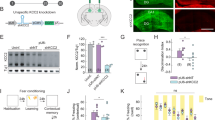
Abstract
An accumulating body of experimental evidence has implicated hippocampal replay occurring within sharp wave ripples (SPW-Rs) as crucial for learning and memory in healthy subjects. This raises speculation that neurological disorders impairing memory disrupt either SPW-Rs or their underlying neuronal activity. We report that mice heterozygous for the gene Scn2a, a site of frequent de novo mutations in humans with intellectual disability, displayed impaired spatial memory. While we observed no changes during encoding, to either single place cells or cell assemblies, we identified abnormalities restricted to SPW-R episodes that manifest as decreased cell assembly reactivation strengths and truncated hippocampal replay sequences. Our results suggest that alterations to hippocampal replay content may underlie disease-associated memory deficits.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
O’Keefe, J. & Dostrovsky, J. The hippocampus as a spatial map. Preliminary evidence from unit activity in the freely-moving rat. Brain Res. 34, 171–175 (1971).
Jones, M. W. & Wilson, M. A. Theta rhythms coordinate hippocampal-prefrontal interactions in a spatial memory task. PLoS Biol. 3, e402 (2005).
Foster, D. J. & Wilson, M. A. Hippocampal theta sequences. Hippocampus 17, 1093–1099 (2007).
Middleton, S. J. & McHugh, T. J. Silencing CA3 disrupts temporal coding in the CA1 ensemble. Nat. Neurosci. 19, 945–951 (2016).
Pfeiffer, B. E. & Foster, D. J. Hippocampal place-cell sequences depict future paths to remembered goals. Nature 497, 74–79 (2013).
Diba, K. & Buzsáki, G. Forward and reverse hippocampal place-cell sequences during ripples. Nat. Neurosci. 10, 1241–1242 (2007).
Skaggs, W. E. & McNaughton, B. L. Replay of neuronal firing sequences in rat hippocampus during sleep following spatial experience. Science 271, 1870–1873 (1996).
Harris, K. D., Csicsvari, J., Hirase, H., Dragoi, G. & Buzsáki, G. Organization of cell assemblies in the hippocampus. Nature 424, 552–556 (2003).
van de Ven, G. M., Trouche, S., McNamara, C. G., Allen, K. & Dupret, D. Hippocampal offline reactivation consolidates recently formed cell assembly patterns during sharp wave-ripples. Neuron 92, 968–974 (2016).
Girardeau, G., Benchenane, K., Wiener, S. I., Buzsáki, G. & Zugaro, M. B. Selective suppression of hippocampal ripples impairs spatial memory. Nat. Neurosci. 12, 1222–1223 (2009).
Ego-Stengel, V. & Wilson, M. A. Disruption of ripple-associated hippocampal activity during rest impairs spatial learning in the rat. Hippocampus 20, 1–10 (2010).
Jadhav, S. P., Kemere, C., German, P. W. & Frank, L. M. Awake hippocampal sharp-wave ripples support spatial memory. Science 336, 1454–1458 (2012).
Cayzac, S. et al. Altered hippocampal information coding and network synchrony in APP-PS1 mice. Neurobiol. Aging 36, 3200–3213 (2015).
Gillespie, A. K. et al. Apolipoprotein E4 causes age-dependent disruption of slow gamma oscillations during hippocampal sharp-wave ripples. Neuron 90, 740–751 (2016).
Nicole, O. et al. Soluble amyloid beta oligomers block the learning-induced increase in hippocampal sharp wave-ripple rate and impair spatial memory formation. Sci. Rep. 6, 22728 (2016).
Planells-Cases, R. et al. Neuronal death and perinatal lethality in voltage-gated sodium channel αII-deficient mice. Biophys. J. 78, 2878–2891 (2000).
Sugawara, T. et al. A missense mutation of the Na+ channel αII subunit gene Na v1.2 in a patient with febrile and afebrile seizures causes channel dysfunction. Proc. Natl. Acad. Sci. USA 98, 6384–6389 (2001).
Kamiya, K. et al. A nonsense mutation of the sodium channel gene SCN2A in a patient with intractable epilepsy and mental decline. J. Neurosci. 24, 2690–2698 (2004).
Rauch, A. et al. Range of genetic mutations associated with severe non-syndromic sporadic intellectual disability: an exome sequencing study. Lancet 380, 1674–1682 (2012).
Carroll, L. S. et al. Mutation screening of SCN2A in schizophrenia and identification of a novel loss-of-function mutation. Psychiatr. Genet. 26, 60–65 (2016).
