Abstract
Complex diseases, such as coronary artery disease (CAD), are often multifactorial, caused by multiple underlying pathological mechanisms. Here, to study the multifactorial nature of CAD, we performed comprehensive clinical and multi-omic profiling, including serum metabolomics and gut microbiome data, for 199 patients with acute coronary syndrome (ACS) recruited from two major Israeli hospitals, and validated these results in a geographically distinct cohort. ACS patients had distinct serum metabolome and gut microbial signatures as compared with control individuals, and were depleted in a previously unknown bacterial species of the Clostridiaceae family. This bacterial species was associated with levels of multiple circulating metabolites in control individuals, several of which have previously been linked to an increased risk of CAD. Metabolic deviations in ACS patients were found to be person specific with respect to their potential genetic or environmental origin, and to correlate with clinical parameters and cardiovascular outcomes. Moreover, metabolic aberrations in ACS patients linked to microbiome and diet were also observed to a lesser extent in control individuals with metabolic impairment, suggesting the involvement of these aberrations in earlier dysmetabolic phases preceding clinically overt CAD. Finally, a metabolomics-based model of body mass index (BMI) trained on the non-ACS cohort predicted higher-than-actual BMI when applied to ACS patients, and the excess BMI predictions independently correlated with both diabetes mellitus (DM) and CAD severity, as defined by the number of vessels involved. These results highlight the utility of the serum metabolome in understanding the basis of risk-factor heterogeneity in CAD.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The raw metagenomic sequencing data per sample of the controls are available from the European Nucleotide Archive (ENA; https://www.ebi.ac.uk/ena): PRJEB11532. The raw metabolomics data and phenotypes per sample of the controls are available from the European Genome-phenome Archive (EGA; https://ega-archive.org/): EGAS00001004512. The raw metabolomics data and full clinical phenotypes for the cohort of individuals with ACS are available from the EGA: EGAS00001005342. Additional data regarding SGB 4712, including the genome sequence, gene annotation and closest references are available at https://github.com/noambar/ACStudy/tree/master/SGB_4712.
Code availability
Analysis source code is available at https://github.com/noambar/ACStudy.
References
Roth, G. A. et al. Global and regional patterns in cardiovascular mortality from 1990 to 2013. Circulation 132, 1667–1678 (2015).
Wilkins, J. T. et al. Lifetime risk and years lived free of total cardiovascular disease. JAMA 308, 1795–1801 (2012).
Poulter, N. Coronary heart disease is a multifactorial disease. Am. J. Hypertens. 12, 92S–95S (1999).
Gaziano, J. M. et al. Use of aspirin to reduce risk of initial vascular events in patients at moderate risk of cardiovascular disease (ARRIVE): a randomised, double-blind, placebo-controlled trial. Lancet 392, 1036–1046 (2018).
Psychogios, N. et al. The human serum metabolome. PLoS ONE 6, e16957 (2011).
Tang, W. H. W. et al. Intestinal microbial metabolism of phosphatidylcholine and cardiovascular risk. N. Engl. J. Med. 368, 1575–1584 (2013).
Brown, J. M. & Hazen, S. L. Metaorganismal nutrient metabolism as a basis of cardiovascular disease. Curr. Opin. Lipidol. 25, 48–53 (2014).
Zhu, W. et al. Gut microbial metabolite TMAO enhances platelet hyperreactivity and thrombosis risk. Cell 165, 111–124 (2016).
Barreto, F. C. et al. Serum indoxyl sulfate is associated with vascular disease and mortality in chronic kidney disease patients. Clin. J. Am. Soc. Nephrol. 4, 1551–1558 (2009).
Meijers, B. K. I. et al. p-Cresol and cardiovascular risk in mild-to-moderate kidney disease. Clin. J. Am. Soc. Nephrol. 5, 1182–1189 (2010).
Zeevi, D. et al. Personalized nutrition by prediction of glycemic responses. Cell 163, 1079–1094 (2015).
Rothschild, D. et al. Environment dominates over host genetics in shaping human gut microbiota. Nature 555, 210–215 (2018).
Jie, Z. et al. The gut microbiome in atherosclerotic cardiovascular disease. Nat. Commun. 8, 845 (2017).
