. J. Neurophysiol. 86, 1916–1936 (2001).
Roitman, J. D. & Shadlen, M. N. Response of neurons in the lateral intraparietal area during a combined visual discrimination reaction time task. J. Neurosci. 22, 9475–9489 (2002).
Newsome, W. T., Britten, K. H. & Movshon, J. A. Neuronal correlates of a perceptual decision. Nature 341, 52–54 (1989).
Newsome, W. T. & Pare, E. B. A selective impairment of motion perception following lesions of the middle temporal visual area (MT). J. Neurosci. 8, 2201–2211 (1988).
Salzman, C. D., Britten, K. H. & Newsome, W. T. Cortical microstimulation influences perceptual judgements of motion direction. Nature 346, 174–177 (1990).
Naumann, E. A. et al. From whole-brain data to functional circuit models: the zebrafish optomotor response. Cell 167, 947–960 (2016).
Portugues, R., Haesemeyer, M., Blum, M. L. & Engert, F. Whole-field visual motion drives swimming in larval zebrafish via a stochastic process. J. Exp. Biol. 218, 1433–1443 (2015).
Ahrens, M. B. et al. Brain-wide neuronal dynamics during motor adaptation in zebrafish. Nature 485, 471–477 (2012).
Kubo, F. et al. Functional architecture of an optic flow-responsive area that drives horizontal eye movements in zebrafish. Neuron 81, 1344–1359 (2014).
Wang, K., Hinz, J., Haikala, V., Reiff, D. F. & Arrenberg, A. B. Selective processing of all rotational and translational optic flow directions in the zebrafish pretectum and tectum. BMC Biol. 17, 29 (2019).
Dunn, T. W. et al. Brain-wide mapping of neural activity controlling zebrafish exploratory locomotion. Elife 5, e12741 (2016).
Chen, X. et al. Brain-wide organization of neuronal activity and convergent sensorimotor transformations in larval zebrafish. Neuron 100, 876–890 (2018).
Pnevmatikakis, E. A. et al. Simultaneous denoising, deconvolution, and demixing of calcium imaging data. Neuron 89, 285–299 (2016).
Randlett, O. et al. Whole-brain activity mapping onto a zebrafish brain atlas. Nat. Methods 12, 1039–1046 (2015).
Satou, C. et al. Transgenic tools to characterize neuronal properties of discrete populations of zebrafish neurons. Development 140, 3927–3931 (2013).
Kunst, M. et al. A cellular-resolution atlas of the larval zebrafish brain. Neuron 103, 21–38 (2019).
Huang, K.-H., Ahrens, M. B., Dunn, T. W. & Engert, F. Spinal projection neurons control turning behaviors in zebrafish. Curr. Biol. 23, 1566–1573 (2013).
Orger, M. B., Kampff, A. R., Severi, K. E., Bollmann, J. H. & Engert, F. Control of visually guided behavior by distinct populations of spinal projection neurons. Nat. Neurosci. 11, 327–333 (2008).
Fisher, D., Olasagasti, I., Tank, D. W., Aksay, E. R. F. & Goldman, M. S. A modeling framework for deriving the structural and functional architecture of a short-term memory microcircuit. Neuron 79, 987–1000 (2013).
Chen, T.-W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Ehrlich, D. E. & Schoppik, D. Control of movement initiation underlies the development of balance. Curr. Biol. 27, 334–344 (2017).
Oteiza, P., Odstrcil, I., Lauder, G., Portugues, R. & Engert, F. A novel mechanism for mechanosensory-based rheotaxis in larval zebrafish. Nature 547, 445–448 (2017).
Orger, M. B., Smear, M. C., Anstis, S. M. & Baier, H. Perception of Fourier and non-Fourier motion by larval zebrafish. Nat. Neurosci. 3, 1128–1133 (2000).
Wolf, S. et al. Sensorimotor computation underlying phototaxis in zebrafish. Nat. Commun. 8, 1–12 (2017).
Haesemeyer, M., Robson, D. N., Li, J. M., Schier, A. F. & Engert, F. A brain-wide circuit model of heat-evoked swimming behavior in larval zebrafish. Neuron 98, 817–831 (2018).
Northmore, D. P. M. Holding visual attention for 400 million years: A model of tectum and torus longitudinalis in teleost fishes. Vision Res. 131, 44–56 (2017).
Akrami, A., Kopec, C. D., Diamond, M. E. & Brody, C. D. Posterior parietal cortex represents sensory history and mediates its effects on behaviour. Nature 554, 368–372 (2018).
Wang, X.-J. Probabilistic decision making by slow reverberation in cortical circuits. Neuron 36, 955–968 (2002).
