Abstract
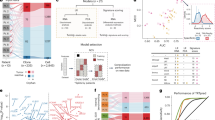
Severe immune-related adverse events (irAEs) occur in up to 60% of patients with melanoma treated with immune checkpoint inhibitors (ICIs). However, it is unknown whether a common baseline immunological state precedes irAE development. Here we applied mass cytometry by time of flight, single-cell RNA sequencing, single-cell V(D)J sequencing, bulk RNA sequencing and bulk T cell receptor (TCR) sequencing to study peripheral blood samples from patients with melanoma treated with anti-PD-1 monotherapy or anti-PD-1 and anti-CTLA-4 combination ICIs. By analyzing 93 pre- and early on-ICI blood samples and 3 patient cohorts (n = 27, 26 and 18), we found that 2 pretreatment factors in circulation—activated CD4 memory T cell abundance and TCR diversity—are associated with severe irAE development regardless of organ system involvement. We also explored on-treatment changes in TCR clonality among patients receiving combination therapy and linked our findings to the severity and timing of irAE onset. These results demonstrate circulating T cell characteristics associated with ICI-induced toxicity, with implications for improved diagnostics and clinical management.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Bulk and single-cell expression data generated in this work have been deposited in GEO under accession no. GSE186144. All requests for raw data will be promptly reviewed by the corresponding authors to determine whether the request is subject to any confidentiality obligations. Any data that can be shared will be released via a material transfer agreement. The published expression datasets analyzed in this work (Supplementary Table 20) are available from GEO with accession nos. GSE50772, GSE61635, GSE72509, GSE126124, GSE3365 and GSE100833. Processed CyTOF, MiXCR and immunoSEQ data are publicly available from https://doi.org/10.25936/f3np-k536. Additional data supporting the findings in this work are available in the main text, figures, extended data and supplementary files.
Code availability
Custom scripts for the training and validation of composite models, evaluating freedom from severe toxicity and generating related figures are publicly available from https://doi.org/10.25936/f3np-k536.
References
Kumar, V. et al. Current diagnosis and management of immune related adverse events (irAEs) induced by immune checkpoint inhibitor therapy. Front. Pharmacol. 8, 49 (2017).
Larkin, J. et al. Combined nivolumab and ipilimumab or monotherapy in untreated melanoma. N. Engl. J. Med. 373, 23–34 (2015).
Hodi, F. S. et al. Nivolumab plus ipilimumab or nivolumab alone versus ipilimumab alone in advanced melanoma (CheckMate 067): 4-year outcomes of a multicentre, randomised, phase 3 trial. Lancet Oncol. 19, 1480–1492 (2018).
Postow, M. A., Sidlow, R. & Hellmann, M. D. Immune-related adverse events associated with immune checkpoint blockade. N. Engl. J. Med. 378, 158–168 (2018).
Wolchok, J. D. et al. Overall survival with combined nivolumab and ipilimumab in advanced melanoma. N. Engl. J. Med. 377, 1345–1356 (2017).
Hamid, O. et al. Five-year survival outcomes for patients with advanced melanoma treated with pembrolizumab in KEYNOTE-001. Ann. Oncol. 30, 582–588 (2019).
Robert, C. et al. Pembrolizumab versus ipilimumab in advanced melanoma (KEYNOTE-006): post-hoc 5-year results from an open-label, multicentre, randomised, controlled, phase 3 study. Lancet Oncol. 20, 1239–1251 (2019).
Robert, C. et al. Pembrolizumab versus ipilimumab in advanced melanoma. N. Engl. J. Med. 372, 2521–2532 (2015).
Schachter, J. et al. Pembrolizumab versus ipilimumab for advanced melanoma: final overall survival results of a multicentre, randomised, open-label phase 3 study (KEYNOTE-006). Lancet 390, 1853–1862 (2017).
Johnson, D. B. et al. Fulminant myocarditis with combination immune checkpoint blockade. N. Engl. J. Med. 375, 1749–1755 (2016).
Wang, D. Y. et al. Fatal toxic effects associated with immune checkpoint inhibitors: a systematic review and meta-analysis. JAMA Oncol. 4, 1721–1728 (2018).
Haanen, J. B. A. G. et al. Management of toxicities from immunotherapy: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann. Oncol. 28, iv119–iv142 (2017).
Das, S. & Johnson, D. B. Immune-related adverse events and anti-tumor efficacy of immune checkpoint inhibitors. J. Immunother. Cancer 7, 306 (2019).
Johnson, D. B. et al. A case report of clonal EBV-like memory CD4+ T cell activation in fatal checkpoint inhibitor-induced encephalitis. Nat. Med. 25, 1243–1250 (2019).
Tahir, S. A. et al. Autoimmune antibodies correlate with immune checkpoint therapy-induced toxicities. Proc. Natl Acad. Sci. USA 116, 22246–22251 (2019).