Tang, W. H. W., Kitai, T. & Hazen, S. L. Gut microbiota in cardiovascular health and disease. Circ. Res. 120, 1183–1196 (2017).
Maier, L. et al. Extensive impact of non-antibiotic drugs on human gut bacteria. Nature 555, 623–628 (2018).
Liu, J. et al. Integration of epidemiologic, pharmacologic, genetic and gut microbiome data in a drug-metabolite atlas. Nat. Med. 26, 110–117 (2020).
Forslund, S. K. et al. Combinatorial, additive and dose-dependent drug-microbiome associations. Nature https://doi.org/10.1038/s41586-021-04177-9 (2021).
Kannel, W. B. & McGee, D. L. Diabetes and cardiovascular disease. The Framingham study. JAMA 241, 2035–2038 (1979).
Bar, N. et al. A reference map of potential determinants for the human serum metabolome. Nature https://doi.org/10.1038/s41586-020-2896-2 (2020).
Winter, S. E. & Bäumler, A. J. Dysbiosis in the inflamed intestine: chance favors the prepared microbe. Gut Microbes 5, 71–73 (2014).
Carnevale, R. et al. Low-grade endotoxaemia enhances artery thrombus growth via Toll-like receptor 4: implication for myocardial infarction. Eur. Heart J. 41, 3156–3165 (2020).
Meijers, B. K. I. et al. Free p-cresol is associated with cardiovascular disease in hemodialysis patients. Kidney Int. 73, 1174–1180 (2008).
Poesen, R. et al. Microbiota-derived phenylacetylglutamine associates with overall mortality and cardiovascular disease in patients with CKD. J. Am. Soc. Nephrol. 27, 3479–3487 (2016).
Nemet, I. et al. A cardiovascular disease-linked gut microbial metabolite acts via adrenergic receptors. Cell 180, 862–877.e22 (2020).
Cheah, I. K. & Halliwell, B. Ergothioneine; antioxidant potential, physiological function and role in disease. Biochim. Biophys. Acta. 1822, 784–793 (2012).
Smith, E. et al. Ergothioneine is associated with reduced mortality and decreased risk of cardiovascular disease. Heart 106, 691–697 (2020).
Leopold, J. A. & Loscalzo, J. Emerging role of precision medicine in cardiovascular disease. Circ. Res. 122, 1302–1315 (2018).
Dhingra, R. & Vasan, R. S. Age as a risk factor. Med. Clin. North Am. 96, 87–91 (2012).
Lusis, A. J. Atherosclerosis. Nature 407, 233–241 (2000).
Weber, C. & Noels, H. Atherosclerosis: current pathogenesis and therapeutic options. Nat. Med. 17, 1410–1422 (2011).
Poirier, P. et al. Obesity and cardiovascular disease: pathophysiology, evaluation, and effect of weight loss: an update of the 1997 American Heart Association Scientific Statement on Obesity and Heart Disease from the Obesity Committee of the Council on Nutrition, Physical Activity, and Metabolism. Circulation 113, 898–918 (2006).
Goossens, G. H. The metabolic phenotype in obesity: fat mass, body fat distribution, and adipose tissue function. Obes. Facts 10, 207–215 (2017).
Cirulli, E. T. et al. Profound perturbation of the metabolome in obesity is associated with health risk. Cell Metab. 29, 488–500.e2 (2019).
Ke, G. et al. LightGBM: a highly efficient gradient boosting decision tree. NeurIPS Proceedings https://papers.nips.cc/paper/6907-lightgbm-a-highly-efficient-gradient-boosting-decision-tree.pdf (2017).
Ringqvist, I. et al. Prognostic value of angiographic indices of coronary artery disease from the Coronary Artery Surgery Study (CASS). J. Clin. Invest. 71, 1854–1866 (1983).
Lundberg, S. M., Erion, G. G. & Lee, S.-I. Consistent individualized feature attribution for tree ensembles. Preprint at arXiv (2018).
Pickens, C. A., Vazquez, A. I., Jones, A. D. & Fenton, J. I. Obesity, adipokines, and C-peptide are associated with distinct plasma phospholipid profiles in adult males, an untargeted lipidomic approach. Sci. Rep. 7, 6335 (2017).
