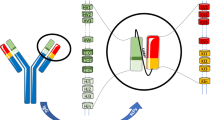
Abstract
Serum antibodies can recognize both pathogens and commensal gut microbiota. However, our current understanding of antibody repertoires is largely based on DNA sequencing of the corresponding B-cell receptor genes, and actual bacterial antigen targets remain incompletely characterized. Here we have profiled the serum antibody responses of 997 healthy individuals against 244,000 rationally selected peptide antigens derived from gut microbiota and pathogenic and probiotic bacteria. Leveraging phage immunoprecipitation sequencing (PhIP-Seq) based on phage-displayed synthetic oligo libraries, we detect a wide breadth of individual-specific as well as shared antibody responses against microbiota that associate with age and gender. We also demonstrate that these antibody epitope repertoires are more longitudinally stable than gut microbiome species abundances. Serum samples of more than 200 individuals collected five years apart could be accurately matched and could serve as an immunologic fingerprint. Overall, our results suggest that systemic antibody responses provide a non-redundant layer of information about microbiota beyond gut microbial species composition.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data generated or analyzed during this study are included within the paper, its Supplementary Information files and public repositories. Detailed information on the cohort, library content and PhIP-Seq data are available in the Supplementary Data files (files: cohort_info.csv, MB_composition.csv, library_content_info.csv and PhIP-Seq_data.zip). Patient-related data not included in the paper may be subject to patient confidentiality. Extended Data Fig. 3, Fig. 1, Extended Data Fig. 5, Fig. 3a,b/Extended Data Fig. 6 and Fig. 4a,c have associated raw data provided, respectively, in Supplementary Table 2, Supplementary Table 3, Supplementary Table 4, Supplementary Table 5 and Supplementary Table 6. Raw data for the PhIP-Seq experiments are deposited in the Harvard Dataverse public repository at https://doi.org/10.7910/DVN/3SOZCQ. Antigens included in the PhIP-Seq library were obtained from the immune epitope database (IEDB, https://www.iedb.org/) and virulence factor database (VFDB, http://www.mgc.ac.cn/VFs/), as well as other sources outlined in the Methods.
Code availability
Custom code used for analyzing the PhIP-Seq data is publicly available at https://github.com/erans99/PhIPSeq_external. The code repository is subdivided into two subfolders: (1) Analyse_Fastq, code to analyze a NextGen Sequencing plate, containing 96 wells, of which 80 are data wells and 16 are different types of controls of well quality (four negative controls, eight mocks and four positive control (‘anchor’) samples). The output of this is a file, per data well, of fold change and −log10(P value); (2) Analysis, code for executing different tests and analyses on the results of the PhIP-Seq output (as cached from files like those in the PhIPSeq_data directory).
References
Sender, R., Fuchs, S. & Milo, R. Are we really vastly outnumbered? Revisiting the ratio of bacterial to host cells in humans. Cell 164, 337–340 (2016).
Gilbert, J. A. et al. Current understanding of the human microbiome. Nat. Med. 24, 392–400 (2018).
Levy, M., Kolodziejczyk, A. A., Thaiss, C. A. & Elinav, E. Dysbiosis and the immune system. Nat. Rev. Immunol. 17, 219–232 (2017).
Bunker, J. J. & Bendelac, A. IgA responses to microbiota. Immunity 49, 211–224 (2018).
Koch, M. A. et al. Maternal IgG and IgA antibodies dampen mucosal T helper cell responses in early life. Cell 165, 827–841 (2016).
Gomez de Agüero, M. et al. The maternal microbiota drives early postnatal innate immune development. Science 351, 1296–1302 (2016).
Zeng, M. Y. et al. Gut microbiota-induced immunoglobulin G controls systemic infection by symbiotic bacteria and pathogens. Immunity 44, 647–658 (2016).
Wilmore, J. R. et al. Commensal microbes induce serum IgA responses that protect against polymicrobial sepsis. Cell Host Microbe 0, 1–10 (2018).
