Abstract
The high-order three-dimensional (3D) organization of regulatory genomic elements provides a topological basis for gene regulation, but it remains unclear how multiple regulatory elements across the mammalian genome interact within an individual cell. To address this, herein, we developed scNanoHi-C, which applies Nanopore long-read sequencing to explore genome-wide proximal high-order chromatin contacts within individual cells. We show that scNanoHi-C can reliably and effectively profile 3D chromatin structures and distinguish structure subtypes among individual cells. This method could also be used to detect genomic variations, including copy-number variations and structural variations, as well as to scaffold the de novo assembly of single-cell genomes. Notably, our results suggest that extensive high-order chromatin structures exist in active chromatin regions across the genome, and multiway interactions between enhancers and their target promoters were systematically identified within individual cells. Altogether, scNanoHi-C offers new opportunities to investigate high-order 3D genome structures at the single-cell level.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The sequencing data of scNanoHi-C have been deposited in the Gene Expression Omnibus under accession code GSE217189. All other published datasets and annotation files used in this study are summarized in Supplementary Table 11. Source data files have been deposited in the figshare database at https://doi.org/10.6084/m9.figshare.23566704 (ref. 72). Source data are provided with this paper.
Code availability
Codes related to this paper can be accessed at https://github.com/LuJiansen/scNanoHi-C. Codes are also deposited into Zenodo (https://doi.org/10.5281/zenodo.7769419)73.
References
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Dekker, J. & Mirny, L. The 3D genome as moderator of chromosomal communication. Cell 164, 1110–1121 (2016).
Dixon, J. R., Gorkin, D. U. & Ren, B. Chromatin domains: the unit of chromosome organization. Mol. Cell 62, 668–680 (2016).
Lin, X. et al. Nested epistasis enhancer networks for robust genome regulation. Science 377, 1077–1085 (2022).
Fudenberg, G. et al. Formation of chromosomal domains by loop extrusion. Cell Rep. 15, 2038–2049 (2016).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Dekker, J., Rippe, K., Dekker, M. & Kleckner, N. Capturing chromosome conformation. Science 295, 1306–1311 (2002).
Oudelaar, A. M. & Higgs, D. R. The relationship between genome structure and function. Nat. Rev. Genet. 22, 154–168 (2021).
Ay, F. et al. Identifying multi-locus chromatin contacts in human cells using tethered multiple 3C. BMC Genom. 16, 121 (2015).
Olivares-Chauvet, P. et al. Capturing pairwise and multi-way chromosomal conformations using chromosomal walks. Nature 540, 296 (2016).
Darrow, E. M. et al. Deletion of DXZ4 on the human inactive X chromosome alters higher-order genome architecture. Proc. Natl Acad. Sci. USA 113, E4504–E4512 (2016).
Allahyar, A. et al. Enhancer hubs and loop collisions identified from single-allele topologies. Nat. Genet. 50, 1151–1160 (2018).
Oudelaar, A. M. et al. Single-allele chromatin interactions identify regulatory hubs in dynamic compartmentalized domains. Nat. Genet. 50, 1744–1751 (2018).
Quinodoz, S. A. et al. Higher-order inter-chromosomal hubs shape 3D genome organization in the nucleus. Cell 174, 744–757 (2018).
Zheng, M. et al. Multiplex chromatin interactions with single-molecule precision. Nature 566, 558–562 (2019).
Beagrie, R. A. et al. Complex multi-enhancer contacts captured by genome architecture mapping. Nature 543, 519–524 (2017).
Deshpande, A. S. et al. Identifying synergistic high-order 3D chromatin conformations from genome-scale nanopore concatemer sequencing. Nat. Biotechnol. 40, 1488–1499 (2022).
Nagano, T. et al. Single-cell Hi-C reveals cell-to-cell variability in chromosome structure. Nature 502, 59–64 (2013).
Flyamer, I. M. et al. Single-nucleus Hi-C reveals unique chromatin reorganization at oocyte-to-zygote transition. Nature 544, 110–114 (2017).
Nagano, T. et al. Cell-cycle dynamics of chromosomal organization at single-cell resolution. Nature 547, 61–67 (2017).
Ramani, V. et al. Massively multiplex single-cell Hi-C. Nat. Methods 14, 263–266 (2017).
Stevens, T. J. et al. 3D structures of individual mammalian genomes studied by single-cell Hi-C. Nature 544, 59 (2017).
Tan, L., Xing, D., Chang, C. H., Li, H. & Xie, X. S. Three-dimensional genome structures of single diploid human cells. Science 361, 924–928 (2018).
