Abstract
Antibiotic screens typically rely on growth inhibition to characterize compound bioactivity—an approach that cannot be used to assess the bactericidal activity of antibiotics against bacteria in drug-tolerant states. To address this limitation, we developed a multiplexed assay that uses metabolism-sensitive staining to report on the killing of antibiotic-tolerant bacteria. This method can be used with diverse bacterial species and applied to genome-scale investigations to identify therapeutic targets against tolerant pathogens.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Code availability
The MATLAB code for the image analysis is available at https://github.com/ajlopatkin/spock-analysis-pipeline.
Data availability
Large screening datasets are provided as Supplementary Information in Supplementary Table 1. Sequencing data are available at the NCBI Sequence Read Archive under accession no. PRJNA517046. The source data for Figs. 1 and 2, and Supplementary Figs. 1, 2, and 4–7 are available online (source data for Supplementary Fig. 3 are available in Supplementary Table 1). Additional data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Brown, E. D. & Wright, G. D. Nature 529, 336–343 (2016).
Wiegand, I., Hilpert, K. & Hancock, R. E. W. Nat. Protoc. 3, 163–175 (2008).
Certain, L. K., Way, J. C., Pezone, M. J. & Collins, J. J. Cell Host Microbe 22, 263–268 (2017).
Cornforth, D. M. et al. Proc. Natl Acad. Sci. USA 115, E5125–E5134 (2018).
Brauner, A., Fridman, O., Gefen, O. & Balaban, N. Q. Nat. Rev. Microbiol. 14, 320–330 (2016).
Meylan, S., Andrews, I. W. & Collins, J. J. Cell 172, 1228–1238 (2018).
Liu, A. et al. Antimicrob. Agents Chemother. 54, 1393–1403 (2010).
Baba, T. et al. Mol. Syst. Biol. 2, 2006.0008 (2006).
Gutierrez, A. et al. Mol. Cell 68, 1147–1154.e3 (2017).
Gutierrez, A., Stokes, J. M. & Matic, I. Antibiotics (Basel) 7, 32 (2018).
Sarkar, S. K., Dutta, M., Kumar, A., Mallik, D. & Ghosh, A. S. PLoS ONE 7, e48598 (2012).
Asadishad, B., Ghoshal, S. & Tufenkji, N. Environ. Sci. Technol. 45, 8345–8351 (2011).
Berridge, M. V., Herst, P. M. & Tan, A. S. Biotechnol. Annu. Rev. 11, 127–152 (2005).
Rohwer, F. & Azam, F. Appl. Environ. Microbiol. 66, 1001–1006 (2000).
Michel, B. PLoS Biol. 3, e255 (2005).
Hurwitz, S. J. & McCarthy, T. J. J. Pharm. Sci. 75, 912–916 (1986).
Tunney, M. M., Ramage, G., Field, T. R., Moriarty, T. F. & Storey, D. G. Antimicrob. Agents Chemother. 48, 1879–1881 (2004).
Fleck, L. E. et al. Antimicrob. Agents Chemother. 58, 1410–1419 (2014).
Shea, A., Wolcott, M., Daefler, S. & Rozak, D. A. Methods Mol. Biol. 881, 331–373 (2012).
Mackie, A. M., Hassan, K. A., Paulsen, I. T. & Tetu, S. G. Methods Mol. Biol. 1096, 123–130 (2014).
Ammor, M. S. J. Fluoresc. 17, 455–459 (2007).
Keseler, I. M. et al. Nucleic Acids Res. 41, D605–D612 (2013).
Karp, P. D. et al. Brief. Bioinform. 17, 877–890 (2016).
Karp, P. D. Science 293, 2040–2044 (2001).
Acknowledgements
We thank D. Hung from the Broad Institute for providing the bacterial pathogens used in this study. This work was supported by the Defense Threat Reduction Agency (grant no. HDTRA1-15-1-0051 to J.J.C.), the Broad Institute (grant no. 800170 to J.J.C.), the Bill and Melinda Gates Foundation (grant no. OPP1182991 to J.M.S. and J.J.C.), the Canadian Institutes of Health Research (grant no. FRN 143215 to E.D.B.), the Canadian Foundation for Innovation (grant no. 229283 to E.D.B.), the Canada Research Chairs program (Tier 1, to E.D.B.), the Fondation pour la Recherche Médicale (grant no. DBF20160635736 to A.G. and I.M.), the Banting Postdoctoral Fellowships Program (fellowship no. 393360 to J.M.S.), the National Science Foundation Graduate Research Fellowship Program (fellowship no. 1122374 to I.W.A.), and a generous gift from A. and J. Bekenstein.
