Abstract
A key unsolved question in the current coronavirus disease 2019 (COVID-19) pandemic is the duration of acquired immunity. Insights from infections with the four seasonal human coronaviruses might reveal common characteristics applicable to all human coronaviruses. We monitored healthy individuals for more than 35 years and determined that reinfection with the same seasonal coronavirus occurred frequently at 12 months after infection.
Similar content being viewed by others
Main
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is a novel coronavirus responsible for an ongoing pandemic. To date, there is limited evidence of reinfection by SARS-CoV-2, although it is generally assumed that reinfections by coronaviruses occur. To prepare for future waves of Coronavirus Disease 2019 (COVID-19), it is important to elucidate the duration of protection to reinfection for which the seasonal coronaviruses might serve as an informative model. There are four species of seasonal coronaviruses—HCoV-NL63, HCoV-229E, HCoV-OC43 and HCoV-HKU1—that all can cause respiratory tract infections but are otherwise genetically and biologically dissimilar. They belong to two distinct taxonomic genera and use different receptor molecules with varying host cell tropism1. Given this large variation, we hypothesize that characteristics shared by these four seasonal coronaviruses, such as the duration of protective immunity, are representative of all human coronaviruses, including SARS-CoV-2. Therefore, the aim of this study was to investigate the duration of protection from seasonal coronavirus reinfections.
Because seasonal coronavirus infections can be asymptomatic, and the duration of viral shedding is limited2, long-term epidemiological studies based on virus detection methods to identify infections (for example, reverse transcriptase-polymerase chain reaction (RT–PCR)) require continuous sampling of respiratory samples, which is not feasible. Alternatively, serological assays can be used, as antibody levels remain elevated up to 1 year after infection2. We, therefore, used serum samples from the Amsterdam Cohort Studies on HIV-1 infection and AIDS, a cohort study following adult males at regular intervals since the 1980s, to investigate how often seasonal coronavirus infections occurred during follow-up. Ten healthy individuals, who did not report any serious illness that could have influenced their immunity, were selected (Methods and ref. 3). Apart from a gap in follow-up between 1997 and 2003, blood collection occurred every 3 months before 1989 and every 6 months afterwards. The cumulative period at which the ten individuals were continuously followed totaled more than 2,473 months.
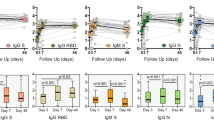
To detect reinfections, we measured increases in antibodies to the carboxyl (C)-terminal region of the nucleocapsid protein (NCt)—an immunodominant region of the structural coronavirus capsid protein4—for each seasonal coronavirus. The choice of the antigen, the serological test, the threshold for infection and the specificity and sensitivity of the tests are supplied in the Methods, Extended Data Figs. 1–4 and Supplementary Tables 1–3. A total of 101 events, ranging from 3 to 17 per individual, were classified as coronavirus infections (Table 1 and Fig. 1a). The time to reinfection was calculated only during continuous follow-up periods (connected dots in Fig. 1a and Supplementary Fig 1). The reinfection times ranged between 6 and 105 months (Fig. 1b). There was no statistically significant difference between the infection interval lengths of the individual viruses (Kruskal–Wallis test, P = 0.256), even though the number of HCoV-HKU1 infections was low and likely underestimated, most probably because of the low sensitivity of the HKU1-NCt-ELISA (Extended Data Fig. 2).
a, Example of antibody dynamics of one individual (#9). Connected dots: follow-up intervals <400 d; asterisks: visits with a ≥ 1.40 ELISA OD fold rise in antibodies with the value of the fold change adjacent to the asterisk; black font and underlined: OD fold rises of ≥1.40 classified as infections; gray font: OD fold rises of ≥1.40 suspected of cross-reactivity and not classified as infections. b, The interval between reinfections determined for the ten individuals. White dots: reinfections without an intermediate decrease in antibody levels; black vertical lines: median reinfection times. c, Changes in antibody levels after infection relative to the follow-up interval duration. Each circle represents an infection. Horizontal line: cutoff between increases (>1.0) or decreases (<1.0) in antibody levels at the next follow-up visit. d, The prevalence of infection of the four coronaviruses across different months. The prevalence per month is shown as a percentage of the total number of infections per coronavirus.