Bintu, B. et al. Super-resolution chromatin tracing reveals domains and cooperative interactions in single cells. Science https://doi.org/10.1126/science.aau1783 (2018).
Arrastia, M. V. et al. Single-cell measurement of higher-order 3D genome organization with scSPRITE. Nat. Biotechnol. 40, 64–73 (2021).
Akgol Oksuz, B. et al. Systematic evaluation of chromosome conformation capture assays. Nat. Methods 18, 1046–1055 (2021).
Lafontaine, D. L., Yang, L., Dekker, J. & Gibcus, J. H. Hi-C 3.0: improved protocol for genome-wide chromosome conformation capture. Curr. Protoc. 1, e198 (2021).
Nollmann, M., Bennabi, I., Gotz, M. & Gregor, T. The impact of space and time on the functional output of the genome. Cold Spring Harb. Perspect. Biol. https://doi.org/10.1101/cshperspect.a040378 (2022).
Hafner, A. & Boettiger, A. The spatial organization of transcriptional control. Nat. Rev. Genet. https://doi.org/10.1038/s41576-022-00526-0 (2022).
Sanborn, A. L. et al. Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc. Natl Acad. Sci. USA 112, E6456–E6465 (2015).
Gabriele, M. et al. Dynamics of CTCF- and cohesin-mediated chromatin looping revealed by live-cell imaging. Science 376, 496–501 (2022).
Mach, P. et al. Cohesin and CTCF control the dynamics of chromosome folding. Nat. Genet. 54, 1907–1918 (2022).
Yu, M. et al. SnapHiC: a computational pipeline to identify chromatin loops from single-cell Hi-C data. Nat. Methods 18, 1056–1059 (2021).
Xie, W. J. et al. Structural modeling of chromatin integrates genome features and reveals chromosome folding principle. Sci. Rep. 7, 2818 (2017).
Boyle, S. et al. The spatial organization of human chromosomes within the nuclei of normal and emerin-mutant cells. Hum. Mol. Genet 10, 211–219 (2001).
Lee, J. T. & Bartolomei, M. S. X-inactivation, imprinting, and long noncoding RNAs in health and disease. Cell 152, 1308–1323 (2013).
Negrini, S., Gorgoulis, V. G. & Halazonetis, T. D. Genomic instability—an evolving hallmark of cancer. Nat. Rev. Mol. Cell Biol. 11, 220–228 (2010).
Sanders, A. D. et al. Single-cell analysis of structural variations and complex rearrangements with tri-channel processing. Nat. Biotechnol. 38, 343–354 (2020).
Zhou, Y. et al. Single-cell multiomics sequencing reveals prevalent genomic alterations in tumor stromal cells of human colorectal cancer. Cancer Cell 38, 818–828 (2020).
Li, R. et al. A body map of somatic mutagenesis in morphologically normal human tissues. Nature 597, 398–403 (2021).
Fu, Y. et al. Uniform and accurate single-cell sequencing based on emulsion whole-genome amplification. Proc. Natl Acad. Sci. USA 112, 11923–11928 (2015).
Chen, C. et al. Single-cell whole-genome analyses by linear amplification via transposon insertion (LIANTI). Science 356, 189–194 (2017).
Zong, C., Lu, S., Chapman, A. R. & Xie, X. S. Genome-wide detection of single-nucleotide and copy-number variations of a single human cell. Science 338, 1622–1626 (2012).
Zhou, B. et al. Comprehensive, integrated, and phased whole-genome analysis of the primary ENCODE cell line K562. Genome Res. 29, 472–484 (2019).
Berger, M. F. et al. Integrative analysis of the melanoma transcriptome. Genome Res. 20, 413–427 (2010).
Zhou, T., Zhang, R. & Ma, J. The 3D Genome Structure of Single Cells. Annu Rev. Biomed. Data Sci. 4, 21–41 (2021).
Fulco, C. P. et al. Activity-by-contact model of enhancer-promoter regulation from thousands of CRISPR perturbations. Nat. Genet. 51, 1664–1669 (2019).
Zhou, H. et al. Epstein-Barr virus oncoprotein super-enhancers control B cell growth. Cell Host Microbe 17, 205–216 (2015).
Ernst, J. & Kellis, M. Chromatin-state discovery and genome annotation with ChromHMM. Nat. Protoc. 12, 2478–2492 (2017).
