Abstract
Genome-wide association studies (GWASs) have provided numerous associations between human single-nucleotide polymorphisms (SNPs) and health traits. Likewise, metagenome-wide association studies (MWASs) between bacterial SNPs and human traits can suggest mechanistic links, but very few such studies have been done thus far. In this study, we devised an MWAS framework to detect SNPs and associate them with host phenotypes systematically. We recruited and obtained gut metagenomic samples from a cohort of 7,190 healthy individuals and discovered 1,358 statistically significant associations between a bacterial SNP and host body mass index (BMI), from which we distilled 40 independent associations. Most of these associations were unexplained by diet, medications or physical exercise, and 17 replicated in a geographically independent cohort. We uncovered BMI-associated SNPs in 27 bacterial species, and 12 of them showed no association by standard relative abundance analysis. We revealed a BMI association of an SNP in a potentially inflammatory pathway of Bilophila wadsworthia as well as of a group of SNPs in a region coding for energy metabolism functions in a Faecalibacterium prausnitzii genome. Our results demonstrate the importance of considering nucleotide-level diversity in microbiome studies and pave the way toward improved understanding of interpersonal microbiome differences and their potential health implications.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Data in this paper are part of the Human Phenotype Project. The raw metagenomic data and basic phenotypes (age, sex and BMI) used in this study are available at the European Genome-phenome Archive (https://ega-archive.org/) under accession EGAS00001007204. The other data are accessible to researchers from universities and other research institutions at https://humanphenotypeproject.org/home.
Code availability
Analysis source code is available at https://github.com/LironZa/MWAS.
References
Lynch, S. V. & Pedersen, O. The human intestinal microbiome in health and disease. N. Engl. J. Med. 375, 2369–2379 (2016).
Manichanh, C. et al. Reduced diversity of faecal microbiota in Crohn’s disease revealed by a metagenomic approach. Gut 55, 205–211 (2006).
Tang, W. H. W. et al. Intestinal microbial metabolism of phosphatidylcholine and cardiovascular risk. N. Engl. J. Med. 368, 1575–1584 (2013).
Turnbaugh, P. J. et al. A core gut microbiome in obese and lean twins. Nature 457, 480–484 (2009).
Karlsson, F. H. et al. Gut metagenome in European women with normal, impaired and diabetic glucose control. Nature 498, 99–103 (2013).
Yoshimoto, S. et al. Obesity-induced gut microbial metabolite promotes liver cancer through senescence secretome. Nature 499, 97–101 (2013).
Gopalakrishnan, V., Helmink, B. A., Spencer, C. N., Reuben, A. & Wargo, J. A. The influence of the gut microbiome on cancer, immunity, and cancer immunotherapy. Cancer Cell 33, 570–580 (2018).
Maruvada, P., Leone, V., Kaplan, L. M. & Chang, E. B. The human microbiome and obesity: moving beyond associations. Cell Host Microbe 22, 589–599 (2017).
De Filippis, F. et al. Distinct genetic and functional traits of human intestinal Prevotella copri strains are associated with different habitual diets. Cell Host Microbe 25, 444–453 (2019).
Brito, I. L. et al. Mobile genes in the human microbiome are structured from global to individual scales. Nature 535, 435–439 (2016).
Greenblum, S., Carr, R. & Borenstein, E. Extensive strain-level copy-number variation across human gut microbiome species. Cell 160, 583–594 (2015).
Zeevi, D. et al. Structural variation in the gut microbiome associates with host health. Nature 568, 43–48 (2019).
Yoshida, H., Bogaki, M., Nakamura, M. & Nakamura, S. Quinolone resistance-determining region in the DNA gyrase gyrA gene of Escherichia coli. Antimicrob. Agents Chemother. 34, 1271–1272 (1990).
Viana, D. et al. A single natural nucleotide mutation alters bacterial pathogen host tropism. Nat. Genet. 47, 361–366 (2015).
Read, T. D. & Massey, R. C. Characterizing the genetic basis of bacterial phenotypes using genome-wide association studies: a new direction for bacteriology. Genome Med. 6, 109 (2014).
Schloissnig, S. et al. Genomic variation landscape of the human gut microbiome. Nature 493, 45–50 (2013).
Shi, Z. J. et al. Fast and accurate metagenotyping of the human gut microbiome with GT-Pro. Nat. Biotechnol. 40, 507–516 (2022).
Olm, M. R. et al. inStrain profiles population microdiversity from metagenomic data and sensitively detects shared microbial strains. Nat. Biotechnol. 39, 727–736 (2021).
Maini Rekdal, V., Bess, E. N., Bisanz, J. E., Turnbaugh, P. J. & Balskus, E. P. Discovery and inhibition of an interspecies gut bacterial pathway for Levodopa metabolism. Science 364, eaau6323 (2019).