Zhang, J. The effects of evidence bounds on decision-making: theoretical and empirical developments. Front. Psychol. 3, 1–19 (2012).
Thura, D. & Cisek, P. Modulation of premotor and primary motor cortical activity during volitional adjustments of speed-accuracy trade-offs. J. Neurosci. 36, 938–956 (2016).
Hanks, T. D., Mazurek, M. E., Kiani, R., Hopp, E. & Shadlen, M. N. Elapsed decision time affects the weighting of prior probability in a perceptual decision task. J. Neurosci. 31, 6339–6352 (2011).
Drugowitsch, J., Moreno-Bote, R., Churchland, A. K., Shadlen, M. N. & Pouget, A. The cost of accumulating evidence in perceptual decision making. J. Neurosci. 32, 3612–3628 (2012).
Goldman, M. S. Memory without feedback in a neural network. Neuron 61, 621–634 (2009).
Lim, S. & Goldman, M. S. Balanced cortical microcircuitry for maintaining information in working memory. Nat. Neurosci. 16, 1306–1314 (2013).
Grama, A. & Engert, F. Direction selectivity in the larval zebrafish tectum is mediated by asymmetric inhibition. Front. Neural Circuits 6, 59 (2012).
Förster, D., Dal Maschio, M., Laurell, E. & Baier, H. An optogenetic toolbox for unbiased discovery of functionally connected cells in neural circuits. Nat. Commun. 8, 116 (2017).
Hildebrand, D. G. C. et al. Whole-brain serial-section electron microscopy in larval zebrafish. Nature 545, 345–349 (2017).
Dal Maschio, M., Donovan, J. C., Helmbrecht, T. O. & Baier, H. Linking neurons to network function and behavior by two-photon holographic optogenetics and volumetric imaging. Neuron 94, 774–789 (2017).
Kim, D. H. et al. Pan-neuronal calcium imaging with cellular resolution in freely swimming zebrafish. Nat. Methods 14, 1107–1114 (2017).
Haug, M. F., Biehlmaier, O., Mueller, K. P. & Neuhauss, S. C. Visual acuity in larval zebrafish: behavior and histology. Front. Zool. 7, 1–8 (2010).
Rohlfing, T. & Maurer, C. R. Jr. Nonrigid image registration in shared-memory multiprocessor environments with application to brains, breasts, and bees. IEEE Trans. Inf. Technol. Biomed. 7, 16–25 (2003).
Pnevmatikakis, E. A. & Giovannucci, A. NoRMCorre: An online algorithm for piecewise rigid motion correction of calcium imaging data. J. Neurosci. Methods 291, 83–94 (2017).
Portugues, R., Feierstein, C. E., Engert, F. & Orger, M. B. Whole-brain activity maps reveal stereotyped, distributed networks for visuomotor behavior. Neuron 81, 1328–1343 (2014).
Avants, B. B., Epstein, C. L., Grossman, M. & Gee, J. C. Symmetric diffeomorphic image registration with cross-correlation: evaluating automated labeling of elderly and neurodegenerative brain. Med. Image Anal. 12, 26–41 (2008).
Marquart, G. D. et al. High-precision registration between zebrafish brain atlases using symmetric diffeomorphic normalization. Gigascience 6, 1–15 (2017).
Ginzburg, I. & Sompolinsky, H. Theory of correlations in stochastic neural networks. Phys. Rev. E. Stat. Phys. Plasmas Fluids Relat. Interdiscip. Topics 50, 3171–3191 (1994).
Choi, S., Kim, T. & Yu, W. Performance evaluation of RANSAC family. Proc. British Machine Vision Conference 2009 1–12 (British Machine Vision Association, 2009)..
Acknowledgements
We are grateful to H. Sompolinsky for discussions, and A. Borst, J. Drugowitsch, M. Haesemeyer, K. Herrera, R. Harpaz, and K. Vogt for discussions and helpful comments on the manuscript. We thank S. Foianini for fish line maintenance and assistance with the behavioral experiments and K. Herrera for help with fish crossings. A.B. was supported by the Human Frontier Science Program Long-Term Fellowship LT000626/2016. F.E. received funding from the National Institutes of Health (U19 NS104653, R43 OD024879, and R24 NS086601), and the Simons Foundation (SCGB 325207 and SCGB 542973).