Wu, S. et al. Circular ecDNA promotes accessible chromatin and high oncogene expression. Nature 575, 699–703 (2019).
Hung, K. L. et al. ecDNA hubs drive cooperative intermolecular oncogene expression. Nature 600, 731–736 (2021).
Zhu, Y. et al. Oncogenic extrachromosomal DNA functions as mobile enhancers to globally amplify chromosomal transcription. Cancer Cell 39, 694–707 (2021).
Burton, J. N. et al. Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions. Nat. Biotechnol. 31, 1119–1125 (2013).
Xie, H. et al. De novo assembly of human genome at single-cell levels. Nucleic Acids Res. 50, 7479–7492 (2022).
Furlong, E. E. M. & Levine, M. Developmental enhancers and chromosome topology. Science 361, 1341–1345 (2018).
Sabari, B. R. et al. Coactivator condensation at super-enhancers links phase separation and gene control. Science https://doi.org/10.1126/science.aar3958 (2018).
Guo, Y. E. et al. Pol II phosphorylation regulates a switch between transcriptional and splicing condensates. Nature 572, 543–548 (2019).
Li, J. et al. Single-gene imaging links genome topology, promoter-enhancer communication and transcription control. Nat. Struct. Mol. Biol. 27, 1032–1040 (2020).
Bhat, P., Honson, D. & Guttman, M. Nuclear compartmentalization as a mechanism of quantitative control of gene expression. Nat. Rev. Mol. Cell Biol. 22, 653–670 (2021).
Bao, S. et al. Epigenetic reversion of post-implantation epiblast to pluripotent embryonic stem cells. Nature 461, 1292–1295 (2009).
Kechin, A., Boyarskikh, U., Kel, A. & Filipenko, M. cutPrimers: a new tool for accurate cutting of primers from reads of targeted next generation sequencing. J. Comput Biol. 24, 1138–1143 (2017).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. https://doi.org/10.1038/nbt.4314 (2018).
Wang, X. et al. Genome-wide detection of enhancer-hijacking events from chromatin interaction data in rearranged genomes. Nat. Methods 18, 661–668 (2021).
Smith, T., Heger, A. & Sudbery, I. UMI-tools: modeling sequencing errors in unique molecular identifiers to improve quantification accuracy. Genome Res. 27, 491–499 (2017).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Boeva, V. et al. Control-FREEC: a tool for assessing copy number and allelic content using next-generation sequencing data. Bioinformatics 28, 423–425 (2012).
Dixon, J. R. et al. Integrative detection and analysis of structural variation in cancer genomes. Nat. Genet. 50, 1388–1398 (2018).
Ghurye, J. et al. Integrating Hi-C links with assembly graphs for chromosome-scale assembly. PLoS Comput. Biol. 15, e1007273 (2019).
Gurevich, A., Saveliev, V., Vyahhi, N. & Tesler, G. QUAST: quality assessment tool for genome assemblies. Bioinformatics 29, 1072–1075 (2013).
Manni, M., Berkeley, M. R., Seppey, M., Simao, F. A. & Zdobnov, E. M. BUSCO update: novel and streamlined workflows along with broader and deeper phylogenetic coverage for scoring of eukaryotic, prokaryotic, and viral genomes. Mol. Biol. Evol. 38, 4647–4654 (2021).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Li, W. et al. scNanoHi-C: a single-cell long-read concatemer sequencing method to reveal high-order chromatin structures within individual cells. figshare https://doi.org/10.6084/m9.figshare.23566704 (2023).
Li, W. et al. scNanoHi-C. Zenodo https://doi.org/10.5281/zenodo.7769419 (2023).
Acknowledgements
We thank the Beijing Advanced Innovation Center for Genomics for support and part of the analysis was performed on the High-Performance Computing Platform of the Center for Life Sciences (Peking University). We thank the staff at the Peking University High-throughput Sequencing Center for help with flow cytometry. We thank D. Xing, Q. Xia, Y. Chen and Z. Liu (Peking University) for helpful suggestions for the Hi-C experiments. This work was supported by the Changping Laboratory, Beijing Advanced Innovation Center for Genomics at Peking University and the National Natural Science Foundation of China (32288102 to F.T.).
Author information
Authors and Affiliations
Contributions
F.T., W.L. and J. Lu conceived the project. W.L. was in charge of the experimental part and developed the scNanoHi-C protocol for the Nanopore platform with the help of J. Lu. Y.B. offered help for DNA FISH. J. Lu was in charge of the bioinformatics analysis with the help of W.L. and P.L. W.L., J. Lu and F.T. wrote the paper. Y.G., K.C., X.S., M.L., J. Liu, Y.C. and L.W. offered suggestions.