Shahabi, V. et al. Gene expression profiling of whole blood in ipilimumab-treated patients for identification of potential biomarkers of immune-related gastrointestinal adverse events. J. Transl. Med. 11, 75 (2013).
Tarhini, A. A. et al. Baseline circulating IL-17 predicts toxicity while TGF-β1 and IL-10 are prognostic of relapse in ipilimumab neoadjuvant therapy of melanoma. J. Immunother. Cancer 3, 39 (2015).
Fujimura, T. et al. Serum levels of soluble CD163 and CXCL5 may be predictive markers for immune-related adverse events in patients with advanced melanoma treated with nivolumab: a pilot study. Oncotarget 9, 15542–15551 (2018).
Das, R. et al. Early B cell changes predict autoimmunity following combination immune checkpoint blockade. J. Clin. Invest. 128, 715–720 (2018).
Subudhi, S. K. et al. Clonal expansion of CD8 T cells in the systemic circulation precedes development of ipilimumab-induced toxicities. Proc. Natl Acad. Sci. USA 113, 11919–11924 (2016).
Chaput, N. et al. Baseline gut microbiota predicts clinical response and colitis in metastatic melanoma patients treated with ipilimumab. Ann. Oncol. 30, 2012 (2019).
Dubin, K. et al. Intestinal microbiome analyses identify melanoma patients at risk for checkpoint-blockade-induced colitis. Nat. Commun. 7, 10391 (2016).
Jing, Y. et al. Multi-omics prediction of immune-related adverse events during checkpoint immunotherapy. Nat. Commun. 11, 4946 (2020).
Lim, S. Y. et al. Circulating cytokines predict immune-related toxicity in melanoma patients receiving anti-PD-1-based immunotherapy. Clin. Cancer Res. 25, 1557–1563 (2019).
Pavan, A. et al. Peripheral blood markers identify risk of immune-related toxicity in advanced non-small cell lung cancer treated with immune-checkpoint inhibitors. Oncologist 24, 1128–1136 (2019).
Andrews, M. C. et al. Gut microbiota signatures are associated with toxicity to combined CTLA-4 and PD-1 blockade. Nat. Med. 27, 1432–1441 (2021).
Hutchinson, J. A. et al. Virus-specific memory T cell responses unmasked by immune checkpoint blockade cause hepatitis. Nat. Commun. 12, 1439 (2021).
Yasuda, Y. et al. CD4+ T cells are essential for the development of destructive thyroiditis induced by anti-PD-1 antibody in thyroglobulin-immunized mice. Sci. Transl. Med. 13, eabb7495 (2021).
Morad, G., Helmink, B. A., Sharma, P. & Wargo, J. A. Hallmarks of response, resistance, and toxicity to immune checkpoint blockade. Cell 184, 5309–5337 (2021).
Marschner, D. et al. MicroRNA-146a regulates immune-related adverse events caused by immune checkpoint inhibitors. JCI Insight 5, e132334 (2020).
Auslander, N. et al. Robust prediction of response to immune checkpoint blockade therapy in metastatic melanoma. Nat. Med. 24, 1545–1549 (2018).
Krieg, C. et al. High-dimensional single-cell analysis predicts response to anti-PD-1 immunotherapy. Nat. Med. 24, 144–153 (2018).
Liu, D. et al. Integrative molecular and clinical modeling of clinical outcomes to PD1 blockade in patients with metastatic melanoma. Nat. Med. 25, 1916–1927 (2019).
Jacquelot, N. et al. Predictors of responses to immune checkpoint blockade in advanced melanoma. Nat. Commun. 8, 592 (2017).
Riaz, N. et al. Tumor and microenvironment evolution during immunotherapy with nivolumab. Cell 171, 934–949.e16 (2017).
Huang, A. C. et al. T-cell invigoration to tumour burden ratio associated with anti-PD-1 response. Nature 545, 60–65 (2017).
Fairfax, B. P. et al. Peripheral CD8+ T cell characteristics associated with durable responses to immune checkpoint blockade in patients with metastatic melanoma. Nat. Med. 26, 193–199 (2020).
Bieber, A. K., Yin, L. & Lo Sicco, K. Pruritus and tense bullae after discontinuation of pembrolizumab in a patient with renal cell carcinoma. JAMA 324, 1453–1454 (2020).
Lopez, A. T. & Geskin, L. A case of nivolumab-induced bullous pemphigoid: review of dermatologic toxicity associated with programmed cell death protein-1/programmed death ligand-1 inhibitors and recommendations for diagnosis and management. Oncologist 23, 1119–1126 (2018).
Singer, S., Nelson, C. A., Lian, C. G., Dewan, A. K. & LeBoeuf, N. R. Nonbullous pemphigoid secondary to PD-1 inhibition. JAAD Case Rep. 5, 898–903 (2019).
Croft, M., So, T., Duan, W. & Soroosh, P. The significance of OX40 and OX40L to T-cell biology and immune disease. Immunol. Rev. 229, 173–191 (2009).