Neale, B. M. et al. Patterns and rates of exonic de novo mutations in autism spectrum disorders. Nature 485, 242–245 (2012).
Ogiwara, I. et al. De novo mutations of voltage-gated sodium channel αII gene SCN2A in intractable epilepsies. Neurology 73, 1046–1053 (2009).
Sanders, S. J. et al. De novo mutations revealed by whole-exome sequencing are strongly associated with autism. Nature 485, 237–241 (2012).
Yamamoto, J., Suh, J., Takeuchi, D. & Tonegawa, S. Successful execution of working memory linked to synchronized high-frequency gamma oscillations. Cell 157, 845–857 (2014).
Osipova, D. et al. Theta and gamma oscillations predict encoding and retrieval of declarative memory. J. Neurosci. 26, 7523–7531 (2006).
Montgomery, S. M., Sirota, A. & Buzsáki, G. Theta and gamma coordination of hippocampal networks during waking and rapid eye movement sleep. J. Neurosci. 28, 6731–6741 (2008).
Wilson, M. A. & McNaughton, B. L. Reactivation of hippocampal ensemble memories during sleep. Science 265, 676–679 (1994).
Ambrose, R. E., Pfeiffer, B. E. & Foster, D. J. Reverse replay of hippocampal place cells is uniquely modulated by changing reward. Neuron 91, 1124–1136 (2016).
Grosmark, A. D. & Buzsáki, G. Diversity in neural firing dynamics supports both rigid and learned hippocampal sequences. Science 351, 1440–1443 (2016).
Suh, J., Foster, D. J., Davoudi, H., Wilson, M. A. & Tonegawa, S. Impaired hippocampal ripple-associated replay in a mouse model of schizophrenia. Neuron 80, 484–493 (2013).
Tang, W., Shin, J. D., Frank, L. M. & Jadhav, S. P. Hippocampal-prefrontal reactivation during learning is stronger in awake compared with sleep states. J. Neurosci. 37, 11789–11805 (2017).
Carr, M. F., Karlsson, M. P. & Frank, L. M. Transient slow gamma synchrony underlies hippocampal memory replay. Neuron 75, 700–713 (2012).
Iaccarino, H. F. et al. Gamma frequency entrainment attenuates amyloid load and modifies microglia. Nature 540, 230–235 (2016).
Witton, J. et al. Disrupted hippocampal sharp-wave ripple-associated spike dynamics in a transgenic mouse model of dementia. J. Physiol. (Lond.) 594, 4615–4630 (2016).
Davidson, T. J., Kloosterman, F. & Wilson, M. A. Hippocampal replay of extended experience. Neuron 63, 497–507 (2009).
Stark, E. et al. Pyramidal cell-interneuron interactions underlie hippocampal ripple oscillations. Neuron 83, 467–480 (2014).
Foster, D. J. Replay comes of age. Annu. Rev. Neurosci. 40, 581–602 (2017).
Roux, L., Hu, B., Eichler, R., Stark, E. & Buzsáki, G. Sharp wave ripples during learning stabilize the hippocampal spatial map. Nat. Neurosci. 20, 845–853 (2017).
Schomburg, E. W. et al. Theta phase segregation of input-specific gamma patterns in entorhinal-hippocampal networks. Neuron 84, 470–485 (2014).
English, D. F. et al. Excitation and inhibition compete to control spiking during hippocampal ripples: intracellular study in behaving mice. J. Neurosci. 34, 16509–16517 (2014).
Gan, J., Weng, S. M., Pernía-Andrade, A. J., Csicsvari, J. & Jonas, P. Phase-locked inhibition, but not excitation, underlies hippocampal ripple oscillations in awake mice in vivo. Neuron 93, 308–314 (2017).
Buzsáki, G. Hippocampal sharp wave-ripple: a cognitive biomarker for episodic memory and planning. Hippocampus 25, 1073–1188 (2015).
Sullivan, D. et al. Relationships between hippocampal sharp waves, ripples, and fast gamma oscillation: influence of dentate and entorhinal cortical activity. J. Neurosci. 31, 8605–8616 (2011).
Cabral, H. O. et al. Oscillatory dynamics and place field maps reflect hippocampal ensemble processing of sequence and place memory under NMDA receptor control. Neuron 81, 402–415 (2014).
Schmitzer-Torbert, N., Jackson, J., Henze, D., Harris, K. & Redish, A. D. Quantitative measures of cluster quality for use in extracellular recordings. Neuroscience 131, 1–11 (2005).
McHugh, T. J., Blum, K. I., Tsien, J. Z., Tonegawa, S. & Wilson, M. A. Impaired hippocampal representation of space in CA1-specific NMDAR1 knockout mice. Cell 87, 1339–1349 (1996).