In a few cases, reinfections occurred as early as 6 months (twice with HCoV-229E and once with HCoV-OC43) and 9 months (once with HCoV-NL63), but reinfections were frequently observed at 12 months (Fig. 1b). For reinfections occurring as early as 6 months, we observed no intermediate reduction in antibodies between infections (Fig. 1b, white circles), but reinfection intervals of more than 6 months did show intermediate reductions between infections (visible as peaks in Fig. 1a and Supplementary Fig. 1). The ability to detect short-term reinfections in this study was limited by the sampling interval, which was, at minimum, 3 months. However, no signs of reinfection were observed at the first subsequent follow-up visit after a 3-month interval (Fig. 1b), as only decreases in antibody levels (optical density (OD) fold changes <1.0) were found (Fig. 1c). We, therefore, concluded that, in our data, the earliest time point for reinfection by seasonal coronaviruses was 6 months.
In theory, antibodies induced by coronavirus infections might have broad coronavirus-recognizing characteristics. To examine this, we performed an additional enzyme-linked immunosorbent assay (ELISA), this time using the complete nucleocapsid (N) protein of SARS-CoV-2, including the more inter-species-conserved N-terminal region to allow detection of broadly recognizing antibodies. Two individuals showed broadly recognizing antibodies (individuals #2 and #9; Extended Data Fig. 5). For individual #9, these antibodies remained present in the following years. The self-reported symptoms of this individual did not reveal any relevant medical history that might explain the induction and maintaining of these antibodies. Notably, these sustained broadly recognizing antibodies did not provide broad protection from subsequent HCoV-NL63, HCoV-229E and HCoV-OC43 infections.
Our serological study is unique because it avoids the sampling bias of previous epidemiologic studies based on symptoms-based testing protocols5. In our study, the months of June, July, August and September show the lowest prevalence of infections for all four seasonal coronaviruses (Fig. 1d; Wilcoxon signed-rank test, P = 0.004), confirming the higher prevalence in winter in temperate countries, and SARS-CoV-2 might share this feature in the post-pandemic era.
We were not able to identify strain variation, which could play a role in susceptibility to reinfection. HCoV-NL63, HCoV-OC43 and HCoV-HKU1 all show different co-circulating genetic clusters1. The situation is even more complicated for HCoV-229E, which shows continuous genetic drift. The sole two volunteer reinfection studies were performed with HCoV-229E. Callow et al. showed reinfection of six of nine volunteers with the same HCoV-229E isolate at a 12-month interval2. In contrast, a study by Reed showed no reinfection of volunteers when the same strain of HCoV-229E was used, whereas reinfection by heterologous strains was successful6. In the current SARS-CoV-2 pandemic, with only slightly varying circulating strains, increased susceptibility to reinfection by divergent SARS-CoV-2-strains is most likely not the case.
Caution should be taken when relying on policies that require long-term immunity, such as vaccination or natural infection to reach herd immunity. Other studies have shown that neutralizing SARS-CoV-2 antibody levels decrease within the first 2 months after infection, especially after mild COVID-197,8, and we observed a similar decrease in anti-nucleocapsid antibodies of seasonal coronaviruses (Extended Data Fig. 6). However, antibodies are only one marker for immunity, which is probably also influenced by B cell- and T cell-mediated immunity. In our study, we monitored reinfections, which can occur only when protective immunity (cellular and/or humoral) is insufficient. We show that reinfections by natural infection occur for all four seasonal coronaviruses, suggesting that it is a common feature for all human coronaviruses, including SARS-CoV-2. Reinfections occurred most frequently at 12 months after infection, indicating that protective immunity is only short-lived.
Methods
Serum samples
A total of 513 serum samples from the Amsterdam Cohort Studies on HIV infection and AIDS3 were examined. The Amsterdam Cohort Studies was initially started to investigate the prevalence, incidence and risk factors of HIV-1 infection. The study population consists of men who have sex with men who live mainly around the city of Amsterdam, the Netherlands. HIV-1 seronegative and HIV-1 seropositive men were enrolled; the individuals in our study were all from the HIV-1-negative arm of the study. The ten individuals were chosen based on more than 10 years of follow-up. All ten individuals remained negative for hepatitis C virus. Four of the ten individuals showed infection by hepatitis B virus (individual #2 before 1984; #8 in 1998; #5 and #10 in 1999). Herpes zoster virus infections were found in three individuals (#3 in 2004 and #9 and #10 in 1985). Whether the study individuals experienced acute infection by Epstein–Barr virus was not tested. One of the individuals (#2) had had insulin-dependent diabetes. No blood disease, cancer, autoimmune disease or neurodegenerative disease was reported during the follow-up period for #1, #2, #3, #4, #5, #8, #9 and #10. For individuals #6 and #7, these data were not available. Four individuals reported receiving a blood product in the 6 months before a visit (exact date or indication not known): #1 in 1991, #2 in 1994, #4 in 1987 and #5 in 1987. The gap in follow-up between 1997 and 2003 was based on limited transmission of HIV-1 in the 1990s among men having sex with men. When HIV-1 infections started re-appearing, the volunteers were asked to rejoin the Amsterdam Cohort Studies on HIV-1 infection and AIDS.