Campbell, J. J. et al. CCR7 expression and memory T cell diversity in humans. J. Immunol. 166, 877–884 (2001).
Shifrut, E. et al. Genome-wide CRISPR screens in primary human T cells reveal key regulators of immune function. Cell 175, 1958–1971.e15 (2018).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e29 (2021).
Chiffelle, J. et al. T-cell repertoire analysis and metrics of diversity and clonality. Curr. Opin. Biotechnol. 65, 284–295 (2020).
Rényi, A. On the foundations of information theory. Rev. Int. Stat. Inst. 33, 1–14 (1965).
Robert, L. et al. CTLA4 blockade broadens the peripheral T-cell receptor repertoire. Clin. Cancer Res. 20, 2424–2432 (2014).
Sims, J. S. et al. Diversity and divergence of the glioma-infiltrating T-cell receptor repertoire. Proc. Natl Acad. Sci. USA 113, E3529–E3537 (2016).
Newman, A. M. et al. Determining cell type abundance and expression from bulk tissues with digital cytometry. Nat. Biotechnol. 37, 773–782 (2019).
Gentles, A. J. et al. The prognostic landscape of genes and infiltrating immune cells across human cancers. Nat. Med. 21, 938–945 (2015).
Newman, A. M. et al. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 12, 453–457 (2015).
Thorsson, V. et al. The immune landscape of cancer. Immunity 48, 812–830.e14 (2018).
Bolotin, D. A. et al. MiXCR: software for comprehensive adaptive immunity profiling. Nat. Methods 12, 380–381 (2015).
Eggermont, A. M. M., Crittenden, M. & Wargo, J. Combination immunotherapy development in melanoma. Am. Soc. Clin. Oncol. Educ. Book 38, 197–207 (2018).
Boutros, C. et al. Safety profiles of anti-CTLA-4 and anti-PD-1 antibodies alone and in combination. Nat. Rev. Clin. Oncol. 13, 473–486 (2016).
Burczynski, M. E. et al. Molecular classification of Crohn’s disease and ulcerative colitis patients using transcriptional profiles in peripheral blood mononuclear cells. J. Mol. Diagn. 8, 51–61 (2006).
Carpintero, M. F. et al. Diagnosis and risk stratification in patients with anti-RNP autoimmunity. Lupus 24, 1057–1066 (2015).
Hung, T. et al. The Ro60 autoantigen binds endogenous retroelements and regulates inflammatory gene expression. Science 350, 455–459 (2015).
Kennedy, W. P. et al. Association of the interferon signature metric with serological disease manifestations but not global activity scores in multiple cohorts of patients with SLE. Lupus Sci. Med. 2, e000080 (2015).
Palmer, N. P. et al. Concordance between gene expression in peripheral whole blood and colonic tissue in children with inflammatory bowel disease. PLoS ONE 14, e0222952 (2019).
Peters, L. A. et al. A functional genomics predictive network model identifies regulators of inflammatory bowel disease. Nat. Genet. 49, 1437–1449 (2017).
Menzies, A. M. et al. Anti-PD-1 therapy in patients with advanced melanoma and preexisting autoimmune disorders or major toxicity with ipilimumab. Ann. Oncol. 28, 368–376 (2017).
Brown, L. J. et al. Combination anti-PD1 and ipilimumab therapy in patients with advanced melanoma and pre-existing autoimmune disorders. J. Immunother. Cancer 9, e002121 (2021).
Johnson, D. B. et al. Ipilimumab therapy in patients with advanced melanoma and preexisting autoimmune disorders. JAMA Oncol. 2, 234–240 (2016).
Tang, S.-Q. et al. The pattern of time to onset and resolution of immune-related adverse events caused by immune checkpoint inhibitors in cancer: a pooled analysis of 23 clinical trials and 8,436 patients. Cancer Res. Treat. 53, 339–354 (2021).
Nabet, B. Y. et al. Noninvasive early identification of therapeutic benefit from immune checkpoint inhibition. Cell 183, 363–376.e13 (2020).
Rizvi, N. A. et al. Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348, 124–128 (2015).
Steele, N. G. et al. Multimodal mapping of the tumor and peripheral blood immune landscape in human pancreatic cancer. Nat. Cancer 1, 1097–1112 (2020).
Amir, E. D. et al. viSNE enables visualization of high dimensional single-cell data and reveals phenotypic heterogeneity of leukemia. Nat. Biotechnol. 31, 545–552 (2013).
Van Gassen, S. et al. FlowSOM: using self-organizing maps for visualization and interpretation of cytometry data. Cytometry A 87, 636–645 (2015).
Amir, E. D. et al. Development of a comprehensive antibody staining database using a standardized analytics pipeline. Front. Immunol. 10, 1315 (2019).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e21 (2019).