Wang, J. & Jia, H. Metagenome-wide association studies: fine-mining the microbiome. Nat. Rev. Microbiol. 14, 508–522 (2016).
Shilo, S. et al. 10 K: a large-scale prospective longitudinal study in Israel. Eur. J. Epidemiol. 36, 1187–1194 (2021).
Leviatan, S., Shoer, S., Rothschild, D., Gorodetski, M. & Segal, E. An expanded reference map of the human gut microbiome reveals hundreds of previously unknown species. Nat. Commun. 13, 3863 (2022).
Rothschild, D. et al. An atlas of robust microbiome associations with phenotypic traits based on large-scale cohorts from two continents. PLoS ONE 17, e0265756 (2022).
Truong, D. T., Tett, A., Pasolli, E., Huttenhower, C. & Segata, N. Microbial strain-level population structure and genetic diversity from metagenomes. Genome Res. 27, 626–638 (2017).
Power, R. A., Parkhill, J. & de Oliveira, T. Microbial genome-wide association studies: lessons from human GWAS. Nat. Rev. Genet. 18, 41–50 (2017).
Earle, S. G. et al. Identifying lineage effects when controlling for population structure improves power in bacterial association studies. Nat. Microbiol. 1, 16041 (2016).
Chen, P. E. & Shapiro, B. J. The advent of genome-wide association studies for bacteria. Curr. Opin. Microbiol. 25, 17–24 (2015).
Gacesa, R. et al. Environmental factors shaping the gut microbiome in a Dutch population. Nature 604, 732–739 (2022).
Leylabadlo, H. E. et al. The critical role of Faecalibacterium prausnitzii in human health: an overview. Microb. Pathog. 149, 104344 (2020).
Xu, Q. et al. Mendelian randomization analysis reveals causal effects of the human gut microbiota on abdominal obesity. J. Nutr. 151, 1401–1406 (2021).
Turnbaugh, P. J. et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444, 1027–1031 (2006).
Das, B., Das, M., Kalita, A. & Baro, M. R. The role of Wnt pathway in obesity induced inflammation and diabetes: a review. J. Diabetes Metab. Disord. 20, 1871–1882 (2021).
Cani, P. D. et al. Metabolic endotoxemia initiates obesity and insulin resistance. Diabetes 56, 1761–1772 (2007).
Natividad, J. M. et al. Bilophila wadsworthia aggravates high fat diet induced metabolic dysfunctions in mice. Nat. Commun. 9, 2802 (2018).
Needham, B. D. & Trent, M. S. Fortifying the barrier: the impact of lipid A remodelling on bacterial pathogenesis. Nat. Rev. Microbiol. 11, 467–481 (2013).
Zhao, S. et al. Adaptive evolution within gut microbiomes of healthy people. Cell Host Microbe 25, 656–667 (2019).
Boehme, M. et al. Microbiota from young mice counteracts selective age-associated behavioral deficits. Nat. Aging 1, 666–676 (2021).
Garud, N. R. & Pollard, K. S. Population genetics in the human microbiome. Trends Genet. 36, 53–67 (2020).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Zeevi, D. et al. Personalized nutrition by prediction of glycemic responses. Cell 163, 1079–1094 (2015).
Seabold, S. & Perktold, J. Statsmodels: econometric and statistical modeling with Python. In Proc. of the 9th Python in Science Conference 92–96 (SciPy, 2010).
Virtanen, P. et al. SciPy 1.0: fundamental algorithms for scientific computing in Python. Nat. Methods 17, 261–272 (2020).
Hunter, J. D. Matplotlib: a 2D graphics environment. Comput. Sci. Eng. 9, 90–95 (2007).
Flyamer, I. et al. Phlya/adjustText: 0.8 beta. Zenodo https://doi.org/10.5281/zenodo.3924114 (2020).
Acknowledgements
We thank members of the Segal laboratory for useful discussions. E.S. is supported by the Crown Human Genome Center; the Larson Charitable Foundation New Scientist Fund; the Else Kröner Fresenius Foundation; the White Rose International Foundation; the Ben B. and Joyce E. Eisenberg Foundation; the Nissenbaum Family; Marcos Pinheiro de Andrade and Vanessa Buchheim; Lady Michelle Michels; Aliza Moussaieff; and grants funded by the Minerva Foundation, with funding from the Federal German Ministry for Education and Research and by the European Research Council and the Israel Science Foundation. S.S. is supported by the Israeli Council for Higher Education via the Weizmann Data Science Research Center. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
L.Z. conceived and designed the study, designed and conducted the analyses, interpreted the results and wrote the manuscript. A.L. developed methods. L.R. interpreted the results and wrote the manuscript. S.S., A.G. and S.L. designed and conducted sample processing. M.R. processed the dietary data. O.W. developed statistical analyses. A.W. designed the project, developed protocols and oversaw sample collection and processing. E.S. conceived, directed and designed the project and analyses, conducted analyses, interpreted the results and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
O.W. is an employee of Eleven Tx. E.S. is a paid consultant for Pheno.AI. The other authors declare no competing interests.