Fadlallah, J. Synergistic convergence of microbiota-specific systemic IgG and secretory IgA. J. Allergy Clin. Immunol. 143, 1575–1585 (2019).
Li, H. et al. Mucosal or systemic microbiota exposures shape the B cell repertoire. Nature 584, 274–278 (2020).
Sterlin, D., Fadlallah, J., Slack, E. & Gorochov, G. The antibody/microbiota interface in health and disease. Mucosal Immunol. 13, 3–11 (2020).
Soto, C. et al. High frequency of shared clonotypes in human B cell receptor repertoires. Nature 566, 398–402 (2019).
Briney, B., Inderbitzin, A., Joyce, C. & Burton, D. R. Commonality despite exceptional diversity in the baseline human antibody repertoire. Nature 566, 393–397 (2019).
Lindner, C. et al. Diversification of memory B cells drives the continuous adaptation of secretory antibodies to gut microbiota. Nat. Immunol. 16, 880–888 (2015).
Bashford-Rogers, R. J. M. et al. Analysis of the B cell receptor repertoire in six immune-mediated diseases. Nature 574, 122–126 (2019).
Meng, W. et al. An atlas of B-cell clonal distribution in the human body. Nat. Biotechnol. 35, 879–884 (2017).
Pasolli, E. et al. Extensive unexplored human microbiome diversity revealed by over 150,000 genomes from metagenomes spanning age, geography and lifestyle. Cell 176, 649–662 (2019).
Moor, K. et al. Analysis of bacterial-surface-specific antibodies in body fluids using bacterial flow cytometry. Nat. Protoc. 11, 1531–1553 (2016).
Palm, N. W. et al. Immunoglobulin A coating identifies colitogenic bacteria in inflammatory bowel disease. Cell 158, 1000–1010 (2014).
Bunker, J. J. et al. Innate and adaptive humoral responses coat distinct commensal bacteria with immunoglobulin A. Immunity 43, 541–553 (2015).
Mohan, D. et al. PhIP-Seq characterization of serum antibodies using oligonucleotide-encoded peptidomes. Nat. Protoc. 13, 1958–1978 (2018).
Larman, H. B. et al. Autoantigen discovery with a synthetic human peptidome. Nat. Biotechnol. 29, 535–541 (2011).
Larman, H. B. et al. PhIP-Seq characterization of autoantibodies from patients with multiple sclerosis, type 1 diabetes and rheumatoid arthritis. J. Autoimmun. 43, 1–9 (2013).
Vazquez, S. E. et al. Identification of novel, clinically correlated autoantigens in the monogenic autoimmune syndrome APS1 by proteome-wide PhIP-Seq. eLife 9, e55053 (2020).
Xu, G. J. et al. Viral immunology. Comprehensive serological profiling of human populations using a synthetic human virome. Science 348, aaa0698 (2015).
Mina, M. J. et al. Measles virus infection diminishes preexisting antibodies that offer protection from other pathogens. Science 366, 599–606 (2019).
Shrock, E. et al. Viral epitope profiling of COVID-19 patients reveals cross-reactivity and correlates of severity. Science 370, 1–23 (2020).
Zeevi, D. et al. Personalized nutrition by prediction of glycemic responses. Cell 163, 1079–1094 (2015).
Chen, L., Zheng, D., Liu, B., Yang, J. & Jin, Q. VFDB 2016: hierarchical and refined dataset for big data analysis—10 years on. Nucleic Acids Res. 44, D694–D697 (2016).
Vita, R. et al. The immune epitope database (IEDB) 3.0. Nucleic Acids Res. 43, D405–D412 (2015).
Lebeer, S. et al. Identification of probiotic effector molecules: present state and future perspectives. Curr. Opin. Biotechnol. 49, 217–223 (2018).
Bunker, J. J. et al. B cell superantigens in the human intestinal microbiota. Sci. Transl. Med. 11, eaau9356 (2019).
Ultsch, M., Braisted, A., Maun, H. R. & Eigenbrot, C. 3-2-1: structural insights from stepwise shrinkage of a three-helix Fc-binding domain to a single helix. Protein Eng. Des. Sel. 30, 619–625 (2017).