Soneson, C., Love, M. I. & Robinson, M. D. Differential analyses for RNA-seq: transcript-level estimates improve gene-level inferences. F1000Res 4, 1521 (2015).
Chen, G. M. et al. Integrative bulk and single-cell profiling of pre-manufacture T-cell populations reveals factors mediating long-term persistence of CAR T-cell therapy. Cancer Discov. 11, 2186–2199 (2021).
Kaech, S. M., Wherry, E. J. & Ahmed, R. Effector and memory T-cell differentiation: implications for vaccine development. Nat. Rev. Immunol. 2, 251–262 (2002).
Sprent, J. & Surh, C. D. T cell memory. Annu. Rev. Immunol. 20, 551–579 (2002).
van den Broek, T., Borghans, J. A. M. & van Wijk, F. The full spectrum of human naive T cells. Nat. Rev. Immunol. 18, 363–373 (2018).
Dixon, P. VEGAN, a package of R functions for community ecology. J. Veg. Sci. 14, 927–930 (2003).
Jiang, H., Lei, R., Ding, S.-W. & Zhu, S. Skewer: a fast and accurate adapter trimmer for next-generation sequencing paired-end reads. BMC Bioinformatics 15, 182 (2014).
Borcherding, N. et al. Mapping the immune environment in clear cell renal carcinoma by single-cell genomics. Commun. Biol. 4, 122 (2021).
Motulsky, H. J. & Brown, R. E. Detecting outliers when fitting data with nonlinear regression—a new method based on robust nonlinear regression and the false discovery rate. BMC Bioinformatics 7, 123 (2006).
Gautier, L., Cope, L., Bolstad, B. M. & Irizarry, R. A. affy—analysis of Affymetrix GeneChip data at the probe level. Bioinformatics 20, 307–315 (2004).
Lipták, T. On the combination of independent tests. Magyar Tud. Akad. Mat. Kutató Int. Közl. 3, 171–197 (1958).
Stouffer, S. A., Suchman, E. A., Devinney, L. C., Star, S. A., & Williams, R. M., Jr. in Studies in Social Psychology in World War II (Princeton Univ. Press, 82–154 1949).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Cohen, J. Statistical Power Analysis for the Behavioral Sciences (L. Erlbaum Associates, 1988).
Acknowledgements
We thank the patients and families involved in this study. We thank A. Nassar and L. Chen for technical assistance with the CyTOF experiments, including provision of resources, staining and processing of samples. We thank G. Ansstas and C. Kaufman for clinical samples. We also thank T. Ley, S. Devarakonda and M. Tal for providing critical feedback on the manuscript. This work was supported by grants from the National Cancer Institute (no. K08CA238711 to A.A.C., no. K08CA237727 to D.Y.C., no. R01CA238471 to K.D., no. R21CA218950 to R.H. and no. R00CA187192 to A.M.N.), National Heart, Lung, and Blood Institute (no. T35HL007649 to A.N.), National Institute of Arthritis and Musculoskeletal and Skin Diseases (no. R01AR077926 to K.D.), a fellowship from the Natural Sciences and Engineering Research Council of Canada (A.X.L.), the Cancer Research Foundation Young Investigator Award (A.A.C.), V Foundation for Cancer Research V Scholar Award (A.A.C.), Washington University Alvin J. Siteman Cancer Research Fund (A.A.C.), Yale Cancer Center Meyers Award (M.S. and R.H.), a 10x Genomics Pilot Program Award (R.H.), the Melanoma Research Alliance (no. 137453 and no. 828544 to R.H.), the Virginia and D.K. Ludwig Fund for Cancer Research (A.M.N.), Stinehart-Reed Foundation (A.M.N.), Stanford Bio-X Interdisciplinary Initiatives Seed Grants Program (IIP) (A.M.N.) and Donald E. and Delia B. Baxter Foundation (A.M.N.).
Author information
Authors and Affiliations
Contributions
A.X.L., A.A.C., A.N., R.H. and A.M.N. conceived of the study, developed strategies for related experiments and wrote the paper. A.X.L., A.A.C., A.N. and A.M.N. performed the data analysis and interpretation with assistance from N.E., C.B.S., B.A.L., G.S.G. and F.K. M.D.V., A.U. and T.B. performed the cytometry experiments with data analysis by A.X.L., N.E., M.D.V. and A.U. with assistance from M.R.V. A.B., P.K.H., D.Y.C. and R.H. collected the patient specimens, which were processed for expression profiling by A.B. D.Y.C., K.D. and M.S. determined the clinical characteristics and outcomes with assistance from A.X.L. and A.A.C. A.X.L., A.A.C., A.B., B.E.T., D.Y.C., K.D., R.H. and A.M.N. curated the clinical data. A.M.N. and R.H. are co-senior authors. All authors commented on the manuscript at all stages.