Peer review
Peer review information
Nature Medicine thanks Sergio Baranzini, Thomas Schmidt and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Alison Farrell, in collaboration with the Nature Medicine team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
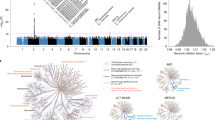
Extended Data Fig. 1 SNPs overview.
(a) Distribution of the 12,686,191 detected SNPs across 348 species. (b) Number of samples covering different SNPs.
Extended Data Fig. 2 Volcano plot.
Volcano plot shows for each SNP the difference between the average BMI in individuals with mostly the alternative allele (major allele frequency ≤ 0.5) and the average BMI in individuals with mostly the major allele (major allele frequency > 0.5; x-axis); and its p-value (y-axis). Red annotations show gene symbols of the protein-coding SNPs left after the clumping stage (if a gene symbol exists). X-axis was truncated to the range of statistically significant associations ±10%.
Extended Data Fig. 3 BMI differences.
For each of the 40 BMI-associated SNPs that remained after the clumping stage, boxplots (center, median; box, interquartile range; whiskers, 5th and 95th percentiles; notches, 95% confidence interval around the median based on 1,000 times bootstrap) compare host BMI distribution of individuals with no bacteria of this species (left box; Methods), hosts of bacteria with the major allele (middle box; major allele frequency ≥ 0.99) and hosts of bacteria with the minor allele (right box; major allele frequency ≤ 0.01). The grey scale indicates the difference between medians. Groups were compared in a two-sided Mann-Whitney test, and p-values were Bonferroni corrected for 120 hypotheses (40 SNPs, 3 comparisons per SNP).
Extended Data Fig. 4 Quantile-quantile (Q-Q) plots.
Expected (uniform distribution between 1/[the total number of tested SNPs] and 1) p-values compared to the SNPs p-values estimated in the MWAS analysis. (a) All tested SNPs. Red dots are the 40 BMI-associated SNPs remaining after the clumping procedure. (b) Each species estimated and plotted separately using a random color. Straight lines connect adjacent SNP dots to increase readability. (c) Species with more than 13 BMI-associated SNPs. Straight lines connect adjacent SNP dots to increase readability.
Extended Data Fig. 5 Number of correlated SNPs in each linkage group.
Histograms show the number of correlated SNPs that were found in the clumping stage in each linkage group. The total number of groups is 40, which is the final number of SNPs that remained post the clumping procedure. (a) Full range of group sizes. (b) Groups with 1 to 100 SNPs.
Extended Data Fig. 6 Power analysis.
Boxplots (center, median; box, interquartile range; whiskers, 1.5 * interquartile range or the most extreme data point) show the calculated power for associating the 40 SNPs with BMI, given the effect size observed in our cohort and various effective sample sizes (N). Alpha was set to 3.9 × 10−9 based on a cutoff of 0.05 and a Bonferroni correction for 12,686,191 hypotheses.
Extended Data Fig. 7 Random replication control.
For 1000 random choices of 40 SNPs from the discovery analysis, showing how many passed the 0.05 Bonferroni adjusted cutoff for association with BMI in the replication cohort. For reference, the red dotted line shows the number of SNPs that passed the cutoff when the 40 SNPs that were associated with BMI in the discovery cohort were tested — 17.
Extended Data Fig. 8 Replication cohort characteristics.
Age, sex, and BMI distribution of the 8,204 study participants.
Supplementary information
Supplementary Tables 1–11
Supplementary Table 1. Number of tested, BMI-associated and post-clumping SNPs per species, in species with BMI-associated SNPs. Table 2. BMI-associated SNPs, post-clumping and their genetic context. SNPs that are within genes. Table 3. BMI-associated SNPs, post-clumping and their genetic context. Intergenic SNPs. Table 4. Associations between bacterial relative abundance and host BMI. Table 5. Replication MWAS in the Dutch cohort. Table 6. Associations between bacterial relative abundance and host BMI in the Dutch cohort. Table 7. BMI-associated SNP (Rep_3066, C_257, pos 6315) and the BMI-associated SNPs that are correlated with it and were excluded in the clumping process. Table 8. Diet, exercise and medication covariates divided by category. Table 9. MWAS analysis with diet covariates. Table 10. MWAS analysis with exercise covariates. Table 11. MWAS analysis with medication covariates.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zahavi, L., Lavon, A., Reicher, L. et al. Bacterial SNPs in the human gut microbiome associate with host BMI. Nat Med 29, 2785–2792 (2023). https://doi.org/10.1038/s41591-023-02599-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-023-02599-8