Korem, T. et al. Bread affects clinical parameters and induces gut microbiome-associated personal glycemic responses. Cell Metab. 25, 1243–1253 (2017).
Mattock, E. & Blocker, A. J. How do the virulence factors of Shigella work together to cause disease? Front. Cell. Infect. Microbiol. 7, 1–24 (2017).
Klotz, C., Goh, Y. J., O’Flaherty, S. & Barrangou, R. S-layer associated proteins contribute to the adhesive and immunomodulatory properties of Lactobacillus acidophilus NCFM. BMC Microbiol. 20, 248 (2020).
Chen, T. & Guestrin, C. XGBoost: a scalable tree boosting system. In Proc. 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining 785–794 (ACM, 2016); https://doi.org/10.1145/2939672.2939785
Landsverk, O. J. B. et al. Antibody-secreting plasma cells persist for decades in human intestine. J. Exp. Med. 214, 309–317 (2017).
Magri, G. et al. Human secretory IgM emerges from plasma cells clonally related to gut memory B cells and targets highly diverse commensals. Immunity 47, 118–134 (2017).
Chen, K., Magri, G., Grasset, E. K. & Cerutti, A. Rethinking mucosal antibody responses: IgM, IgG and IgD join IgA. Nat. Rev. Immunol. 20, 427–441 (2020).
Wilms, E. et al. Intestinal barrier function is maintained with aging—a comprehensive study in healthy subjects and irritable bowel syndrome patients. Sci. Rep. 10, 475 (2020).
Thevaranjan, N. et al. Age-associated microbial dysbiosis promotes intestinal permeability, systemic inflammation and macrophage dysfunction. Cell Host Microbe 21, 455–466 (2017).
Cohen, D. et al. Recent trends in the epidemiology of shigellosis in Israel. Epidemiol. Infect. 142, 2583–2594 (2014).
McCoy, K. D., Burkhard, R. & Geuking, M. B. The microbiome and immune memory formation. Immunol. Cell Biol. 97, 625–635 (2019).
Xu, G. J. et al. Systematic autoantigen analysis identifies a distinct subtype of scleroderma with coincident cancer. Proc. Natl Acad. Sci. USA 113, E7526–E7534 (2016).
Paull, M. L. & Daugherty, P. S. Mapping serum antibody repertoires using peptide libraries. Curr. Opin. Chem. Eng. 19, 21–26 (2018).
Puga, I. et al. B cell-helper neutrophils stimulate the diversification and production of immunoglobulin in the marginal zone of the spleen. Nat. Immunol. 13, 170–180 (2012).
Setliff, I. et al. High-throughput mapping of B cell receptor sequences to antigen specificity. Cell 179, 1636–1646 (2019).
Spitzer, M., Wildenhain, J., Rappsilber, J. & Tyers, M. BoxPlotR: a web tool for generation of box plots. Nat. Methods 11, 121–122 (2014).
Segata, N. et al. Metagenomic microbial community profiling using unique clade-specific marker genes. Nat. Methods 9, 811–814 (2012).
Forsström, B. et al. Dissecting antibodies with regards to linear and conformational epitopes. PLoS ONE 10, e0121673 (2015).
Berglund, L., Andrade, J., Odeberg, J. & Uhlén, M. The epitope space of the human proteome. Protein Sci. 17, 606–613 (2008).
Forsström, B. et al. Proteome-wide epitope mapping of antibodies using ultra-dense peptide arrays. Mol. Cell. Proteom. 13, 1585–1597 (2014).
Li, J. et al. An integrated catalog of reference genes in the human gut microbiome. Nat. Biotechnol. 32, 834–841 (2014).
Rothschild, D. et al. Environment dominates over host genetics in shaping human gut microbiota. Nature 555, 210–215 (2018).
Zeevi, D. et al. Structural variation in the gut microbiome associates with host health. Nature 568, 43–48 (2019).