Vangipurapu, J., Fernandes Silva, L., Kuulasmaa, T., Smith, U. & Laakso, M. Microbiota-related metabolites and the risk of type 2 diabetes. Diabetes Care 43, 1319–1325 (2020).
Zong, G. et al. Associations between linoleic acid intake and incident type 2 diabetes among U.S. men and women. Diabetes Care 42, 1406–1413 (2019).
Pertiwi, K. et al. Plasma and dietary linoleic acid and 3-year risk of type 2 diabetes after myocardial infarction: a prospective analysis in the alpha omega cohort. Diabetes Care 43, 358–365 (2020).
Martínez, I., Muller, C. E. & Walter, J. Long-term temporal analysis of the human fecal microbiota revealed a stable core of dominant bacterial species. PLoS ONE 8, e69621 (2013).
Lozupone, C. A., Stombaugh, J. I., Gordon, J. I., Jansson, J. K. & Knight, R. Diversity, stability and resilience of the human gut microbiota. Nature 489, 220–230 (2012).
Yousri, N. A. et al. Long term conservation of human metabolic phenotypes and link to heritability. Metabolomics 10, 1005–1017 (2014).
Soneson, C., Gerster, S. & Delorenzi, M. Batch effect confounding leads to strong bias in performance estimates obtained by cross-validation. PLoS ONE 9, e100335 (2014).
Friedewald, W. T., Levy, R. I. & Fredrickson, D. S. Estimation of the concentration of low-density lipoprotein cholesterol in plasma, without use of the preparative ultracentrifuge. Clin. Chem. 18, 499–502 (1972).
Soininen, P., Kangas, A. J., Würtz, P., Suna, T. & Ala-Korpela, M. Quantitative serum nuclear magnetic resonance metabolomics in cardiovascular epidemiology and genetics. Circ. Cardiovasc. Genet. 8, 192–206 (2015).
Würtz, P. et al. Quantitative serum nuclear magnetic resonance metabolomics in large-scale epidemiology: a primer on -omic technologies. Am. J. Epidemiol. 186, 1084–1096 (2017).
Marco-Sola, S., Sammeth, M., Guigó, R. & Ribeca, P. The GEM mapper: fast, accurate and versatile alignment by filtration. Nat. Methods 9, 1185–1188 (2012).
Pasolli, E. et al. Extensive unexplored human microbiome diversity revealed by over 150,000 genomes from metagenomes spanning age, geography, and lifestyle. Cell 176, 649–662.e20 (2019).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Korem, T. et al. Growth dynamics of gut microbiota in health and disease inferred from single metagenomic samples. Science 349, 1101–1106 (2015).
Zeevi, D. et al. Structural variation in the gut microbiome associates with host health. Nature 568, 43–48 (2019).
Lundberg, S. & Lee, S.-I. A unified approach to interpreting model predictions. Preprint at arXiv (2017).
Acknowledgements
We thank past and present members of the Segal group and the Cardiology Department at Rabin Medical Center for useful discussions. Y.T.-B. received a research grant from the Tel Aviv University Faculty Funds, and from the Gassner Fund for Medical Research. N.B. received a PhD scholarship for Data Science from the Israeli Council for Higher Education (CHE) via the Weizmann Data Science Research Center and is supported by a research grant from the Estate of Tully and Michele Plesser. E.S. is supported by the Crown Human Genome Center, by D. L. Schwarz, J. N. Halpern and L. Steinberg, and by grants funded by the European Research Council and the Israel Science Foundation. M.-E.D. is supported by the NIHR Imperial Biomedical Research Centre, and by grants from the French National Research Agency (ANR-10-LABX-46 [European Genomics Institute for Diabetes]), from the National Center for Precision Diabetic Medici–e – PreciDIAB, which is jointly supported by the French National Agency for Research (ANR-18-IBHU-0001), by the European Union (FEDER), by the Hauts-de-France Regional Council (Agreement 20001891/NP0025517), by the European Metropolis of Lille (MEL, Agreement 2019_ESR_11) and Isite ULNE (R-002-20-TALENT-DUMAS), also jointly funded by ANR (ANR-16-IDEX-0004-ULNE), the Hauts-de-France Regional Council (20002845) and by the European Metropolis of Lille (MEL). K.C. is supported by Medical Research Council (MRC) Skills Development Fellowship (grant number MR/S020039/1) and Wellcome Trust funded Institutional Strategic Support Fellowship (grant number 204834/Z/16/Z).