Varga, C., Golshani, P. & Soltesz, I. Frequency-invariant temporal ordering of interneuronal discharges during hippocampal oscillations in awake mice. Proc. Natl. Acad. Sci. USA 109, E2726–E2734 (2012).
Fernandez-Ruiz, A. et al. Entorhinal-CA3 dual-input control of spike timing in the hippocampus by theta-gamma coupling. Neuron 93, 1213–1226.e1215 (2017).
Berens, P. CircStat: a MATLAB toolbox for circular statistics. J. Stat. Softw. 31, https://doi.org/10.18637/jss.v031.i10 (2009).
Skaggs, W. E., McNaughton, B. L., Wilson, M. A. & Barnes, C. A. Theta phase precession in hippocampal neuronal populations and the compression of temporal sequences. Hippocampus 6, 149–172 (1996).
Mizuseki, K., Sirota, A., Pastalkova, E. & Buzsáki, G. Theta oscillations provide temporal windows for local circuit computation in the entorhinal-hippocampal loop. Neuron 64, 267–280 (2009).
Trouche, S. et al. Recoding a cocaine-place memory engram to a neutral engram in the hippocampus. Nat. Neurosci. 19, 564–567 (2016).
Peyrache, A., Benchenane, K., Khamassi, M., Wiener, S. I. & Battaglia, F. P. Principal component analysis of ensemble recordings reveals cell assemblies at high temporal resolution. J. Comput. Neurosci. 29, 309–325 (2010).
Marchini, J. L., Heaton, C. & Ripley, B. D. FastICA: fastICA algorithms to perform ICA and projection pursuit. https://CRAN.R-project.org/package=fastICA (2013).
McNamara, C. G., Tejero-Cantero, Á., Trouche, S., Campo-Urriza, N. & Dupret, D. Dopaminergic neurons promote hippocampal reactivation and spatial memory persistence. Nat. Neurosci. 17, 1658–1660 (2014).
Silva, D., Feng, T. & Foster, D. J. Trajectory events across hippocampal place cells require previous experience. Nat. Neurosci. 18, 1772–1779 (2015).
Acknowledgements
We thank all members of the CBP laboratory for their support and the RIKEN Advanced Manufacturing Team for their assistance in microdrive production. This work was supported by NIH grant GM49711 (M.M.), JSPS Kakenhi 26750378 (S.J.M.), JSPS Kakenhi 21791020 and MEXT 25461572 (I.O.), JSPS Kakenhi 17H05986 (T.J.M), SRPBS from MEXT and AMED (K.Y.) and RIKEN BSI (K.Y. and T.J.M.).
Author information
Authors and Affiliations
Contributions
S.J.M, T.J.M. and K.Y. conceived the study. I.O. performed Barnes maze experiments. S.J.M. and E.M.K. performed all other experiments. S.J.M. and S.C. analyzed the data. M.M. created the transgenic mouse. S.J.M. and T.J.M. wrote the manuscript. Lead contact T.J.M.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated Supplementary Information
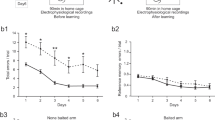
Supplementary Figure 1 Spatial learning, place fields and tetrode locations.
(a) Latencies during Barnes maze learning over a four day period revealed control mice (blue) learned faster than Scn2a+/- mice (red) (P = 0.0027, F = 5.19, 2-way repeated measures ANOVA, followed by post-hoc Bonferroni test P<0.01 on day 3; CTR: N = 14 animals; SCN: N = 14 animals). (b) The spatial receptive fields are shown for all recorded place cells during linear track exploration sessions. Individual grey lines represent single place cell rate maps across the entirety of the track, dark red areas indicate place fields determined as described in the methods section. Control (left) and Scn2a+/- (right) fields are ordered according to their peak position on the track. (c) Reconstructed tetrode tip locations from three animals of each genotype (blue CTR, pink Scn2a+/-) overlaid onto schematic mouse brain slices, numbers indicate posterior distances from bregma. Tetrodes circled in red were excluded as they were either poorly targeted or were devoid of place cells. All boxplots represent the median (black line) and the 25th-75th percentiles, with whiskers extending to the most extreme data points, excluding outliers which are plotted as individual crosses. *P < 0.05.
Supplementary Figure 2 Sharp wave ripple event detection.