Corresponding authors
Ethics declarations
Competing interests
A.A.C. has patent filings related to cancer biomarkers and digital cytometry and has served as an advisor/consultant to Roche, AstraZeneca, Daiichi Sankyo, Tempus, Geneoscopy, NuProbe, Fenix Group International and Guidepoint. A.A.C. has stock options in Geneoscopy, research support from Roche and ownership interests in Droplet Biosciences. M.S. has consulted for Idera Pharmaceuticals, Regeneron Pharmaceuticals, Apexigen, Alligator Bioscience, Verastem Oncology, Agenus, Rubius Therapeutics, Bristol Myers Squibb, Genentech-Roche, Boston Pharmaceuticals, Servier Laboratories, Adaptimmune Therapeutics, Immunocore, Dragonfly Therapeutics, Pierre Fabre Pharmaceuticals, Molecular Partners, Boehringer Ingelheim, Innate Pharma, Nektar Therapeutics, Pieris Pharmaceuticals, Numab Therapeutics, Abbvie, Zelluna Immunotherapy, Seattle Genetics/Seagen, Genocea Biosciences, GI Innovation, Chugai-Roche, BioNTech, Eli Lilly, Modulate Therapeutics, Array Biopharma, AstraZeneca and Genmab. M.S. has stock options in EvolveImmune, NextCure, Repertoire Immune Medicines, Adaptive Biotechnologies, Actym Therapeutics and Amphivena Therapeutics and has stock ownership in GlaxoSmithKline and Johnson & Johnson. A.M.N. has patent filings related to expression deconvolution, digital cytometry and cancer biomarkers and has served as an advisor/consultant to Roche, Merck and CiberMed. The other authors declare no conflicts of interest.
Peer review information
Nature Medicine thanks the anonymous reviewer(s) for their contribution to the peer review of this work. Saheli Sadanand was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Quality control and extended characterization of cell states identified by unsupervised clustering of scRNA-seq data.
a, UMAP representation of pretreatment peripheral blood leukocytes profiled by droplet-based scRNA-seq (10x Genomics) from 13 patients with metastatic melanoma, colored by major cell lineages, severe irAE status, TCR expression by scV(D)J-seq, and BCR expression by scV(D)J-seq (related to Fig. 3a). b, Unsupervised hierarchical clustering (average linkage) of the mean log2 transcriptome per CD4 T cell cluster identified from scRNA-seq data. c, Dot plot showing the average expression of key activation (HLA-DX, MKI67) and lineage markers (SELL, CCR7) in CD4 T cell clusters. d, Same as Fig. 3b but showing all pairwise combinations of scRNA-seq clusters within each of the major cell types analyzed (B cells, CD4 T cells, CD8 T cells, NK cells, monocytes). Across 82 possible pairwise combinations, CD4 T 5 + 3 achieved the highest Spearman correlation against CD4 TEM levels enumerated by CyTOF and the strongest association with severe irAE development. Cells annotated as ‘T/NKT’ were collapsed into CD8 T cells. e, Same as panel d but showing all pairwise combinations ranked by the mean of each feature following unit variance normalization (mean of 0 and standard deviation of 1). In this analysis, the –log10 P-value for the association with severe irAE (two-sided, unpaired Wilcoxon rank sum test) was normalized to unit variance without considering the direction of the association.
Extended Data Fig. 2 Analysis of scRNA-seq states identified by reference-guided annotation.
a, UMAP projections of scRNA-seq data generated in this work, embedded and labeled by Azimuth using a reference PBMC atlas of 162k cells profiled by scRNA-seq and 228 antibodies (Methods). b, Confusion matrix showing the agreement between phenotypic labels determined by marker genes and unsupervised clustering (rows; related to Fig. 3a and Extended Data Fig. 1a) versus reference-guided annotation with Azimuth (columns). In total, 85% of single cells assigned to a major lineage group by Azimuth (B cells, CD4 T, CD8 T, NK cells, monocytes) were assigned to the same identity by canonical marker gene assessment. Given the absence of NKT cells in the reference atlas used for Azimuth, the T/NKT cluster defined by unsupervised analysis was relabeled as CD8 T cells. c, Same analysis as in Fig. 3b but shown for all 27 phenotypic states identified by Azimuth. Among these states, CD4 TEM was most associated with severe irAE and CyTOF-enumerated CD4 TEM. A population combining CD4 TEM and CD4 Proliferating states was also strongly associated with severe irAE. The latter showed the highest expression of HLA-DX and lowest expression of SELL (panel d), consistent with an activated CD4 TEM phenotype. d, Dot plot depicting key activation and lineage markers among CD4 T cell states annotated by Azimuth. e, Violin plots showing protein expression levels imputed by Azimuth using antibody-derived tag (ADT) data, supporting the combination of CD4 TEM and CD4 Proliferating states in panels c and f. f, Performance of top-ranking cell subsets identified by Azimuth and unsupervised clustering for prediction of severe irAEs. The combined CD4 T 5 + 3 clusters (Fig. 3b) were more associated with severe irAE and CyTOF than the top-ranking reference-guided population (panel c). Statistical significance was calculated using a two-sided, unpaired Wilcoxon rank sum test. Data in all panels shown are from the 13 samples profiled by scRNA-seq in Fig. 3.