Author information
Authors and Affiliations
Contributions
Y.T.-B. and N.B. conceived the project, designed and conducted all analyses, interpreted the results, wrote the manuscript and are listed in random order. N.R. performed metabolomics analyses and interpreted the results. A.G. conducted microbiome analysis. Y.B. conducted microbiome analysis and provided additional information regarding SGB 4712. A.A.S., A.S., C.C.-A., Z.A. and Y.H. coordinated and designed data collection. M.L.-P. and A.W. developed protocols, performed microbiome sequencing, and processed serum samples. A.W. designed the project and oversaw sample collection and processing. K.C., S.K.F., S.F., M.-E.D., S.D.E. and O.P. performed the replication analysis on the MetaCardis cohort. R.K. and E.S. conceived and directed the project and analyses, designed the analyses, interpreted the results and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Medicine thanks Matej Oresic, Manuel Mayr, Ellen Blaak and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Michael Basson was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Cohort selection and data acquisition pipeline.
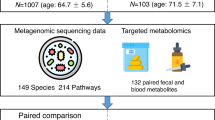
This study includes a total of 199 participants with ACS and 970 non-ACS individuals. Each cell shows the number of individuals who were profiled for the corresponding omic platform indicated on the left. Colored bars connecting cells represent the number of overlapping individuals. For example, there were 169 non-ACS individuals that were profiled both for serum metabolomics using the Metabolon platform and for microbiome composition. The 156 samples of individuals with ACS that were profiled using the Metabolon platform are the first to be enrolled in this study. The 473 samples of non-ACS individuals that were profiled using the Metabolon platform, were profiled as part of our previous study (Bar et al. 2020). All samples of individuals with ACS (n = 191) and of non-ACS individuals (n = 961) for which we had available serum obtained during their recruitment, were profiled using the Nightingale platform. While microbiome data were available for all individuals with and without ACS, we only considered samples for which the collection, DNA extraction and sequencing procedures were identical (n = 199 for ACS; n = 340 for non-ACS). Differential abundance analysis was performed based on subcohorts resulting from 1:1 matching for age, sex, BMI, DM, and smoking status. ACS, Acute Corony Syndrome; BMI Body Mass Index; DM, Diabetes Mellitius.
Extended Data Fig. 2 Breakdown of ACS serum metabolomics pattern by the origin of metabolites and biological pathway.
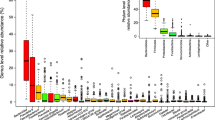
(a) Box plots (y axis: center, median; box, IQR; whiskers, 1.5×IQR) showing the explained variance of metabolites by different feature groups (x-axis) separated to metabolites enriched in ACS (N = 175; orange) and enriched in matched non-ACS controls (N = 358; blue). (b) explained variance of metabolites (y axis: center, median; box, IQR; whiskers, 1.5×IQR) by their super pathways (x axis) separated to metabolites enriched in ACS (orange) and enriched in matched non-ACS controls (blue). The number of metabolites per group is shown below each box. Trad., Traditional; C&V, cofactors and vitamins.
Extended Data Fig. 3 Depletion of ACS-related bacteria SGB 4712 replicates in an independent validation cohort.
(a) Box plots showing the relative abundance of the unknown bacterial species SGB 4712 (y-axis: center, median; box, IQR; whiskers, 1.5×IQR; log scaled) in our ACS and matched controls (x-axis; n = 80 each). The P-value shown is computed using the two-sided Mann–Whitney U-test. (b) Relative abundance of the unknown bacterial species SGB 4712 (y-axis: center, median; box, IQR; whiskers, 1.5×IQR; log scaled) in four groups from the MetaCardis validation cohort (x-axis; HC, healthy controls, n = 275; MMC, metabolically matched controls, n = 218; UMCC, untreated metabolically compromised controls, n = 211; IHD, ischaemic heart disease, n = 319). The P-value shown is computed using the two-sided Mann–Whitney U-test. r.a., relative abundance.