Truong, D. T. et al. MetaPhlAn2 for enhanced metagenomic taxonomic profiling. Nat. Methods 12, 902–903 (2015).
Babu, M. et al. Global landscape of cell envelope protein complexes in Escherichia coli. Nat. Biotechnol. 36, 103–112 (2018).
Götz, S. et al. High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res. 36, 3420–3435 (2008).
Rothschild, D. et al. An atlas of robust microbiome associations with phenotypic traits based on large-scale cohorts from two continents. Preprint at bioRxiv https://doi.org/10.1101/2020.05.28.122325 (2020).
Wozniak, J. M. et al. Mortality risk profiling of Staphylococcus aureus bacteremia by multi-omic serum analysis reveals early predictive and pathogenic signatures. Cell 182, 1311–1327 (2020).
Acknowledgements
A.W. is the Louis H. Sackin Research Fellow Chair in Computer Science. E.S. is supported by grants from the European Research Council, the Israel Science Foundation and by the Seerave Foundation. T.V. gratefully acknowledges support from the Austrian Science Fund (FWF, Erwin Schrödinger fellowship J4256). R.K.W. is supported by the Seerave Foundation and the Netherlands Organization for Scientific Research. A.Z. is supported by ERC Starting Grant 715772, the Netherlands Organization for Scientific Research NWO-VIDI grant 016.178.056, the Netherlands Heart Foundation CVON grant 2018-27 and the NWO Gravitation grant ExposomeNL 024.004.017. J.F. is supported by NWO Gravitation Netherlands Organ-on-Chip Initiative (024.003.001), ERC Consolidator grant 101001678 and the Netherlands Heart Foundation CVON grant 2018-27. C.W. is supported by NWO Gravitation grant 024.003.001 and NWO Spinoza Prize SPI 92-266. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
T.V. and S.K. conceived the project and designed the library. S.L. designed and implemented the coding of the library. T.V. and S.K. planned and calibrated the experimental system and performed biological experiments. S.L. designed and implemented the computational pipeline. S.L. and I.N.K. performed high-throughput data analysis; T.V. analyzed additional data and wrote the manuscript. E.S. and A.W. conceived and directed the project. T.V., S.K., S.L., I.N.K., A.W., C.W., J.F., A.Z., R.K.W. and E.S. reviewed and edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Medicine thanks Rachael Bashford-Rogers, George Georgiou, Andrea Cerutti and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Saheli Sadanand was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Control experiments of optimizing the ratio of phage/antibody amounts in IPs (a), the reproducibility of technical duplicates (b), examples of high technical (c) as well as biological reproducibility (d,e), and the > 1,000 samples reported in the main manuscript were processed in batches of 96 well plates, that were not biased by batch effects (f).
a, ‘Phages per variant’ refers to the number of phages per library variant, ‘Phages per reaction’ refers to the number of total phages in a reaction mixture of the microbiota library (244,000 variants times the number of phages per variant). IP reactions were performed in duplicates (R1, R2), the numbers of significantly bound peptides are shown normalized as percent of the highest binding phage/antibody combination (4,000-fold phage coverage and 4 µg of IgG antibodies). A mixed pool of human serum samples was used as antibody material for this calibration. b, Technical replicates (n = 191 samples measured in duplicates) were in excellent agreement with an average Pearson R² (of FCs) of 0.96 between duplicates. 95% of duplicates correlated with R² greater than 0.90 (181/191) and 78% of duplicates even with an R² greater than 0.95 (149/191). Given this high reproducibility and little added information gained from duplicates, the exploratory experiments reported in this manuscript were carried out in single reactions. For potential diagnostic applications of PhIP-Seq technical replicates may be valuable to validate results. c-e, Examples of high technical reproducibility and low background binding (e) as well as biological reproducibility of samples collected 6 days (f) and 1.5 years apart (g). In red in panel e low background binding of a negative control without antibodies (‘Mock IP’21) is illustrated. Samples collected days (f) or years (g) apart and processed in different PhIP-Seq runs show excellent reproducibility. f, Principle component analysis (PCA) of samples measured in different batches of PhIP-Seq experiments. PCs were computed on signals (log FC) against bound peptides of the entire antigen, the first six PCs are shown. Samples measured in the same batch do not cluster separately from other batches indicating no clear bias of batch effects for these samples.