Extended Data Fig. 3 Analysis of activated, resting, and parental T cell subsets in relation to severe irAE development.
a, Association between severe irAE development and pretreatment levels of T cell states identified by unsupervised clustering (left) and memory-like T cell states identified by Azimuth (right) in 13 PBMC samples profiled by scRNA-seq (Figs. 1 and 3a). Activated cells were defined as those expressing HLA-DX or MKI67 (CPM > 0); resting cells were defined by the absence of HLA-DX and MKI67 expression (CPM = 0). b, Left: Association between severe irAE development and pretreatment levels of memory T cell subsets, total CD4 and CD8 T cells, and total T cells quantified by CyTOF, for all 18 patients analyzed in the single-cell discovery cohort (Figs. 1 and 2a). Activated phenotypes were defined as CD38+ or HLA-DR+ or Ki67+. Resting phenotypes were defined as CD38–HLA-DR–Ki67–. Right: ROC plot showing the performance of activated and resting CD4 TEM subsets (left panel) for predicting severe irAE development. Cell fractions were assessed relative to total PBMC content. Statistical significance in a, b was determined by a two-sided, unpaired Wilcoxon rank sum test and nominal –log10 P-values are displayed. –log10 P-values were further multiplied by –1 for associations with no severe irAE. See also Supplementary Table 6.
Extended Data Fig. 4 Extended characterization of immune repertoire diversity from single-cell V(D)J sequencing data.
a, Key TCR diversity measures and the impact of cell abundance, TCR richness, and distinct clonal repertoires on such measures. Hypothetical CD4 naïve and TEM cell subsets are shown as examples. Triangles depicting differences in magnitude are not drawn to scale. b, Mean Shannon entropy versus mean clonality (1 – Pielou’s evenness) for each CD4 T cell state identified by unsupervised clustering of scRNA-seq data. CD4 T 5 + 3 (Fig. 3b,c), a TEM state enriched for activated cells, shows elevated clonality relative to other CD4 states, as expected for this phenotype77, while also showing higher diversity (Shannon entropy), indicating elevated richness. c, Distribution of EM-like CD4 T cell states (from Fig. 3f) with available scTCR clonotype data. d, Association between severe irAE development and TCR diversity (Shannon entropy) in pseudo-bulk T cells from pretreatment blood, shown for all T cell states identified by scRNA-seq (left) and after the removal of the EM-like states indicated in panel c (no severe irAE, n = 5 patients; severe irAE, n = 4 patients). e, Same as d but shown for EM-like states alone. f, Area under the curve (AUC) for the association between pretreatment peripheral TCR diversity (Shannon entropy) and severe irAE development, shown for all combinations of the constituent cell states in e, including the combined CD4 T 5 + 3 cluster after restricting to activated cells (CPM > 0 for HLA-DX or MKI67). Of note, no other combination of activated EM-like states achieved an AUC > 0.85 in this analysis. g, BCR clonotype diversity (Shannon entropy), shown for each B cell state identified by unsupervised clustering (Fig. 3a). In b, d–f, only patients with at least 100 TCR clones were analyzed (n = 9; Methods). The same patients were analyzed in g for consistency. In panels d, e, and g, center lines, bounds of the box, and whiskers indicate medians, 1st and 3rd quartiles, and minimum and maximum values, respectively.
Extended Data Fig. 5 Validation of CIBERSORTx by single-cell analysis.
a, Expression of developmentally-regulated marker genes in major CD4 T cell subsets from the LM22 signature matrix (MAS5 normalized), showing that the LM22 reference signature for activated CD4 memory T cells has a TEM profile. b, CIBERSORTx versus mass cytometry for enumeration of activated CD4 memory T cells in the pretreatment peripheral blood of 17 metastatic melanoma patients (Supplementary Table 1). A linear regression line with 95% confidence band is shown. Concordance and significance were determined by Pearson r and a two-sided t test, respectively. While activated CD4 memory T cells quantitated by CyTOF were defined by CD38 expression in this plot, other activated CD4 TEM subsets were also significantly correlated with CIBERSORTx (panel c). c, Cross correlation plot of lymphocyte subset frequencies determined by CyTOF and CIBERSORTx. Act., Activated. d, Correlation between activated CD4 memory T cell levels inferred by CIBERSORTx and 14 memory T cell states profiled by CyTOF, including CD38+ activated subsets manually gated within each population, in PBMCs from 17 metastatic melanoma patients (Supplementary Table 1). e, Scatter plot depicting the global correlation of lymphocyte subsets enumerated by CIBERSORTx and flow cytometry in peripheral blood samples from five healthy subjects profiled by bulk RNA-seq (Supplementary Table 1). A linear regression line with 95% confidence band is shown. Concordance and significance were determined by Pearson r and a two-sided t test, respectively. As monocytes were variably underestimated by cytometry compared to complete blood counts, all results in b–e are expressed as a function of total lymphocytes. f, Distribution of activated CD4 memory T cell levels quantitated by CyTOF (CD38+, HLA-DR+ or Ki67+ CD4 TEM cells, n = 28 patients), scRNA-seq (HLA-DX+ or MKI67+ cells within CD4 T clusters 5 and 3, n = 13 patients), and CIBERSORTx (n = 60 patients) across all irAE-evaluable samples profiled by each modality in this work (Supplementary Table 1). Box center lines, bounds of the box, and whiskers indicate medians, 1st and 3rd quartiles, and minimum and maximum values, respectively. Statistical significance was determined by a Kruskal-Wallis test. n.s., not significant (P > 0.05).