Extended Data Fig. 4 Clinical data correlates with metabolic deviations.
(a) The mean weighted R2 of genetics for ACS-enriched metabolites (y-axis) versus chronological age (x-axis). Dots are colored by sex. Spearman correlation is computed over all samples (Spearman ⍴ = 0.18; p = 0.032). (b) The mean weighted R2 of traditional risk factors for ACS-depleted metabolites (y axis) versus chronological age (x axis; Spearman ⍴ = 0.33; p = 7.7 × 10−5). (c) The mean weighted R2 of genetics for ACS-enriched metabolites (y axis: center, median; box, IQR; whiskers, 1.5×IQR) in ACS patients who had a combined CVD outcome (defined as either: acute myocardial infarction, acute stroke, unplanned PCI, or cardiovascular-related death; x axis) versus not (two-sided Mann–Whitney U-test, p = 0.002).
Extended Data Fig. 5 Replication of higher predicted BMI in ACS individuals based on NMR metabolomics.
Figure panels refer to results of serum metabolomics-based prediction model of BMI trained in a non-ACS control cohort (n = 763) and evaluated on held-out test sets consisting of both controls (n = 179) and individuals with ACS (n = 179; Methods). (a) Measured (x axis) versus predicted (y-axis) BMI for both controls (blue) and ACS (orange) individuals. Line and shaded coloring represent the fitting of a linear model and the 95% confidence interval. (b) Difference between predicted and measured BMI (y axis: center, median; box, IQR; whiskers, 1.5×IQR) of individuals, binned into three BMI groups (<25, 25-30, >30; x-axis). The P-values shown are computed using the two-sided Mann–Whitney U-test. BMI, body mass index.
Extended Data Fig. 6 Replication of higher predicted BMI in IHD individuals in the MetaCardis study.
Figure panels refer to results of serum metabolomics-based prediction model of BMI trained in a cohort of individuals without IHD and evaluated on held-out test sets consisting of both 319 IHD and 319 non-IHD individuals (Methods). (a) Measured (x axis) versus predicted (y axis) BMI for healthy controls (HC; blue), metabolically matched controls (MMC; blue), and untreated metabolically compromised controls (UMCC; blue), and individuals with ischaemic heart disease (IHD; orange). Line and shaded coloring represent the fitting of a linear model and the 95% confidence interval. (b) Difference between predicted and measured BMI (y axis: center, median; box, IQR; whiskers, 1.5×IQR) of individuals, binned into three BMI groups (<25, 25-30, >30; x axis). (c) Same as in (b) only for individuals with IHD, and each bin is separated into normoglycemic versus T2DM patients. Higher predicted BMI is associated with an increased incidence of T2DM (OR = 1.13, 95% CI = 1.05-1.22, p = 0.002; a logistic regression model adjusted for BMI and age; Methods). The p values shown are computed using the two-sided Mann–Whitney U-test. BMI, body mass index; T2DM, type 2 diabetes mellitus; OR, odds ratio; CI, confidence interval.
Supplementary information
Supplementary Information
Supplementary Notes 1 and 2.
Supplementary Tables 1–20
Guide and legend are in the ‘Guide’ tab of the spreadsheet.
Rights and permissions
About this article
Cite this article
Talmor-Barkan, Y., Bar, N., Shaul, A.A. et al. Metabolomic and microbiome profiling reveals personalized risk factors for coronary artery disease. Nat Med 28, 295–302 (2022). https://doi.org/10.1038/s41591-022-01686-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-022-01686-6
This article is cited by
-
Oral-gut microbial transmission promotes diabetic coronary heart disease
Cardiovascular Diabetology (2024)
-
Chinmedomics: a potent tool for the evaluation of traditional Chinese medicine efficacy and identification of its active components
Chinese Medicine (2024)
-
Effects of dietary fibre on metabolic health and obesity
Nature Reviews Gastroenterology & Hepatology (2024)
-
Effects of a personalized nutrition program on cardiometabolic health: a randomized controlled trial
Nature Medicine (2024)
-
Plasma metabolomic differences in early-onset compared to average-onset colorectal cancer
Scientific Reports (2024)