Author information
Authors and Affiliations
Contributions
A.B. and F.E. conceived the project. A.B. designed and performed the experiments, built and programmed the behavioral setups and the two-photon microscope, analyzed the data, and carried out the modeling. A.B. wrote the manuscript with help from F.E.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Behavior in freely swimming larval zebrafish.
a, Turn angle larvae accumulate over time (positive, right; negative, left). b, Probability distribution of inter-bout intervals during presentation of different coherence levels. c, Time-binned inter-bout intervals as a function of time. d, Probability distributions of turn angles per bout relative to motion direction. Bouts are defined as correct when they follow motion direction (positive) and incorrect otherwise (negative). e, Time-binned precision (absolute turn angle) of correct and incorrect bouts over time. Gray shaded areas in a,c,e indicate motion presentation. Before and after, we always show 0% coherence. Bin sizes in c,e are 2 s. All error bars are mean ± s.e.m. over fish. n = 60 fish in a–e, same fish as in Fig. 1b–d.
Extended Data Fig. 2 Model alternatives for freely swimming larval zebrafish.
a–d, Schematics and simulation results of alternative models. For simplicity, none of these models had visual feedback. Quantification as in Fig. 1b–f,h–l. n = 16 model runs for each model. e, Pearson’s correlation coefficient between each model feature and the respective experimental data. We use the average of these values to rank the models. All error bars in a–d are mean ± s.e.m. over model runs.
Extended Data Fig. 3 Behavior and modeling in head-fixed larval zebrafish.
a, 100 randomly selected experimental example trials with periods of coherent motion (shades of gray) and 0% coherence (lightest gray), sorted by response delay. Note that after each correct (green dots) or incorrect (blue dots) bout, coherence levels are immediately set to 0%. b, Probability density distributions of response delays for correct (solid lines) and incorrect (dashed lines) bouts, for experiment (black lines) and models (red, orange, and blue lines). Models have the same structure as in Extended Data Fig. 2a,c,d but without spontaneous bouts below the bound and different parameters. c, Accuracy as a function of response delay (short, 0–2 s; medium, 2–4 s; long, 4–6 s) for experiment (*P < 0.05 for both comparisons at 50% coherence) and models. Behavioral data in a–c come from the experiment with constant motion coherence (Fig. 2b). d–g, Behavior quantification for experiment and models, as in Fig. 2c,e,g–i,k–m. h, Pearson’s correlation coefficient between each model feature and the respective experimental data. We use the average of these values to rank the models. n = 13 fish in a–d, n = 10 fish in e, n = 8 fish (left panels) and n = 6 fish (right panel) in f, n = 13 fish in g, same fish as in Fig. 2c,e,g–i,k–m. n = 8 model runs for each model in b–h. All error bars are mean ± s.e.m. over fish. P values in (c) are based on one-sided t-tests comparing response differences to zero. Asterisks in c indicate significance (*P < 0.05).
Extended Data Fig. 4 Detailed quantification of responsive brain areas identified during brain-wide calcium imaging.
a, All brain areas with >1% responsive cells, sorted by fitted onset time constant during 50% coherent motion. Text label colors relate to colors in (b) and Fig. 3c,d. Bar colors represent fraction of responsive cells within a brain region. Black arrows indicate anterior hindbrain regions with slow dynamics and a large fraction of responsive cells. b, Brain areas with >15% responsive cells sorted by temporal dynamics (top, fast; bottom, slow) characterized from left to right. Column 1: Peak-normalized calcium dynamics, relative to baseline (C0) averaged over all cells responding to coherent motion in preferred- (PD) or null-direction (ND), respectively. Column 2: Average (last 5 s of coherent motion) calcium response amplitude (comparisons between 50% and 100%, from top to bottom: P = 0.05, P = 0.06, P = 0.16, P < 0.05, P < 0.01, P < 0.05, P < 0.05). Column 3: Variance (over time), calculated in individual cells and trials, then averaged, during 0% coherence Column 4: Same as column 3 but time-binned for regions preferred- and null-direction. As variances for preferred- and null-direction motion quickly converge after motion stimulation, the last time bin reflects a motion-memory-independent variance at 0% coherence. c, Preferred- and null-direction dynamics of all identified anterior hindbrain cells functionally clustered by regressor analysis (Fig. 3e). Preferred motion direction refers to motion to the left or right for cells in the left or right hemisphere, respectively, null-direction motion the other way around. d, Spatial arrangement of trial-to-trial reliability for all cells without functional clustering, as in Fig. 3g but for all three coherence levels. n = 6 fish for 50% and n = 6 fish for 100% motion coherence stimulation in a,b. Open circles in a,b indicate individual fish. Note that in some fish not all brain areas were imaged and, hence, fish number per brain region is variable. n = 6 fish in c,d. All error bars in a,b indicate mean ± s.e.m. over fish. Shaded gray areas in b and dashed vertical lines in c indicate motion stimulation. Before and after 0% coherence is shown. All P values are based on two-sided t-tests.