Extended Data Fig. 6 Extended analysis of TCR diversity from pretreatment peripheral blood expression profiles.
a–d, Association between baseline bulk TCR diversity and the highest irAE grade observed for each patient in bulk cohorts 1 and 2 (Supplementary Tables 7 and 9), shown for two diversity measures (a and c, Shannon entropy; b and d, Gini-Simpson index) and stratified by therapy type. In a and b, patients treated with combination therapy are stratified by future irAE status: no severe irAE (n = 10) versus severe irAE (n = 14 patients) (left) and irAE grade (right): 0/1 (n = 3), 2 (n = 7), 3 (n = 12), and 4 (n = 2). In c and d, patients treated with PD1 monotherapy are stratified by future irAE status: no severe irAE (n = 26) versus severe irAE (n = 3 patients) (left) and irAE grade (right): 0/1 (n = 19), 2 (n = 7), 3 (n = 2), and 4 (n = 1). Two-group comparisons were assessed by a two-sided, unpaired Wilcoxon rank sum test. n.s., not significant (P > 0.05). Linear regression was applied to evaluate the median value of each measure grouped by irAE grade (insets). The significance of linear concordance was determined by a two-sided t test. Grades 0 and 1 reflect no toxicity and asymptomatic toxicity, respectively, and were combined. In all panels, the box center lines, bounds of the box, and whiskers denote medians, 1st and 3rd quartiles, and minimum and maximum values within 1.5 × IQR (interquartile range) of the box limits, respectively.
Extended Data Fig. 7 Composite model performance across patients, key patient subgroups, the number of symptomatic irAEs per patient, and organ system involvement.
a, Same as Fig. 4d, but applied to both bulk cohorts (n = 53 patients) using leave-one-out cross-validation (LOOCV) (Methods). b, Same as Fig. 4c, but shown for model scores determined by LOOCV. c, Performance of the composite model versus other candidate pretreatment factors for predicting severe irAE development (Methods). The composite model was trained in bulk cohort 1 (BC1) and validated in bulk cohort 2 (BC2) or vice versa, as indicated. d, Performance of the composite model trained on bulk cohort 1 for predicting severe irAEs in different patient subgroups from bulk cohort 2. DCB, durable clinical benefit; NDB, no durable clinical benefit; GI, gastrointestinal. e, Composite model scores determined by LOOCV for all bulk cohort patients treated with combination therapy (n = 24), stratified by future irAE grade: 0/1 (n = 3), 2 (n = 7), 3 (n = 12), and 4 (n = 2). f, Model performance for predicting grade 2 + , 3 + , or 4 irAE development in combination therapy patients using the scores in e. g,h, Composite model scores determined by LOOCV in both bulk cohorts (n = 53 patients) versus the number of symptomatic irAEs (grade 2 + ) per patient (g) and the number of organ system toxicities per patient (h). i, Distribution of irAEs across patients and organ systems (Supplementary Table 15). Patients from bulk cohorts 1 and 2 are organized by decreasing composite model scores determined via LOOCV (Methods). The line distinguishing high/low scores was optimized using LOOCV (Methods). j, Fraction of patients in both bulk cohorts that developed irAEs in at least 2 organ systems versus those that did not, stratified by the threshold in panel i (Methods). Significance was determined by a two-sided Fisher’s exact test. In e, g, and h, center lines, bounds of the box, and whiskers indicate medians, 1st and 3rd quartiles, and minimum and maximum values within 1.5 × IQR (interquartile range) of the box limits, respectively. Statistical significance in e, g, and h was determined by a Kruskal-Wallis test.