Author information
Authors and Affiliations
Contributions
J.M.S. and A.G. conceptualized the study. J.M.S., A.G., A.J.L., I.W.A., and S.F. carried out the data acquisition and analysis. J.M.S. and A.G. wrote the manuscript. J.M.S., A.G., A.J.L., I.W.A., S.F., E.D.B., and J.J.C. edited the manuscript. J.M.S., A.G., I.M., E.D.B., and J.J.C. acquired the funding. I.M., E.D.B., and J.J.C. supervised the work.
Corresponding author
Ethics declarations
Competing interests
J.J.C. is scientific cofounder and SAB chair of EnBiotix, an antibiotic drug discovery company.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Killing of E. coli colonies on nitrocellulose membranes by ciprofloxacin.
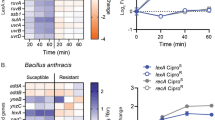
a, Killing of wild-type E. coli colonies by ciprofloxacin. Cells were seeded onto nitrocellulose membranes and grown for 24 h, then transferred to LB supplemented with ciprofloxacin for 24 h, and plated (mean ± range; n = 2 or n = 4 biologically independent experiments). b, Same as in a, except colony area was calculated (mean ± s.d.; n = 3 or n = 4 biologically independent experiments). Note that at concentrations as high as 5,000-fold MIC, only ~1 log of cell death occurred with insignificant changes in colony size. c, 24-point MIC curves of bactericidal antibiotics against wild-type E. coli. Overnight cultures were inoculated 1/10,000 into fresh LB media in final volumes of 100 µl in 96-well microtiter plates, and antibiotic was added to the concentrations indicated. Cultures were incubated until untreated control cultures reached stationary phase, and optical density at 600 nm was measured (mean ± range; n = 2 biologically independent experiments). d, 24-point MIC curves of bacteriostatic antibiotics against wild-type E. coli. Overnight cultures were inoculated 1/10,000 into fresh LB media in final volumes of 100 µl in 96-well microtiter plates, and antibiotic was added to the concentrations indicated. Cultures were incubated until untreated control cultures reached stationary phase, and optical density at 600 nm was measured (mean ± range; n = 2 biologically independent experiments). e, Killing of mature wild-type E. coli colonies by diverse antibiotics (top panel; mean ± range; n = 2 biologically independent experiments). Amp, ampicillin; cef, cefsulodin; mec, mecillinam; azt, aztreonam; cip, ciprofloxacin; gen, gentamicin; rif, rifampicin; cm, chloramphenicol; spec, spectinomycin. Numbers show fold-MIC concentrations used. Amp, cef, mec, azt, cip, and gen are regarded as bactericidal against E. coli; rif, cm, and spec are bacteriostatic. The bottom panel shows metabolic staining of wild-type E. coli colonies after treatment with the corresponding antibiotics on the top (mean ± range; n = 2 biologically independent experiments). f, Kinetics of TTC staining of E. coli. Wild-type cells were seeded onto nitrocellulose membranes and grown for 24 h on solid LB under aerobic conditions. Membranes were then transferred to LB under aerobic conditions, LB under anaerobic conditions, or LB supplemented with KCN at the indicated concentrations for 24 h. Membranes were then transferred to LB supplemented with 50 µg/ml TTC (aerobically for aerobic and KCN conditions; anaerobically for anaerobic conditions) and incubated for the noted durations prior to imaging (mean ± range; n = 2 biologically independent experiments). One hour of staining was selected as the experimental endpoint for the Keio screen to acquire a sufficiently high signal while avoiding overstaining. Fluorescence was normalized on the basis of maximum fluorescence (time = 0). g, Bacterial cell killing and metabolic staining of diverse species—E. coli, Klebsiella pneumoniae, Enterobacter cloacae, Acinetobacter baumannii, Pseudomonas aeruginosa, Staphylococcus aureus, Enterococcus faecalis, Bacillus simplex, and Mycobacterium smegmatis—subjected to heat treatment at 80 °C for 30 min. Blue bars show CFU quantification of colonies that were not heat treated (mean ± range; n = 2 biologically independent experiments). For CFU quantification of heat-treated samples, asterisks represent cell densities below the detection limit of 102 CFU/ml.
Supplementary Figure 2 Time dependence of colony killing by ciprofloxacin against 384 strains in the Keio collection.