Corresponding author
Ethics declarations
Competing interests
F.T., W.L. and J. Lu are investors on a patent that covers scNanoHi-C. The other authors declare no competing interests.
Peer review
Peer review information
Nature Methods thanks Bing Ren and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Lei Tang and Hui Hua, in collaboration with the Nature Methods team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
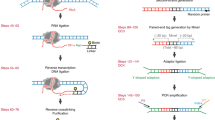
Extended Data Fig. 1 The detailed experimental process of scNanoHi-C.
a. Schematic diagram for detailed experimental procedure and principle of scNanoHi-C. b. The length distribution of fragments after MboI digestion (left), proximity ligation (medium) and amplification (right), respectively. c. Collisions rate of the whole experiment (left, n = 96) and permeabilized nuclei during amplification (right, n = 192). We identified collision cells as those in which less than 95% of monomers mapped to the mouse or to human genome (Supplementary Text 3).
Extended Data Fig. 2 The quality control of scNanoHi-C data.
a. An overview of data statistics of merged high-, medium-, and low-depth scNanoHi-C for GM12878. b. Composition of contacts of each single-cell from high-, medium- and low-depth data, respectively. Contacts were colored by their assigned groups during filtration, including adjacent, close, duplicate, promiscuous, isolated, and passed contacts. c. Composition of duplicate sets in each cell from high-, medium- and low-depth data, respectively. Duplicate sets were colored based on their group. ‘Identical’ means all concatemers are the same within the set, ‘included’ means all other contacts are included in the concatemer with the highest cardinality, and ‘others’ are the remaining duplicate sets. d. Boxplot showed the percentage of contacts within a duplicate set that were included in the concatemer with the highest cardinality, related to the ‘others’ group of (c). e-i. Total bases after removal of sequencing adaptors (e), ratio of passed contacts (f), ratio of trans-chromosomal contacts (g), the number of contacts (h), and ratio of high-order concatemers (i) in single-cell of high-, medium-, low-depth scNanoHi-C. j. The coverage of high-, medium- and low-depth scNanoHi-C in different resolutions (50 kb, 100 kb, 500 kb, and 1 Mb, respectively). The n = 24, 192 and 456 biologically independent cells for high-, medium- and low-depth scNanoHi-C, respectively (d-j). From left to right are the number of monomers (left) and contacts (middle) detected in each bin and the percentage of covered bins in each cell. The central lines of boxplots are median of data. The lower and upper hinges correspond to the 25th and 75th percentiles. The end of lower and upper whiskers are 1.5 * IQR (inter-quartile range). Data beyond the end of the whiskers are plotted as outliers (d-j).
Extended Data Fig. 3 Detecting 3D genome structures in merged scNanoHi-C.
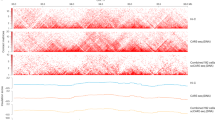
a, b.(i) Correlation of 1-Mb raw contacts between merged high-, medium-depth scNanoHi-C and bulk Hi-C for GM12878 (r = 0.96, r = 0.96). (ii) Correlation of 1-Mb CS between merged high-, medium-depth scNanoHi-C and bulk Hi-C for GM12878 (r = 0.85, r = 0.93). (iii) Correlation of 50-kb IS between merged high-, medium-depth scNanoHi-C and bulk Hi-C for GM12878 (r = 0.78, r = 0.88). (iv) Comparison of aggregate peak analysis (APA) for merged high-, medium-depth scNanoHi-C virtual pair-wise contacts and bulk Hi-C pair-wise contact density within 100 kb of Hi-C loop anchors. c. Dependence of contact probability on genomic distance, P(s), for bulk Hi-C, low-, medium-, high-depth scNanoHi-C. d. Derivative of the P(s) plots from c. e. Dependence of normalized contact probability on genomic distance for bulk Hi-C, low-, medium-, high-depth scNanoHi-C.