Extended Data Fig. 8 Composite model performance for predicting time to severe irAE in validation bulk cohort 2.
a–c, Kaplan-Meier analysis for freedom from severe irAE in bulk cohort 2 for patients treated with combination or PD1 immune checkpoint blockade (a), combination therapy (b), or PD1 monotherapy (c), stratified by the composite model score (Methods). Statistical significance was calculated by a two-sided log-rank test. In all panels, training was performed in bulk cohort 1 and the cut-point predicting severe irAE was optimized for bulk cohort 1 using Youden’s J statistic (Supplementary Table 10; Methods). Notably, the analyses in a–c were landmarked between treatment initiation and three months following treatment initiation, with all severe irAEs occurring within this period. The Kaplan-Meier plots are shown out to four months given the extended follow-up of patients that did not develop any severe irAE (Supplementary Table 9).
Extended Data Fig. 9 Peripheral blood TCR-β profiling with immunoSEQ®.
a, Evenness (Pielou’s index) of TCR repertoires assembled by MiXCR (bulk RNA-seq) and immunoSEQ® (genomic DNA) from paired pretreatment PBMC samples (n = 15 combination therapy patients) (Supplementary Tables 1 and 18). Concordance and significance were determined by Spearman ρ and a two-sided t test, respectively. b, Similar to Fig. 5b but showing clonality for each pre- and on-treatment PBMC sample (Supplementary Table 18). Statistical significance was determined by a two-sided, paired Wilcoxon rank sum test. ns, not significant (P > 0.05). c, Fraction of pretreatment peripheral blood TCR clonotypes detected on-treatment in 15 combination therapy patients (Supplementary Table 18), stratified by no severe (n = 6) and severe (n = 9) irAE status. Clonotypes with matching productive CDR3 β-chain nucleotide sequences were considered identical. Center lines, bounds of the box, and whiskers indicate medians, 1st and 3rd quartiles, and minimum and maximum values, respectively. Significance was determined by a two-sided, unpaired Wilcoxon rank sum test. d–g, Clonal dynamics in circulating T cells following combination therapy initiation. d, Persistent T cell clones identified by immunoSEQ® were cross-referenced with scTCR-seq and scRNA-seq data of pretreatment PBMCs from the same three patients (YUALOE, YUNANCY, YUHONEY), all of whom received combination therapy and developed severe ICI-induced toxicity (Supplementary Table 18; Methods). e, Log2 expression of key lineage and activation markers across major T cell states annotated by Azimuth along with persistent clones classified into CD4 and CD8 T cells (Methods). f, Aggregate change from baseline in the productive frequencies of persistent clonotypes, stratified by lineage (n = 2 cell types) and patient (n = 3). The sum of the difference in productive frequencies (on-treatment % – pretreatment %) was calculated from immunoSEQ® data. Bars denote mean + /- SD. g, Top: Change in bulk TCR clonality from baseline (Fig. 5b). Bottom: Same as f but showing the underlying clonotypes, where circle size is proportional to pretreatment clone frequency (immunoSEQ®). h, Same as Fig. 5d but restricted to blood draws taken cycle 1 day 1 of combination therapy and <1 month later (n = 7 patients; Supplementary Table 18).
Extended Data Fig. 10 Schema of large-scale assessment of peripheral blood leukocytes in autoimmune disorders versus healthy controls.
Schema describing the workflow and statistical meta-analysis for evaluating the enrichment of individual circulating leukocyte subsets in autoimmune disorders relative to healthy controls (Fig. 6; Methods). In brief, CIBERSORTx was applied to enumerate 15 leukocyte subsets in bulk RNA-seq or microarray profiles of peripheral blood samples from patients with either systemic lupus erythematosus57,58,59 (SLE; n = 239) or inflammatory bowel disease56,60,61 (IBD; n = 348) compared to healthy controls (Supplementary Table 20). For each dataset and cell subset, a two-sided, unpaired Wilcoxon rank sum test was applied to assess the difference in relative abundance between healthy and disease phenotypes. Results were subsequently combined across studies by meta-z statistics (Methods).
Supplementary information
Supplementary Information
Supplementary Figs. 1–3
Supplementary Table
Supplementary Tables 1–20
Rights and permissions
About this article
Cite this article
Lozano, A.X., Chaudhuri, A.A., Nene, A. et al. T cell characteristics associated with toxicity to immune checkpoint blockade in patients with melanoma. Nat Med 28, 353–362 (2022). https://doi.org/10.1038/s41591-021-01623-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-021-01623-z
This article is cited by
-
S100A9+CD14+ monocytes contribute to anti-PD-1 immunotherapy resistance in advanced hepatocellular carcinoma by attenuating T cell-mediated antitumor function
Journal of Experimental & Clinical Cancer Research (2024)
-
The predictive value of peripheral blood CD4 cells ATP concentration for immune-related adverse events in advanced non-small cell lung cancer patients
BMC Immunology (2024)
-
Computational immunogenomic approaches to predict response to cancer immunotherapies
Nature Reviews Clinical Oncology (2024)
-
Clinical and translational attributes of immune-related adverse events
Nature Cancer (2024)
-
Comparing anti-tumor and anti-self immunity in a patient with melanoma receiving immune checkpoint blockade
Journal of Translational Medicine (2024)