Self-reported symptoms of influenza-like illnesses (ILIs) were documented at each study visit (Supplementary Table 4). The Amsterdam Cohort Studies on HIV infection and AIDS was approved by the Medical Ethics Committee of the Amsterdam University Medical Center of the University of Amsterdam, the Netherlands (MEC 07/182). Participation is voluntary and without incentive. Written informed consent of each participant was obtained at enrollment.
To determine the sensitivity and specificity of the serological assay, serum samples of patients with RT-PCR-confirmed coronavirus infection were included in the study. The patients were recruited in 16 primary care networks9,10. Inclusion criteria for patients were as follows: age 18 years or older with an acute or worsened cough (≤28-d duration) as the main symptom or any clinical presentation considered by the general practitioner to be caused by a lower respiratory tract infection and consulting for the first time for this illness episode. Within 24 h of the first presentation, a serum sample and a nasopharyngeal flocked swab (COPAN) were collected (V1). At days 28–35 after V1, serum sampling was repeated (V2). Respiratory samples were tested for respiratory viruses, including the four seasonal coronaviruses11. The study was approved by the local ethics committees in all participating centers and by the competent authority in each country: Cardiff and Southampton (United Kingdom): Southampton & South West Hampshire Research Ethics Committee A; Utrecht (Netherlands) Medisch Ethische Toetsingscommissie Universitair Medisch Centrum Utrecht; Barcelona (Spain): Comitè ètic d’investigació clínica Hospital Clínic de Barcelona; Mataro (Spain): Comitè d’Ètica d’Investigació Clínica (CEIC) del Consorci Sanitari del Maresme; Rotenburg (Germany): Ethik-Kommission der Medizinischen Fakultät der Georg-August-Universität Göttingen; Antwerpen (Belgium): UZ Antwerpen Comité voor Medische Ethiek; Lodz, Szeczecin and Bialystok (Poland): Komisja Bioetyki Uniwersytetu Medycznego W Lodzi; Milano (Italy): IRCCS Fondazione Cà Granda Policlinico; Jonkoping (Sweden): Regionala etikprövningsnämnden i Linköping; Bratislava (Slovakia): Etika Komisia Bratislavskeho; Gent (Belgium): Ethisch Comité Universitair Ziekenhuis Gent; Nice (France): Comité de Protection des Personnes Sud-Méditerranée II, Hôpital Salvator; and Jesenice (Slovenia): Komisija Republike Slovenije za Medicinsko Etiko. Written informed consent was obtained from each individual before inclusion. Detailed information on experimental design and reagents is supplied in the Life Sciences Reporting Summary.
Choice of serological assay
An ELISA-based methodology was chosen for this study. Of note is that virus neutralization tests could not be used. There is no cell line facilitating replication of HCoV-HKU1, and the only available HCoV-229E and HCoV-OC43 cultured virus strains are from the 1960s and lab adapted, which might not be proper representatives of wild-type viruses. As a result, HCoV-NL63 (strain Amsterdam-1) is the only virus that can be used in neutralization tests. Because neutralization is a biological assay, each serum sample has to be tested at least twice with eight-fold serial dilutions to be able to calculate the 50% neutralization titer12. This requires a minimum of 300 µl of serum per time point, a volume that was not available for each individual in our study. For some time points of individuals #5 and #7, sufficient serum required was available and used for neutralization as a confirmation of the HCoV-NL63 ELISA (Extended Data Fig. 4).
Choice of antigen
In ELISA, a careful choice for the antigen has to be made, considering the tradeoff between sensitivity and cross-reactivity in a serological assay. Although the spike protein elicits neutralizing antibodies, it is the least conserved within a seasonal coronavirus species (Supplementary Table 2 and ref. 1). The N protein, and specifically its NCt, was chosen for this study because it is conserved within each species yet is the least conserved between species (Supplementary Table 1). The NCt protein has proven to be an immunogenic and sensitive protein to monitor seasonal coronavirus infections4,13,14,15,16,17,18.
HCoV-NL63, HCoV-229E, HCoV-OC43 and HCoV-HKU1 C-terminal N proteins (NCt) and SARS-CoV-2 complete N protein
The antigens for ELISA were produced and purified as described previously13. Strains Amsterdam-1 (NC_005831), Inf-1 (NC_002645), VR-759 (AY391777), Caen1 (HM034837) and Wuhan-Hu-1 (NC_045512.2) were the input virus strains to generate proteins of HCoV-NL63, HCoV-229E, HCoV-OC43, HCoV-HKU1 and SARS-CoV-2, respectively. The following primers were used to amplify the full N protein of SARS-CoV-2: forward primer, ATG TCT GAT AAT GGA CCC CAA AAT C; reverse primer, TTA GGC CTG AGT TGA GTC AGC.