Extended Data Fig. 2 Analysis of negative controls for estimating nonspecific background signal (a) and comparison of viral positive controls of this study to population wide responses previously reported25 (b).
a, Negative controls indicate little nonspecific background-binding impacting population-scale interpretation of the measured antibody epitope repertoires. We had included negative controls of proteins that were expected to elicit little binding in healthy individuals. These included random amino acid sequences (100 peptides), as well as human proteins (autoimmune disease targets reported in the IEDB30 and various abundant housekeeping genes such as histones and glycolytic enzymes represented as 364 peptides). Analyzing binding to these negative controls in the cohort showed that a few random peptides were significantly enriched in up to 0.5% (5/997 individuals), indicating a low background of unspecific binding (or cross-reactivity) which can be eliminated by using a threshold for peptides bound in >1% of individuals. Peptides of human proteins were bound in up to 3.3% (33/997) of individuals, similar to results previously reported using PhIP-Seq23. It has been speculated that such antibody binding against human proteins may arise from cross-reactivity and are unlikely to have detrimental consequences in healthy individuals23. The following machine learning based predictions in this work were limited to peptides bound in at least 2% of the population. b, Controls of viral epitopes measured on our cohort match previously reported seroprevalences from a population scale study25. Xu et al25. employed a PhIP-Seq workflow with a library covering viral antigens (‘VirScan’) and detected near universal population wide targeting of certain viral peptides. They had reported a list of 11 viral peptides including the amino acid sequences and seroprevalences (supporting information, Table S2 of their publication25, % seroprevalences are reproduced from this table). We had included the exact same peptides and detected similar rates of seroprevalence, demonstrating the reproducibility of the PhIP-Seq workflow and sensitivity of our implementation. *: Xu et al. have analyzed in total sera of 569 individuals, although the exact number of individuals for calculating the seroprevalence is not specified in the caption of their supporting table S2.
Extended Data Fig. 3 The PhIP-Seq workflow robustly identifies peptide targets of antibodies generated against immunogens of full-length proteins and bacterial cells.
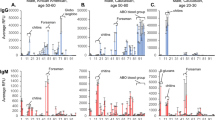
a-i, Commercially available antibodies were measured with our PhIP-Seq microbiota library following the same standard approach applied to human serum samples. Antibody amounts were normalized by the concentrations specified by the manufacturers. Panels a-g represent antibody preparations targeting microbial antigens, panels h and i represent negative controls of monoclonal antibodies targeting human proteins. See the first sheet of supporting file.xlsx file Supplementary table 2 for details on immunogens and properties related to each antibody. Measurements of each antibody were performed in triplicates and peptides appearing in all replicates were used in the analysis. The correlation of Fold change values for two random replicates [Rep. 1, Rep. 2] are shown (Pearson R²). ‘Fold change’ refers to the ratio between reads in the IP reaction with antibodies vs. input sequencing of the phage library (a proxy for binding strength). Note the different scales on the axes and note the use of a logarithmic scale for the axes of panel f (to adequately represent weakly enriched peptides). ‘*’ in panel c denotes an antibody preparation, that was protein A purified according to the manufacturer. The exact bound peptides are listed in the second sheet of supporting file.xlsx file Supplementary table 2. Only peptides related to the bound antigens are listed (background reactivity of the whole sera from rabbit/goat omitted). j, Assessment of potential cross-reactivity or background reactivity of the antibody preparations. The list of bound peptides by each antibody preparation (marked in panels a-i) was searched among the bound peptides by every other antibody preparation (and is marked in all plots, only nearly identical E. coli and Shigella peptides show up in the other sample as well). The numbers of bound peptides are listed. Thereby we have verified that the marked peptides in panels a-i are not appearing due to background/cross-reactivity of the whole animal sera, as they only appear in reactions of the respective antibodies.