Extended Data Fig. 4 Comparisons between scNanoHi-C and scSPRITE.
a. Distribution of cardinality at concatemer/cluster levels (left) and monomer/read levels (right) in scNanoHi-C (top) and scSPRITE (bottom), respectively. b. Distribution of cardinality at the cluster (left) and read (right) levels in single cells of scNanoHi-C (top) and scSPRITE (bottom). Cells were ordered by the number of contacts and clusters for scNanoHi-C and scSPRITE, respectively. Clusters or reads were grouped by their cardinality. c. The decay of normalized contacts probability as a function of genomic distance separation for single cells in scNanoHi-C and scSPRITE. Contacts from each cell were grouped by their cardinality, the lines and shadows represent the mean and 25% to 75% quantiles of normalized contacts probability among all cells, respectively. The black line represents the result of bulk in situ Hi-C. d. The ratio of trans-chromosomal contacts (trans-ratio) in single cells of scNanoHi-C and scSPRITE (left). The trans-ratios of scSPRITE were further calculated by each cardinality group (right). The n = 1,000, 288 and 288 biologically independent cells for scSPRITE, medium- and low-depth scNanoHi-C, respectively. The central lines of boxplots are the median of data. The lower and upper hinges correspond to the 25th and 75th percentiles. The end of the lower and upper whiskers are 1.5 * IQR (inter-quartile range). e. Aggregate peak analysis (APA) for mESC of bulk in situ HiC (top), scSPRITE (middle) and scNanoHi-C (bottom) within 100 kb on either side of loop anchors identified from bulk Hi-C. Contacts of scSPRITE and scNanoHi-C were merged according their cardinality. The number on the upper left corner represents the enrichment score of contacts in the central pixel.
Extended Data Fig. 5 Detecting 3D genome structures in single cells using scNanoHi-C data.
a–c. Distribution of average normalized detection scores (nDS) across all cells in each pairs of chromosome territories (n = 253, 1-Mb resolution, a), A/B compartment switching triplets (A-B-A or B-A-B, n = 630, 1-Mb resolution, b) and TAD boundaries (n = 2,503, 50-kb resolution, c) of GM12878 scNanoHi-C data; score = 0 (red line). Representative structures from a medium- and low-depth GM12878 scNanoHi-C cell were shown at the right. All chromosome territories (chr1 to chr22 and chrX) in a (ii) was visulazed at 5-Mb resolution. d. Percentage of cells with TAD nDS > 0 for each TAD boundary; e. Correlation between average TAD nDSs and TAD boundary strength of each TAD boundary. Spearman’s correlation was calculated and tested by ‘cor.test’ function in R with default two-sided test. f. comparison between the average TAD normalized detection scores in boundaries across 672 biologically independent cells with (n = 1,506) and without (n = 997) the presence of CTCF binding (inferred from CTCF ChIP-seq peaks). Paired two-side t-test was used to determine the significance. The central lines of boxplots are median of data. The lower and upper hinges correspond to the 25th and 75th percentiles. The end of lower and upper whiskers are 1.5 * IQR (inter-quartile range). Data beyond the end of the whiskers are plotted as outliers.
Extended Data Fig. 6 Identifying loops with scNanoHi-C.
a. Information about the datasets used for detecting loops with SnapHiC and the number of detected loops in each dataset. b. Track view of WashU epigenome browser showing two representative clusters of loops around MIR155HG and RUNX3. From top to bottom are refGene and repeatMasker annotations, ChIP-seq signals of CTCF, H3K4me3, H3K27ac, and RAD21, loops identified from scNanoHi-C, cohesin HChIP (called with FitHiChIP), cohesin HiChIP (called with MAPS), H3K27ac HiChIP (called with FitHiChIP), H3K27ac HiChIP (called with MAPS) and in situ Hi-C (called with HiCCUPS). Super-enhancers were highlighted with orange shadows. c. Track view of WashU epigenome browser showing two representative clusters of loops around Sox2 and Nanog. From top to bottom are refGene and repeatMasker annotations, ChIP-seq signals of H3K4me1, H3K4me3, and H3K27ac, loops identified from scNanoHi-C, cohesin HChIP (called with FitHiChIP) and in situ Hi-C (called with HiCCUPS).
Extended Data Fig. 7 Reconstruction of the 3D genome structure model in single cells using scNanoHi-C data.
a. The distribution of median monomer length of merged high-, medium-, low-depth scNanoHi-C for GM12878 (left); the distribution of median concatemer length of merged high-, medium-, low-depth scNanoHi-C for GM12878 (right). b. The distribution of median concatemer length of a representative single cell (left); the distribution of median monomer length of a representative single cell (right). c. Semi-phased (left) and full-phased (right) percentage of monomers in Dip-C (n = 16) and low- (n = 456), medium- (n = 192), high-depth (n = 24) scNanoHi-C (left). The central lines of boxplots are median of data. The lower and upper hinges correspond to the 25th and 75th percentiles. The end of lower and upper whiskers are 1.5 * IQR (inter-quartile range). Data beyond the end of the whiskers are plotted as outliers. d. Rank-normalized 1-Mb 3D scA/B values of maternal (red dot) and paternal alleles (blue dot) in GM12878 at chr1 to chr22 and chrX (RMS r.m.s.d. < 1.5, n = 28). e. Radial position of maternal (red dot) and paternal alleles (blue dot) in GM12878, as measured by average distances to the nuclear center (RMS r.m.s.d. < 1.5, n = 28).