Amino acid sequences of the proteins used in ELISA
-
HCoV-NL63 isolate Amsterdam-1, C-terminal N protein based on NL63 AA 215–377
KPNKPLSQPRADKPSQLKKPRWKRVPTREENVIQCFGPRDFNHNMGDSDLVQNGVDAKGFPQLAELIPNQAALFFDSEVSTDEVGDNVQITYTYKMLVAKDNKNLPKFIEQISAFTKPSSIKEMQSQSSHVAQNTVLNASIPESKPLADDDSAIIEIVNEVLH
-
HCoV-229E isolate Inf-1, C-terminal N protein based on 229E: AA 213–389
PSRNQSPASSQTSAKSLARSQSSETKEQKHEMQKPRWKRQPNDDVTSNVTQCFGPRDLDHNFGSAGVVANGVKAKGYPQFAELVPSTAAMLFDSHIVSKESGNTVVLTFTTRVTVPKDHPHLGKFLEELNAFTREMQQHPLLNPSALEFNPSQTSPATAEPVRDEVSIETDIIDEVN
-
HCoV-OC43 isolate VR-759, C-terminal N protein based on AY391777.1
RLELAKVQNLSGNPDEPQKDVYELRYNGAIRFDSTLSGFETIMKVLNENLNAYQQQDGMMNMSPKPQRQRGHKNGQGENDNISVAVPKSRVQQNKSRELTAEDISLLKKMDEPYTEDTSEI
-
HCoV-HKU1 isolate Caen1, C-terminal N protein based on HM034837.1
KLELVKRESEADSPVKDVFELRYSGSIRFDSTLPGFETIMKVLKENLDAYVNSNQNTVSGSLSPKPQRKRGVKQSPELLDPLNLSADTQHISNDFTPEDHSLLATLDDPYVEDSVA
-
SARS-CoV-2 isolate Wuhan-Hu-1, full nN protein based on NC_045512.2
MSDNGPQNQRNAPRITFGGPSDSTGSNQNGERSGARSKQRRPQGLPNNTASWFTALTQHGKEDLKFPRGQGVPINTNSSPDDQIGYYRRATRRIRGGDGKMKDLSPRWYFYYLGTGPEAGLPYGANKDGIIWVATEGALNTPKDHIGTRNPANNAAIVLQLPQGTTLPKGFYAEGSRGGSQASSRSSSRSRNSSRNSTPGSSRGTSPARMAGNGGDAALALLLLDRLNQLESKMSGKGQQQQGQTVTKKSAAEASKKPRQKRTATKAYNVTQAFGRRGPEQTQGNFGDQELIRQGTDYKHWPQIAQFAPSASAFFGMSRIGMEVTPSGTWLTYTGAIKLDDKDPNFKDQVILLNKHIDAYKTFPPTEPKKDKKKKADETQALPQRQKKQQTVTLLPAADLDDFSKQLQQSMSSADSTQA
ELISA seasonal coronavirus NCt proteins and SARS-CoV-2 N protein
The ELISAs were performed in 96 half-area microplates (Greiner Bio-One) coated overnight at 4 °C with 3 μg ml−1 of protein diluted in 0.1 M carbonate buffer, pH 9.6. Non-specific binding sites were blocked with PBS and 0.1% Tween 20 (PBST) supplemented with 5% skim milk (Fluka), mildly shaking for 1 h at room temperature. Longitudinal serum samples were diluted 1:200 in PBST containing 1% skim milk and incubated in the plate, mildly shaking for 2 h at room temperature. After a washing, alkaline phosphatase-conjugated AffiniPure Goat Anti-Human IgG, Fc Fragment Specific (Jackson ImmunoResearch, catalog no. 109–055–170), diluted (1:1,500) in 1% skim milk–PBST, was added. After mildly shaking for 1 h at room temperature, the plates were washed, and the signal was developed with Lumi-Phos Plus (Lumigen), which was incubated for 1 h in the dark at room temperature. Measurements were done with a GloMax 96 Microplate Luminometer (Promega). All sera were tested in duplicate or triplicate and normalized (replicates were performed on newly made dilutions of the same serum sample) to correct for differences in lumination times. All samples of one individual were tested within one ELISA plate or, when spread across two plates, with at least six samples on both plates to allow normalization of the signals.