Extended Data Fig. 4 Bacterial strains of different functional groups within the library (Fig. 1b, methods section and Supplementary table 1) all elicit substantial population wide antibody responses.
The fraction of peptides per strain (out of all the strain’s peptides) bound in >3% of the cohort (n = 997) is shown. See Supplementary table 1 for details on the bacterial strains listed. Antibody responses are not limited to pathogenic strains, but extend to strains selected from healthy individuals’ gut microbiota (from metagenomics sequencing, see the methods section), probiotic strains, and strains previously reported to be coated by antibodies19. A large fraction of Staphylococcus aureus peptides were bound, possibly owing to its ubiquitous role in the upper respiratory tract and human skin microbiome along its large number of virulence factors potentially eliciting antibody responses29,61.
Extended Data Fig. 5 Antibody binding with protein A and protein G coated beads separately (a-c) and antibody coated beads capturing IgA and IgG separately (d).
Supplementary table 4 provides detailed lists on the respective peptides bound. a-c Relying on different binding affinities of protein A and G for antibody classes, we processed 80 serum samples each with 1.) a mixture of protein A and G, 2.) protein A alone, and 3.) protein G alone. a, Comparison of peptides bound by protein A vs. protein G. b, Comparison of peptides bound by a mixture of protein A and G vs. protein G. c, Comparison of peptides bound by a mixture of protein A and G vs. protein A. In panels a-c data of 78/80 samples are shown, as samples with <200 significantly enriched peptides per sample were excluded (same cutoff as for the other human sera measured). d Experimental workflow to detect IgA and IgG subclasses separately (following procedures reported in the literature27 and Methods). In panel d, a comparison of peptides bound by IgA vs. IgG specific beads is shown. Samples with IgG specific beads were sequences with 0.8 million reads, however we do not expect a strong impact thereof, as the number of detected peptides typically saturates22. e Comparison of peptides bound by a mixture of protein A and G vs. IgG specific beads. f Comparison of peptides bound by a mixture of protein A and G vs. IgA specific beads. In panels d-f data of 80 samples is shown (as for IgA many samples would not have passed the threshold of >200 peptides applied in other figures, see panel g). For the IgA vs. IgG experiments a different batch of phages was used. In panels a-f Pearson R² is shown. g Number of bound peptides per sample with each set of magnetic beads used. Center lines show the medians; box limits indicate the 25th and 75th percentiles as determined by R software; whiskers extend 1.5 times the interquartile range from the 25th and 75th percentiles, outliers are represented by dots. n = 80 sample points.
Extended Data Fig. 6 Associations between serum antibody responses and abundances inferred from metagenomics sequencing.
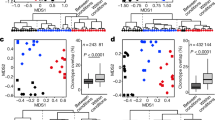
a, Testing antibody bound peptides which appeared in >2% of individuals (4469 peptides) vs. relative abundances of species (SGBs50) which appear in >2% of individuals (1056 SGBs). Of them, 1706 pairs (listed in Supplementary table 5) passed FDR correction (p-value <0.05) for multiple hypothesis testing (approximately 4.7 million tests). Most of these associations were from peptides and species that appeared in a small percentage (2-5%) of individuals. We also performed the same test of peptides and species which appear in >5% of individual (745 species and 1566 peptides) with 12 pairs passing FDR correction. Some of the species abundances are correlated with the fold change of up to 23 peptides per species (histogram in Fig. 3b). This analysis includes also associations of multiple SGBs with the same peptide. For example antibody binding of the Shigella IpaC protein (antibody binding against which we had found to be associated with age [Fig. 4c]) was associated with abundances of various SGBs, suggesting multiple factors contributing to its biological effects (for example potentially increased translocation as well as effects mediated by the adaptive immune system). These results are affected by detection thresholds of PhIP-Seq and metagenomics sequencing and we cannot rule out that small amounts of bacteria undetectable in microbiome sequencing eliciting weak antibody responses would associate in a larger fraction of individuals. Another technical consideration beyond the detection threshold is the library content size, with the option of creating PhIP-Seq antigen libraries specific to individuals potentially allowing to capture links between metagenomics data and antibody epitope repertoires at greater depth. b, Representation of the 1,706 significant population scale associations between antibody binding against peptides (x-axis) and detection in metagenomics sequencing (y-axis). Every dot represents one of the significant associations listed in (Supplementary table 5). Each dot is colored by the FDR-corrected p-value of the Spearman correlation (also listed in Supplementary table 5). c,d Correlation of every person’s metagenomics gut microbiome sequencing data with the antibody repertoire data (similar to Fig. 6a,b) on gut metagenomics antigens/genes (methods section).