(a) Upper traces (WB) show five examples of raw CA1 stratum pyramidale local field potentials recorded from control mice during a rest period in a small sleep box. Blue traces represent activity centered on events detected as SPW-Rs, with the period detected as a ripple (as described in the methods section) highlighted in red. Below these are the same LFPs band-pass filtered between 80–200 Hz. The bottom panel shows multi-unit activity of all clustered place cells for the corresponding time periods. (b) Shows the same but for Scn2a+/- mice. Scale bars represent 100 ms (horizontal) and 100 µV (vertical) throughout.
Supplementary Figure 3 Detection of cell assemblies.
(a) Session example showing assembly spatial maps (of 12 cell assemblies) ordered from left to right across the entirety of the linear track. (b) Corresponding data showing the 57 CA1 place cells recorded during this session, each neurons weight in a given assembly is indicated by its deviation on the x-axis. Cells with a significant weight are color coded to the assembly to which they belong (from a). (c) Mean assembly strength during linear track exploration did not differ between genotypes (P = 0.83, Z = 0.22, two-sided Wilcoxon Rank-sum; CTR: N = 7 animals, n = 96 cell assemblies; SCN: N = 7 animals, n = 132 cell assemblies). (d) Cells classified as forming part of an assembly had significantly higher co-firing coefficients (r) with assembly partners, than cells which did not form part of any assembly (P = 3.32 × 10−23, Z = 9.92, two-sided Wilcoxon Rank-sum; n = 59 assembly cell pairs, n = 105 non-assembly cell pairs), relating to example session shown in a & b. (e) Place-field similarity was also significantly greater for assembly member pairs, compared with non-members (P = 1.62 × 10−16, Z = 8.25, two-sided Wilcoxon Rank-sum; n = 59 assembly cell pairs, n = 105 non-assembly cell pairs). (f) Across all subjects during SPW-Rs, cell pairs forming part of an assembly had consistently higher co-firing coefficients independent of genotype (CTRs, P = 1.28 × 10−52, Z = 15.3, two-sided Wilcoxon Rank-sum; SCN, P = 7.82 × 10−67, Z = 17.3, two-sided Wilcoxon Rank-sum; CTR: N = 7 animals; SCN: N = 7 animals), when compared to non-members. (g) Place field similarity was also significantly higher for cell-assembly members for both controls (P = 1.25 × 10−21, Z = 9.55, two-sided Wilcoxon Rank-sum, N = 7 animals) and mutants (P = 4.31 × 10−25, Z = 10.3, two-sided Wilcoxon Rank-sum, N = 7 animals). Cell-assembly members (P = 0.31, Z = 1.02, two-sided Wilcoxon Rank-sum; CTR: N = 7 animals; SCN: N = 7 animals) and non-members (P = 0.73, Z = 0.35, two-sided Wilcoxon Rank-sum; CTR: N = 7 animals; SCN: N = 7 animals) were not significantly different when compared across genotype. All boxplots represent the median (black line) and the 25th-75th percentiles, with whiskers extending to the most extreme data points, excluding outliers which are plotted as individual crosses. *P < 0.05.
Supplementary Figure 4 Spike waveforms and Bayesian decoding of position.
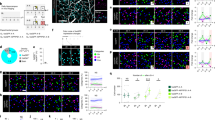
(a) Left panel shows decoding accuracy for a full linear track exploration session, with hot colors indicating highest probabilities and high probabilities along the diagonal demonstrating decoded positions accurately reflect the real world positon of the animal. Middle panel shows multiple traversals of the linear track using the Bayesian decoder, hot colors representing highest probability, red line indicates true position and is shifted vertically for clarity. Right panel shows the mean decoding error for both control (blue) and Scn2a+/- mice (red), with individual points representing single animals (P = 0.99, two-sided Wilcoxon Rank-sum; CTR: N = 7 animals; SCN: N = 7 animals). (b) Example spike waveforms recorded from control mice in CA1 stratum pyramidale, for both neurons classified as both pyramidal cells (top left) and parvalbumin-like interneurons (top right). Middle panel (blue) shows average waveforms for all pyramidal (left) and PV-like (right) neurons from control mice. Scn2a+/- average waveforms are shown in the bottom panel (red) with pyramidal left and PV-like to the right, shaded areas represent SEM. (c) Pyramidal (blue) and PV-like neurons (red) were separable into two distinct clusters based on waveform asymmetry and firing frequency, however within cell types, genotypes (control and Scn2a+/- are represented by dark and light colors respectively) were indistinguishable. (d) Example phase precession plots for three place cells from each genotype (control and Scn2a+/- in blue and red respectively), showing spikes moving to progressively earlier phases with increasing passage into place fields. (e) Histograms showing the relative frequency of all place cells relative to LFP theta, with positive values indicate phase precession of cells (P = 0.24, Z = 1.16, two-sided Wilcoxon Rank-sum; CTR: N = 7 animals, n = 224 cells; SCN: N = 7 animals, n = 274 cells). All boxplots represent the median (black line) and the 25th-75th percentiles, with whiskers extending to the most extreme data points, excluding outliers which are plotted as individual crosses. *P < 0.05.