Quantitative measles virus IgG ELISA
To determine natural fluctuation of antibodies, a quantitative measles virus antibody ELISA was used according to the manufacturer’s instructions (IBL International). Samples were examined in a 1:100 dilution if signals remained lower than the highest standard supplied with the assay. In case values were above the highest standard, a 1:200, 1:500 or 1:1,000 dilution was used, according to the manufacturer’s instructions, allowing quantification. All samples were tested in duplicate from new dilutions of the same serum sample.
HCoV-NL63 neutralization test
LLC-MK2 cells were cultured in minimal essential medium with Earle’s salt (Gibco), non-essential amino acids (Sciencell), L-glutamine 200 µM (Lonza) and penicillin–streptomycin (Lonza), 100 U ml−1 each, supplemented with 10% heat-inactivated fetal calf serum (Greiner Bio-One). Twenty-four hours before HCoV-NL63 infection, cells were plated onto 96-well plates at a density of 4 × 104 cells per well. Strain Amsterdam-119, passage 10 or passage 12 was used to infect LLC-MK2 cells at a multiplicity of infection of 0.01, in the presence of two-fold serial dilutions (highest concentration, 1:20 serum dilution). The infection medium was equal to the culture medium of LLC-MK2 cells, apart from the percent of heat-inactivated fetal calf serum, which was 3%. The 96-well plates were incubated at 33 °C, and, after 7 d, cytopathic effect was scored visually and confirmed with a cell viability assay (CellTiter 96 AQueous One Solution Cell Proliferation Assay, Promega).
Determination of threshold for infection
Coronavirus infections were determined by measuring fold changes in OD in NCt antibody recognition between two subsequent visits. We first measured the natural fluctuation among consecutive visits in measles virus antibodies for all ten individuals, assuming that measles infections did not occur during follow-up, as all individuals were vaccinated during childhood. Fold changes in antibody OD for measles virus ranged between 0.85 and 1.28 (Extended Data Fig. 1). Subsequently, a threshold for coronavirus infection was determined by evaluating the distribution of the OD fold change for each of the seasonal coronaviruses. To do so, we assumed that, during most intervals, no coronavirus infection occurred and that infections would, therefore, appear as outliers. Extended Data Fig. 1 shows that outliers were found for OD fold changes ≥1.40. We next determined whether these serological infection criteria could be confirmed with self-reported ILIs in the interval directly preceding the rise in antibodies. Indeed, reporting of any ILI symptom and fever (>38 °C > 3 d) alone were significantly associated with a ≥ 1.40 rise in antibodies (Fisher’s exact test, P = 0.028 and P = 0.024, respectively; Supplementary Table 4). Finally, we compared the ELISA results of HCoV-NL63 with neutralization titers for HCoV-NL63 for two individuals (#5 and #7, three infections). The infections showed an increase in neutralization titer accompanying the ≥1.40 fold rise in NCt antibodies (Extended Data Fig. 4).
Sensitivity
To determine the sensitivity of the NCt-ELISA, using the aforementioned OD fold change infection threshold, a total of 59 RT–PCR-confirmed coronavirus infections were investigated. This set included 11 HCoV-NL63, 16 HCoV-229E, 14 HCoV-OC43 and 18 HCoV-HKU1 infections, with Ct values less than 30 for HCoV-NL63, HCoV-229E and HCoV-OC43. For HCoV-HKU1 (which was assayed with a separate test (RespiFinder)), only positive and negative information was available, and no selection for low Ct values could be done. OD signals were tested on the V1 and V2 serum samples in duplicate (from separate dilutions of the same serum samples) for each coronavirus.
A sensitivity of 91% (10 of 11; one positive sample was later classified as a false positive because of cross-reactivity with HCoV-229E; see below) for HCoV-NL63, 81% (13 of 16) for HCoV-229E, 71% (10 of 14) for HCoV-OC43 and 22% (4 of 18) for HCoV-HKU1 was found (Extended Data Fig. 2, right panel). The low sensitivity for HCoV-HKU1 cannot be fully explained, but the unexpected low OD fold changes below 1.0 indicate that antibody levels were already elevated on V1 and started decreasing before V2, which might be a specific feature for this virus. If antibodies to HCoV-HKU1 wane more rapidly than antibodies to other coronaviruses, it might explain the low frequency of detecting an HCoV-HKU1 infection in individuals #1–#10 (Supplementary Fig. 1), and serology might, for this virus, be less sensitive to call infections for HCoV-HKU1.
Specificity
To determine the specificity of the ELISA assays, a total of 47 coronavirus-negative respiratory infections (but infections that were positive for other respiratory viruses) were added to the investigation. Seven rhinovirus, seven influenza A virus, six respiratory syncytial virus, eight parainfluenzavirus, seven adenovirus, five bocavirus and seven metapneumovirus infections were included. Sampling at V1 and V2 was as described above, and levels of antibodies were tested on the V1 and V2 serum samples as biological replicates (in duplo) for each coronavirus.