Extended Data Fig. 7 Peptide ELISAs validations of PhIP-Seq results.
Sections of 20 amino acids (aa) of six peptides included within our PhIP-Seq library were chemically synthesized and tested in a peptide ELISA against sera of 80 individuals, for whom also PhIP-Seq data was available (see M&Ms section “Peptide ELISAs” for the selection criteria and sequences of the peptides). a-f, Comparison of peptide ELISA and PhIP-Seq data for each peptide (as indicated by the title above each panel). Each dot represents data of one individual. Absorption values of ELISA data and p-values of significance of enrichment of binding in PhIP-Seq (for one peptide) are shown on the x and y axes respectively. Absorption values below the average of the negative control were normalized to 0. Spearman correlation (R) with associated p-value (Spearman rank-order correlation coefficient, nonparametric measure) was computed for each pair of PhIP-Seq and ELISA data (shown in each panel). The negative control peptides were not bound in PhIP-Seq, hence Spearman R/p-val are not applicable (n.a.). g, Summary of the results shown in panels a-f. The percentage of ELISA or PhIP-Seq binding in the 80 individuals was calculated for each peptide with the standard Generalized Poisson cutoffs for PhIP-Seq (with binding of multiple peptides summarized if applicable, see text below) and the ELISA data was counted as positive when the absorption value was greater than the average of the negative control. The calculated Spearman correlation (R) between the frequency of antibody responses in PhIP-Seq and ELISA is shown in the panel.
Extended Data Fig. 8 Antibody epitope repertoires against the microbiota antigen library significantly predict C-reactive protein (CRP) levels (measured with a wide CRP range test) by machine learning, albeit with lower predictive power than age (Fig. 5a) or gender (Fig. 5b).
a, Age and gender alone are confounding factors of machine learning based predictions of serum CRP levels (measured with a wide range test for ca. 400 individuals). As antibody epitope repertoires also carry a wealth of age/gender related information (Fig. 5) the contribution of age and gender alone vs. antibody epitope repertoires was assessed here. CRP levels were predicted using age and gender alone as features and with a combination of age, gender, and microbiota antibody epitope repertoires as features, using Ridge Regression 10-fold cross validation. Each model was repeated 100 times (different cross validation sets) and a histogram of the resulting 100 Pearson correlation coefficients (correlation of actual vs. predicted CRP values) are shown in panel a. The analysis has been corrected for multiple hypothesis testing: Pearson correlation of predicted (on antibody bound peptides + age + gender) to actual value is 0.12, with p-value of 0.011, which after FDR correction becomes 0.018 (<0.05, i.r. passes FDR correction). Pearson correlation of machine learning based prediction on age & gender alone to actual value is 0, so that all predictive power comes directly from antibody bound peptides, and not from their prediction of age and gender. Thereby a significant added predictive value of microbiota antibody epitope repertoires is demonstrated. b, For both 1.) other blood tests beyond CRP or 2.) anthropometrics such as body mass index (BMI), adding microbiota antibody epitope repertoire information rather worsens machine learning based predictions compared to age and gender alone (as additional meaningless features increase noise, see methods section) or did not pass FDR correction. This notion is exemplified with machine learning based prediction of BMI in panel b.
Extended Data Fig. 9 In contrast to antibody responses targeting microbiota antigens, reactivities against human self-proteins and random peptides carry virtually no predictive power by machine learning for age (a) and gender (b).