Extended Data Fig. 8 Detecting CNVs in single cells using scNanoHi-C data.
a. CNVs of merged GM12878 cells at in 1-Mb resolution, copy numbers across the genome are plotted in merged GM12878 cells from high-, medium-, low-depth of scNanoHi-C. b. The relationship between numbers of contacts and CV at different resolutions of GM12878 scNanoHi-C data. c. The heat map of CNV profiles in GM12878 scNanoHi-C single cells without clear clonal CNVs (contacts > 100k, n = 169),1-Mb windows (left); CNV profiles of MALBAC GM12878 single cells (n = 147, number of reads > 1 million, CV < 0.25, left). Cells without and with CNVs (length > 20 Mb, supported by at least 5 cells) were selected and visualized at the top and bottom, respectively. d. The average distance of particles within paternal and maternal chr1 (left, n = 25,200 particles at 20 kb, P-value was determined by paired two-side t-test), the long-arm (n = 7,260) and short-arm (n = 5,151) of paternal (middle, P-value was determined by unpaired one-side t-test) and maternal (right, P-value was determined by unpaired one-side t-test) chr1 of the same cell mentioned in Fig. 3g. The central lines of boxplots are the median of data. The lower and upper hinges correspond to the 25th and 75th percentiles. The end of the lower and upper whiskers are 1.5 * IQR (inter-quartile range). e. The relationship between numbers of contacts and CV at different resolutions of K562 scNanoHi-C data. f. CNV profiles of K562 WGS, merged scNanoHi-C, bulk Hi-C and three representative single cells of scNanoHi-C at 1-Mb resolution.
Extended Data Fig. 9 Analyses of the relationships between high-order ratios and other factors.
a, b. Heat map showed the partial Spearman’s correlation analyses between the ratio of high-order concatemers and other factors among each 50-kb genomic windows in GM12878 (a.) and K562 (b), respectively. c. Partial Pearson’s correlations between the conditioned ratio of high-order concatemers and the average length of monomers in K562. d. Partial Pearson’s correlations between the conditioned ratio of high-order concatemers and compartment scores in K562. e–f. the distribution of supported concatemers and cells in one enhancer bin regulating multiple promoter bins (e.) and one promoter bin regulated by multiple enhancer bins (f.).
Extended Data Fig. 10 Detecting synergies in single GM12878 cells and single COLO320DM cells.
a. The LOLA enrichment of GM12878 regulatory elements synergies (n = 917) relative to backgrounds (n = 1481); RNAPII, RNA polymerase II. The values of each bar represent log2(odd ratios) of enrichment for each feature, error bars represent the 95% confidence intervals. See also in Methods. b. The DNA FISH of MYC in COLO320DM. The experiment were repeated two times independently with similar results in multiple views. c. The contacts map of COLO320DM in merged scNanoHi-C and single-cell scNanoHi-C data; red triangles, regions of MYC gene amplification. d. The CNVs pattern of single COLO320DM cells, 1-Mb resolution. e. The adjnTIF of merged single COLO320DM cells at genome-wide scale. f. Single−cell averaged adjnTIF of COLO320DM at genome-wide scale. g. AdjnTIF of ecDNA region in single COLO320DM cells. The central lines of boxplots are the median of data. The lower and upper hinges correspond to the 25th and 75th percentiles. The end of the lower and upper whiskers are 1.5 * IQR (inter-quartile range).
Supplementary information
Supplementary Information
Supplementary Figs. 1–7 and Text.
Supplementary Table 1
Combined supplementary tables.
Supplementary Data 1
Combined source data for supplementary figures.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, W., Lu, J., Lu, P. et al. scNanoHi-C: a single-cell long-read concatemer sequencing method to reveal high-order chromatin structures within individual cells. Nat Methods 20, 1493–1505 (2023). https://doi.org/10.1038/s41592-023-01978-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-023-01978-w
This article is cited by
-
From sequence to consequence: Deciphering the complex cis-regulatory landscape
Journal of Biosciences (2024)