For HCoV-NL63, HCoV-229E and HCoV-OC43, only one of 47 paired sera tested positive using our serological assay. For HCoV-HKU1, two of 47 individuals tested positive. The specificities of the assays were, therefore, 98%, 98%, 98% and 96%, respectively (Extended Data Fig. 2, left panel). The false-positive HCoV-NL63 and HCoV-229E results were from the same sample.
Cross-reactivity
The cohort of coronavirus RT–PCR-positive samples also allowed us to evaluate the presence of cross-reactivity:
-
Intra-alphacoronavirus cross-reactivity: Cross-reactive antibody increases were seen for viruses within the same genus of the RT–PCR-confirmed virus. For example, four of the 11 (36%) RT-PCR-confirmed HCoV-NL63 infections showed, besides significant antibody reactivity (≥1.40 OD fold change) to NL63-NCt, also reactivity to HCoV-229E (Extended Data Fig. 3 and Supplementary Table 3). However, in all patients, the fold rise in OD was the highest for the coronavirus confirmed by RT–PCR by more than 10% in OD fold change difference (Extended Data Fig. 3), except for one patient. This patient, whose respiratory sample was RT–PCR positive for HCoV-NL63, had a slightly higher OD fold rise for HCoV-229E than for HCoV-NL63 (OD fold change of 10.28 versus 9.83, respectively, a 5% difference). To exclude false calling of HCoV-229E infections that were actually HCoV-NL63 infections, a cutoff was added for the calling of infections in individuals #1–#10: a minimum of 10% difference in fold change was needed to classify a ≥1.40 OD fold rise in antibodies as an infection for a specific virus within the same genus.
-
Intra-betacoronavirus cross-reactivity: Cross-reactive antibody increases were observed for both seasonal betacoronaviruses, but, in all cases, a higher OD rise was observed to the NCt of the matching RT-PCR-confirmed virus (Extended Data Fig. 3). Notably, all serum samples that were negative for the HCoV-HKU1 NCt-ELISA (the ELISA suspected to be of lower sensitivity) also showed no reactivity to the HCoV-OC43 NCt. Therefore, there was no possibility of mis-classification of infections by seasonal betacoronaviruses.
-
Inter-genus cross-reactivity: Cross-reactivity across genera was only for one of the 59 RT-PCR-confirmed HCoV infections. This outlier was seen for an RT–PCR-confirmed HCoV-229E infection that had a 6.16-OD fold rise for 229E but also a 1.50 fold rise for HCoV-HKU1. We, therefore, concluded that there are no significant signs of cross-reactivity between the seasonal alpha- and betacoronaviruses.
Statistical analysis
All analyses were performed using R version 3.6.2, with the following packages: reshape2, plyr, dplyr, zoo, EnvStats, survival and survminer. For the calculation of infections based on antibody rises, we determined that the interval between visits should be no longer than 400 d. Similarly, for the calculation of time interval between infections, intervals were determined only when all intermittent visits fell within the same 400-d limit.
The seasonality analysis was done by giving each infection a similar weight for the prevalence calculation. Because the infection might have occurred at any time during the interval preceding the rise in antibodies, the weight per infection was distributed uniformly over the interval, to the level of individual days. Doing so, it was assumed that the probability of infection onset was distributed over the interval. Per month, the total weight of all the infections was added and displayed as a percentage of the total of all infections across all months.
For the Kaplan–Meier estimator, each infection per individual was counted as a single observation; all infections from all individuals were subsequently used for this analysis. The visit at which the infection was established was counted as time point 0; individuals were subsequently followed until the antibody levels dropped below the indicated level (event). Three different event types were calculated per infection: a 50%, a 75% and a 100% return to pre-infection antibody levels (defined by the antibody level at the last visit before infection). When the antibody level did not decrease to the indicated levels by the end of a continuous follow-up period, an observation was censored at the last visit.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
We have provided all of the de-identified raw data used in this paper, including all figures, in the Supplementary Data. Materials are available for study via the open and ongoing Amsterdam Cohort Studies on HIV among men who have sex with men (MSM): https://www.ggd.amsterdam.nl/beleid-onderzoek/projecten/amsterdamse-cohort/. Source data are provided with this paper.
References
Pyrc, K. et al. Mosaic structure of human coronavirus NL63, one thousand years of evolution. J. Mol. Biol. 364, 964–973 (2006).
Callow, K. A., Parry, H. F., Sergeant, M. & Tyrrell, D. A. J. The time course of the immune response to experimental coronavirus infection of man. Epidemiol. Infect. 105, 435–446 (1990).