This finding precludes that self-reactivity or potential cross-reactivity against random peptides underlie the strong associations observed. The machine learning based predictions based on human proteins and random peptides encompassed ca. 6,300 peptides included in the antigen library as part of the IEDB (autoantigens) or as controls (covering abundant proteins such histones, glycolytic enzymes etc., see the methods section and Supplementary table 1 for details). Average and standard deviation derived by 10 repeats of XGBoost with 10-fold cross validation (as in Fig. 5). Ideally, the same number of controls as microbial peptides should have been used. However, that would have doubled the cost of the PhIP-Seq library as well as doubling the cost of every assay performed (as we would have had to sequence deeper and use more beads to retain the same signal strength). Given these cost considerations, we could not afford to include a set of nearly 250,000 controls. However, we believe that also the set of only 6,300 peptides serves as an important control: We detected very little binding against these controls (discussed in more detail in S2/S3), and they do not carry any predictive power by machine learning, demonstrating that there is no exceedingly large cross-reactivity or background signal with our PhIP-Seq system.
Extended Data Fig. 10 Additional analyses of longitudinal stability.
Complete correlation diagrams for the five-year longitudinal antibody stability results of 213 individuals shown in Fig. 6 (panels a, c, e) and additional subgroups of the antigen library (c,d) as well as two different approaches to assess gut microbiome stability from metagenomics sequencing data (g,h). Pearson correlations of log fold changes of all baseline (t = 0) and follow up (t = 5 years) samples compared with each other are shown. Correlations based on antigens of the entire microbiota library (a) [also shown in Fig. 6a], only the VFDB (b), and the microbiota library excluding VDFB (c) are shown. In addition to these antigens obtained from databases, two analyses with antigens from microbiome sequencing of this cohort28 (Methods) were performed (d,e). f, Summary figure on the correlation coefficients of the stability of antibody epitope repertoires from antigen subgroups shown in panel a to e of this figure, comparing correlation of random pairs of samples and pairs of matched individual’s samples collected five years apart. Mean values and standard deviations of n = 213. Sample sizes: random pairs of samples: 213²-213 comparisons; individuals’ matched samples: 213 comparisons (see Fig. 6b,c for details). Antigen groups sizes for panels a-f: All microbiota – 231,975 peptides, VFDB – 24,164 peptides, Library excluding VFDB – 207,811 peptides, Metagenomics antigens - 147,061 peptides. g,h Gut microbiome stability inferred from metagenomics sequencing of stool samples collected five years apart of 188 individuals. In panel g, stability is calculated from gene abundances. The Bray Curtis distances for all baseline (t = 0) and follow up (t = 5 years) samples compared with each other are shown (the higher the value, the closer the samples resemble). In panel h, stability is calculated based on presence/absence (existence) of genes appearing in individuals (by applying a cutoff threshold to the gene abundances). The Normalized Hamming distances for all baseline (t = 0) and follow up (t = 5 years) samples compared with each other are shown (the higher the value, the closer the samples resemble).
Supplementary information
Supplementary Information
Supplementary Figs. 1–4, Supplementary Table 1 and legends for Supplementary Tables 2–6.
Supplementary Data 1
Supplementary Tables 2–6.
Supplementary Data 2
Cohort info.
Supplementary Data 3
Library content info.
Supplementary Data 4
MB composition.
Supplementary Data 5
PhIP-Seq data.
Rights and permissions
About this article
Cite this article
Vogl, T., Klompus, S., Leviatan, S. et al. Population-wide diversity and stability of serum antibody epitope repertoires against human microbiota. Nat Med 27, 1442–1450 (2021). https://doi.org/10.1038/s41591-021-01409-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-021-01409-3
This article is cited by
-
Efficient encoding of large antigenic spaces by epitope prioritization with Dolphyn
Nature Communications (2024)
-
Detecting antibody reactivities in Phage ImmunoPrecipitation Sequencing data
BMC Genomics (2022)