Supplementary Figure 5 Awake replay events are truncated in a similar manner to sleep replays.
(a) Cumulative distribution of sequence scores (rZ) for events detected by both MUA and ripple power, the means ± SEM are displayed in the inset (P = 0.33, Z = 0.98, two-sided Wilcoxon Rank-sum; CTR: N = 7 animals, n = 241 events; SCN: N = 7 animals, n = 255 events). (b) Group data showing mean distances spanned by replay events detected by a combination of both MUA and ripple power (P = 2.4 × 10−5, Z = 4.23, two-sided Wilcoxon Rank-sum; CTR: N = 7 animals, n = 241 events; SCN: N = 7 animals, n = 255 events). (c) Average speed in m/s of progression from starting position to end position for all replay events detected by MUA and ripple power (P = 4.1 × 10−4, Z = 3.5, two-sided Wilcoxon Rank-sum; CTR: N = 7 animals, n = 241 events; SCN: N = 7 animals, n = 255 events). (d) The average spatial jump in centimeters for decoded positons in adjacent temporal bins, for each replay event (P = 0.016, Z = 2.41, two-sided Wilcoxon Rank-sum; CTR: N = 7 animals, n = 241 events; SCN: N = 7 animals, n = 255 events). (e) Mean z-scored LFP ripple power concurrent with all significant replay events, lined up to the time of initial detection of the events (CTR: N = 7 animals, n = 241 events; SCN: N = 7 animals, n = 255 events), control (blue) and Scn2a+/- (red) are represented by their respective colors throughout (a–e). (f) Examples of trajectory sequences derived from Bayesian decoding of spiking activity during awake SPW-Rs, top row represents controls, beneath are Scn2a+/- examples. (g) Representative examples of unfiltered SPW-Rs during the awake exploratory session, for both control (blue) and Scn2a+/- (red). (h) Mean replay event sequence scores (rZ) for control (blue) and Scn2a+/- (red) in the awake exploratory session (P = 0.97, Z = 0.03, two-sided Wilcoxon Rank-sum; CTR: N = 7 animals, n = 59 events; SCN: N = 7 animals, n = 88 events). (i) Group data showing mean distances spanned by awake replay events detected by a combination of both MUA and ripple power (P = 0.044, Z = 2.0, two-sided Wilcoxon Rank-sum; CTR: N = 7 animals, n = 59 events; SCN: N = 7 animals, n = 88 events). (j) Average speed in m/s of progression from starting position to end position for all awake replay events detected by MUA and ripple power (P = 0.027, Z = 2.2, two-sided Wilcoxon Rank-sum; CTR: N = 7 animals, n = 59 events; SCN: N = 7 animals, n = 88 events). (k) Mean z-scored LFP ripple power concurrent with all significant awake replay events, lined up to the time of initial detection (combined ripple power and MUA), control and Scn2a+/- are represented by blue and red respectively (CTR: N = 7 animals, n = 59 events; SCN: N = 7 animals, n = 88 events). All boxplots represent the median (black line) and the 25th–75th percentiles, with whiskers extending to the most extreme data points, excluding outliers which are plotted as individual crosses. *P < 0.05.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Table 1
Rights and permissions
About this article
Cite this article
Middleton, S.J., Kneller, E.M., Chen, S. et al. Altered hippocampal replay is associated with memory impairment in mice heterozygous for the Scn2a gene. Nat Neurosci 21, 996–1003 (2018). https://doi.org/10.1038/s41593-018-0163-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-018-0163-8
This article is cited by
-
Identification of common core ion channel genes in epilepsy and Alzheimer’s disease
Irish Journal of Medical Science (1971 -) (2024)
-
Inversed Effects of Nav1.2 Deficiency at Medial Prefrontal Cortex and Ventral Tegmental Area for Prepulse Inhibition in Acoustic Startle Response
Molecular Neurobiology (2024)
-
Microglial over-pruning of synapses during development in autism-associated SCN2A-deficient mice and human cerebral organoids
Molecular Psychiatry (2024)
-
Phenotypes, mechanisms and therapeutics: insights from bipolar disorder GWAS findings
Molecular Psychiatry (2022)
-
SCN2A channelopathies in the autism spectrum of neuropsychiatric disorders: a role for pluripotent stem cells?
Molecular Autism (2020)