Van Bilsen, W. P. H. et al. Diverging trends in incidence of HIV versus other sexually transmitted infections in HIV-negative MSM in Amsterdam. AIDS 34, 301–309 (2020).
Blanchard, E. G., Miao, C., Haupt, T. E., Anderson, L. J. & Haynes, L. M. Development of a recombinant truncated nucleocapsid protein based immunoassay for detection of antibodies against human coronavirus OC43. J. Virol. Methods 177, 100–106 (2011).
Li, Y., Wang, X. & Nair, H. Global seasonality of human seasonal coronaviruses: a clue for postpandemic circulating season of severe acute respiratory syndrome coronavirus 2? J. Infect. Dis. 222, 1090–1097 (2020).
Reed, S. E. The behaviour of recent isolates of human respiratory coronavirus in vitro and in volunteers: evidence of heterogeneity among 229E‐related strains. J. Med. Virol. 13, 179–192 (1984).
Long, Q. X. et al. Clinical and immunological assessment of asymptomatic SARS-CoV-2 infections. Nat. Med. 26, 1200–1204 (2020).
Ibarrondo, F. J. et al. Rapid decay of anti–SARS-CoV-2 antibodies in persons with mild Covid-19. N. Engl. J. Med. https://doi.org/10.1056/NEJMc2025179 (2020).
Ieven, M. et al. Aetiology of lower respiratory tract infection in adults in primary care: a prospective study in 11 European countries. Clin. Microbiol. Infect. 24, 1158–1163 (2018).
Gobat, N. et al. Preparedness for clinical research during pandemics: a perspective from the Platform for European Preparedness Against (Re-)emerging Epidemics (PREPARE). Lancet 392, S38 (2018).
Loens, K. et al. Performance of different mono- and multiplex nucleic acid amplification tests on a multipathogen external quality assessment panel. J. Clin. Microbiol. 50, 977–987 (2012).
Reed, L. J. & Muench, H. A simple method of estimating fifty per cent endpoints. Am. J. Epidemiol. 27, 493–497 (1938).
Dijkman, R. et al. The dominance of human coronavirus OC43 and NL63 infections in infants. J. Clin. Virol. 53, 135–139 (2012).
Severance, E. G. et al. Development of a nucleocapsid-based human coronavirus immunoassay and estimates of individuals exposed to coronavirus in a U.S. metropolitan population. Clin. Vaccin. Immunol. 15, 1805–1810 (2008).
Sastre, P. et al. Differentiation between human coronaviruses NL63 and 229E using a novel double-antibody sandwich enzyme-linked immunosorbent assay based on specific monoclonal antibodies. Clin. Vaccin. Immunol. 18, 113–118 (2011).
Dijkman, R. et al. Human coronavirus NL63 and 229E seroconversion in children. J. Clin. Microbiol. 46, 2368–2373 (2008).
Lehmann, C. et al. A line immunoassay utilizing recombinant nucleocapsid proteins for detection of antibodies to human coronaviruses. Diagn. Microbiol. Infect. Dis. 61, 40–48 (2008).
Gorse, G. J., Donovan, M. M. & Patel, G. B. Antibodies to coronaviruses are higher in older compared with younger adults and binding antibodies are more sensitive than neutralizing antibodies in identifying coronavirus‐associated illnesses. J. Med. Virol. 92, 512–517 (2020).
van der Hoek, L. et al. Identification of a new human coronavirus. Nat. Med. 10, 368–373 (2004).
Acknowledgements
The authors gratefully acknowledge the Amsterdam Cohort Studies (ACS) on HIV infection and AIDS, a collaboration among the Public Health Service of Amsterdam, the Amsterdam University Medical Center of the University of Amsterdam, the Sanquin Blood Supply Foundation, the Medical Center Jan van Goyen and the HIV Focus Center of the DC-Clinics. It is part of the Netherlands HIV Monitoring Foundation and financially supported by the Center for Infectious Disease Control of the Netherlands National Institute for Public Health and the Environment. The authors thank all ACS participants for their contribution, as well as the ACS study nurses, data managers and lab technicians. We thank K. Jacobs for his help with lifting of GRACE consortium samples and D. Eggink for PREPARE consortium samples. We acknowledge the GRACE network (LSHM-CT-2005–518226), the European Science Foundation Research Networking Programmes network TRACE and the PREPARE project (funded through the European Union FP7 Programme no. 602525). This work was supported by a grant from the European Union’s Horizon 2020 Research and Innovation Programme, under Marie Skłodowska-Curie Actions grant agreement no. 721367 (HONOURs), Amsterdam University Medical Center funding connected to HONOURs and the Amsterdam University Medical Center PhD scholarship of A. W. D. Edridge.