Extended Data Fig. 5 Neurotransmitter identity and neuronal morphology in the anterior hindbrain.
a, Overlay of neurotransmitter type-specific masks from the Z-brain atlas (refs. 24,25) (red outline, glutamate; blue outline, GABA) with the functionally characterized cell types (same cells as in Fig. 3f) in the same coordinate system. Note that almost all identified dynamic threshold neurons lie within the Gad1b cluster 2 and are therefore likely to be inhibitory. Please also note that the expression pattern of the vglut2a-driver line (ref. 25) suggests that the anatomical mask of the VGlut2 cluster 1 is likely to extend even further into the anterior part of the hindbrain. b, Simultaneous imaging of DsRed (pink), expressed only in excitatory vglut2a+ neurons (left panel) or of DsRed expressed only in inhibitory gad1b+ neurons (right panel), and cytosolic or nuclear-localized GCaMP6s (green, pan-neuronal expression). Colored ellipses are manually added to highlight regions of interest. c, Single-cell morphologies with somata in the identified regions in the anterior hindbrain. Cells were mapped from the Max-Planck Zebrafish Brain Atlas (ref. 26) into the Z-brain coordinate system and overlaid with the available masks (gray) on a GCaMP5G reference larval zebrafish (green). n = 6 fish in a, same data as in Fig. 3f. n = 1 fish for each plot in b.
Extended Data Fig. 6 Neural correlates of behavioral choices.
a, Measured cluster dynamics aligned to swim bouts (same data as in Fig. 4b, but showing only ipsilateral dynamics for medium response delays). The thick black line illustrates the difference between the dynamic threshold cluster and the evidence integrator cluster, which crosses zero (baseline) around the same time the motor command cluster reaches its maximum. This event occurs slightly after the bout, probably owing to delays introduced by the relatively slow dynamics of the GCaMP6s indicator (ref. 30). The transparent thin lines are single-trial responses. b,c, Same analysis as in Fig. 4b,c but for bout-aligned network model simulations (*P < 0.001 for the integrator and dynamic threshold unit comparisons; P = 0.58 and P = 0.06 for the motor command cluster comparisons). d, Bout-aligned preferred- and null-direction dynamics of all identified cells from all n = 5 fish, functionally clustered by the behavior-based classification method. Preferred motion direction refers to motion to the left or right for cells in the left or right hemisphere, respectively; null-direction motion the other way around. e, Probability density distributions of response delays for correct (solid lines) and incorrect (dashed lines) bouts, for experiment (black) and network model (brown). f, Illustration of two methods for trial-by-trial prediction of individual behavioral choices based on the experimentally obtained cluster dynamics. Bouts are predicted when the smoothed and extrapolated integrator cluster activity (red dashed lines are exponential fits of the experimental data, solid red lines) crosses the threshold (cyan lines). Three fixed thresholds (left panel) and the dynamic threshold (right panel) are tested. The third method, which uses a sudden rise in the motor command slope as a predictor, is not illustrated. g, Quantification of the fraction of trials in which a threshold crossing event is detected and quantification of predictive quality (coefficient of determination, R2) for the different threshold models (*P < 0.05 for each model compared to the dynamic threshold model). Gray lines are individual fish, black lines are fish averages. h, Trial-by-trial predictions of bout-timing for individual fish using the dynamic threshold, and robust linear regression analysis results (RANSAC, see Methods). Gray shaded areas indicate confidence intervals of the regression fits. n = 5 fish in a,d,e,g,h, same fish as in Fig. 4. n = 8 model runs in b,c,e. All error bars are mean ± s.e.m. over simulated trials in b, model runs in c, or fish in g. P values in c,g are based on one-sided t-tests comparing differences to zero. Asterisks (*) in c,g indicate significance (*P < *0.05, or *P < 0.001).
Extended Data Fig. 7 Speculative network model implementation of urgency-related signals in freely swimming larval zebrafish.
a, Network model as in Fig. 3k but with inhibitory bout clock attached to the dynamic threshold clusters. We speculate that the bout clock or the system for keeping balance act as urgency-related signals here. These signals lead to rapidly collapsing bounds, allowing for spontaneous swimming. Also, in this model, each simulated bout induced opposing visual feedback, as in Fig. 1g–l. b, Copy of behavior data from Fig. 1b–f. c, Network model simulation results, quantified as in Fig. 1b–f, h–l. n = 8 model runs in c. All error bars are mean ± s.e.m. over fish in b or model runs in c. Asterisks in b indicate significance (*P < 0.05, *P < 0.01, or *P < 0.001). See Fig. 1b–f for more details on P values and statistics.