Overnight cultures of 384 strains in the Keio collection (example plate shown in Fig. 2a) were seeded onto nitrocellulose membranes in 384-density in biologically independent duplicate experiments, and the membranes were placed onto LB media and incubated to stationary phase for 24 h. Nitrocellulose membranes containing mature colonies were then transferred to LB media supplemented with ciprofloxacin and incubated for the duration noted. Ciprofloxacin-treated colonies were then transferred to LB media supplemented with 50 µg/ml TTC for 1 h to stain live colonies. Fluorescence values of colonies treated with ciprofloxacin for varying durations were averaged and normalized to the non-drug-treated control (t = 0). Darker green represents greater fluorescence. These data show that increasing the duration of exposure to ciprofloxacin enhanced colony killing of sensitive strains. 24 h of ciprofloxacin exposure was selected to sufficiently increase the fluorescence of sensitive strains relative to that of non-sensitive strains.
Supplementary Figure 3 Screens of the E. coli Keio collection for killing and growth inhibition by ciprofloxacin.
a, Image and example analysis of the plate shown in Fig. 2a, depicting the contrast between highly fluorescent colonies (dead cells) and those that have been masked by the reduction of TTC to TPF (live cells). b, Replicate plot of normalized fluorescence values of the Keio collection in the absence of ciprofloxacin. In biologically independent duplicate experiments, colonies were seeded onto nitrocellulose membranes in 384-density and grown for 24 h. Membranes were transferred to LB plates without ciprofloxacin and incubated for an additional 24 h. Membranes were then transferred to LB supplemented with 50 µg/ml TTC for 1 h prior to imaging. Fluorescence values were normalized according to row, column, and plate as described in the Methods section. c, Replicate plot of normalized fluorescence values of the Keio collection treated with 1 µg/ml ciprofloxacin. In biologically independent duplicate experiments, colonies were seeded onto nitrocellulose membranes in 384-density and grown for 24 h. Membranes were transferred to LB plates containing 1 µg/ml ciprofloxacin and incubated for an additional 24 h. Membranes were then transferred to LB supplemented with 50 µg/ml TTC for 1 h prior to imaging. Fluorescence values were normalized according to row, column, and plate as described in the Methods section. d, Normalized data of the screen of the Keio collection for colony fluorescence using the SPOCK assay. The yellow region shows hit strains based on a 3σ hit cutoff. With this criterion, 38 strains were identified as being sensitive to killing by ciprofloxacin under resource-depleted, tolerant conditions. e, Replicate plot of normalized optical density values of the Keio collection in the absence of ciprofloxacin. In biologically independent duplicate experiments, strains were grown in liquid LB in 384-density for 18 h and read at 600 nm. Data were normalized according to row, column, and plate as described in the Methods section. f, Replicate plot of normalized optical density values of the Keio collection in the presence of ciprofloxacin. In biologically independent duplicate experiments, strains were grown in liquid LB supplemented with 1/8 × MIC ciprofloxacin in 384-density for 18 h and read at 600 nm. Data were normalized according to row, column, and plate as described in the Methods section. g, Normalized data of the screen of the Keio collection for growth inhibition in liquid culture. The blue region shows strains that displayed enhanced growth inhibition in the presence of ciprofloxacin based on a 1.5σ hit cutoff. 27 strains were identified as being ciprofloxacin sensitive.
Supplementary Figure 4 Characterization of strains that display sensitivity to killing by ciprofloxacin.
a, MIC values of 38 strains that displayed as hits in the SPOCK screen of the Keio collection. 12-point concentration curves were generated, and MIC values were plotted (mean ± range; n = 2 biologically independent experiments). Note that no strains that displayed high fluorescence showed significant shifts in ciprofloxacin MIC relative to wild-type E. coli. b, Ciprofloxacin-induced killing in liquid LB of the 38 strains that displayed as hits in the SPOCK screen of the Keio collection. Strains were grown to saturation for 24 h in final volumes of 100 µl in 96-well microtiter plates, at which time ciprofloxacin was added at a concentration of 1 µg/ml. Cultures were incubated for an additional 24 h prior to plating. Survival is calculated as the viability of each strain after ciprofloxacin treatment relative to untreated control cultures of each strain, which is depicted as the red dotted line (mean ± range; n = 2 biologically independent experiments). c, Culture density of each Keio strain after 24 h of incubation, prior to the addition of ciprofloxacin. The red dotted line shows the absolute density of wild-type E. coli (mean ± range; n = 2 biologically independent experiments). d, Stationary phase sensitivity of the 38 SPOCK assay hits to ciprofloxacin in liquid media, relative to colony fluorescence values. Strains were grown for 24 h in liquid LB, at which time ciprofloxacin (1 µg/ml) was added and cells were grown for an additional 24 h prior to plating (mean; n = 2 biologically independent experiments). Light blue circles show strains that were used in follow-up experiments. The red dotted line depicts wild-type E. coli survival. e, Growth kinetics of 38 SPOCK assay hit strains. Overnight cultures were inoculated 1/10,000 into fresh LB media in final volumes of 100 µl in 96-well microtiter plates. Optical density at 600 nm was read every 10 min over 24 h, and the mean of every growth curve was plotted (mean; n = 2 biologically independent experiments). Wild-type E. coli is shown in blue. The five strains that were selected for follow-up investigations are shown in light blue.