Author information
Authors and Affiliations
Contributions
A.W.D.E: conceptualization, writing (original draft, review and editing), investigation, visualization, formal analysis and validation; J.K.: investigation and writing (original draft, review and editing); A.C.R.H.: resources; M.B.: investigation and resources; M.K.: investigation; K.L: resources; M.F.J.: investigation and methodology; A.M.: formal analysis; C.M.K.: writing (original draft, review and editing) and formal analysis; P.R.: resources; M.I.: resources; H.G.: resources; M.P.: resources; P.S.: resources; M.D.: investigation, visualization and methodology; L.vd.H.: conceptualization, writing (original draft, review and editing) and supervision.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Alison Farrell was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Threshold for infection based on distribution of fold changes in OD.
Boxplots show median values with the interquartile range (IQR), the whiskers extend up to the range after which samples are considered outliers (1.5 x IQR below the first or above the third quantile). The 1.40 threshold for infection is indicated by the dotted line.
Extended Data Fig. 2 Specificity and sensitivity of the seasonal coronavirus ELISA assays.
Left panel: specificity analysis. Forty-seven subjects infected by a respiratory virus (non-coronavirus) were tested for significant increases in anti-NCt antibodies directed to the four seasonal coronaviruses. V1 (acute symptomatic infection) and V2 (28–35 days later) sera were compared. On the Y-axis the V2/V1 OD fold change is shown and a ratio ≥ 1.40 is regarded as positive. Right panel: sensitivity analysis. Fifty-nine subjects with an RT-PCR confirmed coronavirus infection were tested for development of antibodies directed to the four seasonal coronaviruses.
Extended Data Fig. 3 Lack of misassignment due to cross reactivity in the ELISAs.
The V1 and V2 serum samples were tested in the ELISAs for each virus. The OD value V2/V1 was calculated for each infection (= fold change). For those fold changes that were above 1.40 we calculated the ratio between the fold changes for the viruses that may or may not have cross-reacted. In each panel, the ratio of the fold changes is displayed on the Y-axis. A ratio below one indicates that the fold change measured in ELISA is the highest for the virus that was found by RT-PCR in the corresponding respiratory sample. Only one patient (infected by HCoV-NL63) displayed, besides a large rise in HCoV-NL63, also a substantial rise in HCoV-229E OD, and for this patient the fold changes were similar (9.7 versus 10.3) resulting in a 1.1 ratio (far left panel, the one data-point above 1). For all other patients with ≥1.40 fold rises matched with the RT-PCR confirmed infection.
Extended Data Fig. 4 Comparison of HCoV-NL63-NCt ELISA and HCoV-NL63 neutralization.
Red: antibody dynamics measured by ELISA, blue: neutralization titers.
Extended Data Fig. 5 Broadly recognizing coronavirus antibodies.
From top to bottom: SARS-CoV2 N-protein, HCoV-NL63, HCoV-229E, HCoV-OC43, and HCoV-HKU1 NCt, measles virus, and negative control his-tagged HIV-1 envelope SOSIP protein (van Gils et al Nature Microbiology 2016, doi: 10.1038/nmicrobiol.2016.199).
Extended Data Fig. 6 Kaplan-Meier curve of post-infection antibody depletion.
Red continuous line: 50% decline of antibody levels to pre-infection levels, blue dotted line with short dots: 75% decline, green dotted line with log dots: 100% decline.
Supplementary information
Supplementary Information
Supplementary Fig. 1 and Supplementary Tables 1–4.
Supplementary Data
Source data of all ELISA experiments and the data underlying Supplementary Fig. 1.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Rights and permissions
About this article
Cite this article
Edridge, A.W.D., Kaczorowska, J., Hoste, A.C.R. et al. Seasonal coronavirus protective immunity is short-lasting. Nat Med 26, 1691–1693 (2020). https://doi.org/10.1038/s41591-020-1083-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-020-1083-1
This article is cited by
-
Recombination analysis on the receptor switching event of MERS-CoV and its close relatives: implications for the emergence of MERS-CoV
Virology Journal (2024)
-
Clinical course and management of COVID-19 in the era of widespread population immunity
Nature Reviews Microbiology (2024)
-
Computational design and evaluation of mRNA- and protein-based conjugate vaccines for influenza A and SARS-CoV-2 viruses
Journal of Genetic Engineering and Biotechnology (2023)
-
SARS-CoV-2 and innate immunity: the good, the bad, and the “goldilocks”
Cellular & Molecular Immunology (2023)
-
SARS-CoV-2 variant biology: immune escape, transmission and fitness
Nature Reviews Microbiology (2023)