Supplementary information
Supplementary Video 1
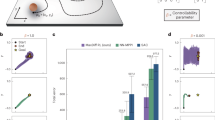
Freely swimming larval zebrafish with example dot motion stimulus. We present 5 s of 0% coherence, followed by 10 s of 25% coherence and 5 s of 0% coherence. Dot motion is always perpendicular (rightward or leftward) locked to the body orientation of the fish. Dots have a short lifetime and stochastically disappear and reappear at a random location within the arena. Left panel shows the perspective of the experimenter (world-centric view). Right panel shows the visual stimulus from the perspective of the fish (fish-centric view). Note that sizes are not to scale. The body length of a larva zebrafish at this age (6 days post fertilization) is about 4 mm. The diameter of the visual arena is 12 cm.
Supplementary Video 2
Simulation of the leaky integrator model with visual feedback for freely swimming larval zebrafish. We present different levels of coherence (moving rightward or leftward), indicated by the orange and blue shaded areas. Bouts are initiated by a stochastic bout clock operating at a low probability below the threshold and a higher probability above it. Orange dots indicate bouts to the right, blue dots indicate bouts to the left. Note that in this model, each bout slightly resets the integrator, mimicking self-created optic flow and visual noise.
Supplementary Video 3
Simulation of the stochastic model for freely swimming larval zebrafish. Same as in Supplementary Video 2 but for the stochastic model. Note that threshold crossing events are rarely visible because we slightly low-pass filter the sensory variable over time. Otherwise, the trace would not be resolvable in the movie.
Supplementary Video 4
Simulation of the stochastic model with motor memory for freely swimming larval zebrafish. Same as in Supplementary Video 2 but for the stochastic model with motor memory. Bouts are initiated according to the same rules as in the stochastic model. Additionally, a motor memory component allows larvae to sometimes simply repeat the last motor action.
Supplementary Video 5
Simulation of the leaky integrator model without visual feedback for freely swimming larval zebrafish. Same as in Supplementary Video 2 but for a simpler version of the bounded leaky integrator model in which we did not include self-created optic flow or visual noise.
Supplementary Video 6
Simulation of the non-leaky integrator model with reset and motor memory for freely swimming larval zebrafish. Same as in Supplementary Video 2 but for an integrator model without a leak component in which bouts completely reset the sensory variable to zero. Similarly, as in Supplementary Video 4, an additional motor memory component allows larvae to sometimes repeat the last motor action.
Supplementary Video 7
Simulation of the leaky integrator model for head-fixed larval zebrafish. Following the behavior of head-fixed larvae, bouts (orange and blue dots) are only triggered when the integrator variable reaches the threshold and not stochastically. We present different levels of coherence (moving rightward or leftward), indicated by the orange and blue shaded areas. Note that whenever we detect a bout, we immediately set coherence levels to zero, as in the experiments.
Supplementary Video 8
Simulation of the stochastic model for head-fixed larval zebrafish. Same as in Supplementary Video 7 but for the stochastic model. Note that threshold crossing events are rarely visible because we slightly low-pass filter the sensory variable over time. Otherwise, the trace would not be resolvable in the movie.
Supplementary Video 9
Simulation of the non-leaky integrator model with reset and motor memory for head-fixed larval zebrafish. Same as in Supplementary Video 7 but for an integrator model without a leak component in which bouts completely reset the sensory variable to zero.
Supplementary Video 10
Denoised activity in the anterior hindbrain during motion integration. We created the video by reconstituting fluorescent activity from the segmented and calcium-extracted neuronal activity using the CaImAn framework. The video is not aligned to visual stimulation onset or bout events and is slightly sped up to enhance the visibility of bilateral circuit dynamics.
Rights and permissions
About this article
Cite this article
Bahl, A., Engert, F. Neural circuits for evidence accumulation and decision making in larval zebrafish. Nat Neurosci 23, 94–102 (2020). https://doi.org/10.1038/s41593-019-0534-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-019-0534-9
This article is cited by
-
Functional neuronal circuits emerge in the absence of developmental activity
Nature Communications (2024)
-
An optofluidic platform for interrogating chemosensory behavior and brainwide neural representation in larval zebrafish
Nature Communications (2023)
-
Complex computation from developmental priors
Nature Communications (2023)
-
Large-scale F0 CRISPR screens in vivo using MIC-Drop
Nature Protocols (2023)
-
Spatiotemporal visual statistics of aquatic environments in the natural habitats of zebrafish
Scientific Reports (2023)