Supplementary Figure 5 Dose-response analyses of ciprofloxacin against select strains that display sensitivity to killing by ciprofloxacin.
a–e, 24-point MIC curves of the five SPOCK assay hit strains that were used for follow-up investigation. Overnight cultures were inoculated 1/10,000 into fresh LB media in final volumes of 100 µl in 96-well microtiter plates, and ciprofloxacin was added to the concentrations indicated. Cultures were incubated until untreated control cultures reached stationary phase, and optical density at 600 nm was measured (mean ± range; n = 2 biologically independent experiments). In all panels, wild-type E. coli is shown in blue and mutants are shown in red. Note that no shifts in MIC are observed. f–j, 12-point kill curves of the five strains described above. Cultures were grown to exponential phase (squares) or stationary phase (circles) in LB media in final volumes of 100 µl in 96-well microtiter plates. Ciprofloxacin was introduced at the concentrations noted, and cells were incubated for an additional 24 h prior to plating (mean ± s.d.; n = 2 (exponential phase) or n = 3 (stationary phase) biologically independent experiments). In all panels, wild-type E. coli is shown in blue and mutants are shown in red. Note that little difference in killing is observed between wild-type E. coli and each mutant during exponential phase treatment, whereas significant divergence occurs at stationary phase. LdcA (EMBO J. 18, 4108–4117; 1999), NlpI (Proc. Natl. Acad. Sci. USA 112, 10956–10961; 2015), and YibP (EnvC) (Mol. Microbiol. 52, 1255–1269; 2004) modulate peptidoglycan cross-linking and recycling, whereas OmpR (J. Biol. Chem. 277, 24155–24161; 2002) and YfgL (BamB) (Cell 121, 235–245; 2005) play roles in governing membrane protein composition.
Supplementary Figure 6 Dose-response analyses of functionally diverse antibiotics against select strains that display sensitivity to killing by ciprofloxacin.
a–c, 24-point MIC curves of wild-type E. coli and the five SPOCK assay hit strains that were used for follow-up investigation. Overnight cultures were inoculated 1/10,000 into fresh LB media in final volumes of 100 µl in 96-well microtiter plates, and antibiotic was added to the concentrations indicated. Cultures were incubated until untreated control cultures reached stationary phase, and optical density at 600 nm was measured (mean ± range; n = 2 biologically independent experiments). d–f, 12-point kill curves of wild-type E. coli and the five strains described above. Cultures were grown to stationary phase in LB media in final volumes of 100 µl in 96-well microtiter plates. Antibiotic was introduced at the concentrations noted, and cells were incubated for an additional 24 h prior to plating (mean ± range; n = 2 biologically independent experiments). We note here that perhaps because of its lack of cell envelope integrity in stationary phase, the ΔldcA strain displays promiscuous sensitivity to killing by ampicillin and gentamicin, in addition to ciprofloxacin, whereas ΔnlpI, ΔompR, ΔyfgL, and ΔyibP display stationary phase sensitivity to ciprofloxacin exclusively, relative to the wild type. LdcA (EMBO J. 18, 4108–4117; 1999), NlpI (Proc. Natl. Acad. Sci. USA 112, 10956–10961; 2015), and YibP (EnvC) (Mol. Microbiol. 52, 1255–1269; 2004) modulate peptidoglycan cross-linking and recycling, whereas OmpR (J. Biol. Chem. 277, 24155–24161; 2002) and YfgL (BamB) (Cell 121, 235–245; 2005) play roles in governing membrane protein composition.
Supplementary Figure 7 Phenotypic analysis of ciprofloxacin-sensitive gene deletion mutants.
a, Example TUNEL staining as observed by flow cytometry of wild-type E. coli treated with hydrogen peroxide during exponential phase (left), ciprofloxacin during exponential phase (middle), or ciprofloxacin during stationary phase (right). This is a representative experiment of three biologically independent experiments. In all panels, gray regions show untreated control samples, and blue regions show treated samples. Note the difference in TUNEL stained populations of E. coli with ciprofloxacin between exponential phase and stationary phase. b, DNA double-strand breaks induced by ciprofloxacin during exponential growth. Cells were grown to exponential phase in LB and treated with 100 ng/ml ciprofloxacin (–C) or vehicle for 1 h prior to TUNEL analysis by flow cytometry (mean ± s.d.; n = 3 biologically independent experiments, except for YibP, where n = 2 biologically independent experiments). Hydrogen peroxide (100 mM for 10 min) was used as a positive control. c–h, Micrographs of E. coli strains treated with ciprofloxacin during exponential phase and stationary phase. These are representative images of two biologically independent experiments. For exponential phase images, cells were grown to OD ~0.1 and treated with ciprofloxacin (100 ng/ml) for 24 h. For stationary phase images, cells were grown to saturation for 24 h and treated with ciprofloxacin (1 µg/ml) for 24 h. Cells were then fixed, stained with FM4–64, and imaged. Note the lack of filamentation by ciprofloxacin against resource-depleted stationary phase cells, despite their sensitivity to killing. i, 24-point MIC curves of wild-type E. coli and E. coli harboring a gyrA S83A ciprofloxacin-resistance mutation. Overnight cultures were inoculated 1/10,000 into fresh LB media in final volumes of 100 µl in 96-well microtiter plates, and ciprofloxacin was added to the concentrations indicated. Cultures were incubated until untreated control cultures reached stationary phase, and optical density at 600 nm was measured (mean ± range; n = 2 biologically independent experiments). j, 24-point MIC curves of ΔyfgL E. coli and ΔyfgL E. coli harboring a gyrA D87Y ciprofloxacin-resistance mutation. Overnight cultures were inoculated 1/10,000 into fresh LB media in final volumes of 100 µl in 96-well microtiter plates, and ciprofloxacin was added to the concentrations indicated. Cultures were incubated until untreated control cultures reached stationary phase, and optical density at 600 nm was measured (mean ± range; n = 2 biologically independent experiments). Note that gyrA S83A and gyrA D87Y confer the same level of resistance. k, Sensitivity of the four strains shown in i and j to killing by ciprofloxacin in exponential phase. Cells were grown to ~107 CFU/ml and treated with ciprofloxacin for 24 h prior to plating (mean ± s.d.; n = 3 biologically independent experiments). l, Growth kinetics of the four strains in i and j. Overnight cultures were inoculated 1/10,000 into fresh LB media in final volumes of 100 µl in 96-well microtiter plates. Optical density at 600 nm was read every 10 min over 24 h, and curves were plotted (mean ± s.d.; n = 3 biologically independent experiments). m, 12-point kill curves of the four strains in i and j. Cultures were grown to stationary phase in LB media in final volumes of 100 µl in 96-well microtiter plates. Ampicillin was introduced at the concentrations noted, and cells were incubated for an additional 24 h prior to plating (mean ± range; n = 2 biologically independent experiments). Wild-type E. coli is shown in blue, and ΔyfgL E. coli is shown in red. Circles represent wild-type gyrA strains, and squares represent ciprofloxacin-resistant gyrA strains. Ampicillin requires high metabolic activity for killing, and thus these data support the notion that all strains were in stationary phase prior to the introduction of ciprofloxacin.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7
Supplementary Table 1
Screening data for growth inhibition and colony fluorescence of the E. coli Keio collection treated with ciprofloxacin.
Supplementary Table 2
Gene deletion mutant strains of E. coli with enhanced colony fluorescence after treatment with ciprofloxacin.
Supplementary Table 3
Gene ontology (GO), biosynthetic pathway, and promoter activation enrichment by the screen for enhanced fluorescence of the E. coli Keio collection after treatment with ciprofloxacin.
Supplementary Table 4
Description: Gene deletion mutant strains of E. coli with enhanced growth inhibition after treatment with ciprofloxacin.
Supplementary Table 5
Gene ontology (GO), biosynthetic pathway, and promoter activation enrichment by the screen for enhanced growth inhibition of the E. coli Keio collection after treatment with ciprofloxacin.
Rights and permissions
About this article
Cite this article
Stokes, J.M., Gutierrez, A., Lopatkin, A.J. et al. A multiplexable assay for screening antibiotic lethality against drug-tolerant bacteria. Nat Methods 16, 303–306 (2019). https://doi.org/10.1038/s41592-019-0333-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-019-0333-y
This article is cited by
-
A high-throughput and low-waste viability assay for microbes
Nature Microbiology (2023)
-
Combining CRISPRi and metabolomics for functional annotation of compound libraries
Nature Chemical Biology (2022)
-
Expanding the search for small-molecule antibacterials by multidimensional profiling
Nature Chemical Biology (2022)
-
Pareto optimality between growth-rate and lag-time couples metabolic noise to phenotypic heterogeneity in Escherichia coli
Nature Communications (2021)