Abstract
PD-1 blockade has transformed the management of advanced clear cell renal cell carcinoma (ccRCC), but the drivers and resistors of the PD-1 response remain incompletely elucidated. Here, we analyzed 592 tumors from patients with advanced ccRCC enrolled in prospective clinical trials of treatment with PD-1 blockade by whole-exome and RNA sequencing, integrated with immunofluorescence analysis, to uncover the immunogenomic determinants of the therapeutic response. Although conventional genomic markers (such as tumor mutation burden and neoantigen load) and the degree of CD8+ T cell infiltration were not associated with clinical response, we discovered numerous chromosomal alterations associated with response or resistance to PD-1 blockade. These advanced ccRCC tumors were highly CD8+ T cell infiltrated, with only 27% having a non-infiltrated phenotype. Our analysis revealed that infiltrated tumors are depleted of favorable PBRM1 mutations and enriched for unfavorable chromosomal losses of 9p21.3, as compared with non-infiltrated tumors, demonstrating how the potential interplay of immunophenotypes with somatic alterations impacts therapeutic efficacy.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All relevant data are available from the authors and/or are included with the manuscript. Clinical data about the patients and tumor immunophenotyping are listed in Supplementary Table 1. Somatic mutations are available in Supplementary Table 2. Significantly recurrent mutations (by MutSig2CV) and copy number alterations (by GISTIC2) are available in supplementary table 3. Normalized RNA-seq expression data, single sample gene set enrichment scores, immune deconvolution (by CIBERSORTx), and ERV expression (inferred from RNA-seq data) are available in Supplementary Table 4. WES data from patients who consented to deposition have been submitted to the European Genome-phenome Archive (Accession numbers EGAS00001004290, EGAS00001004291, EGAS00001004292).
Code availability
Algorithms used for data analysis are all publicly available from the indicated references in the Methods section.
References
Ribas, A. & Wolchok, J. D. Cancer immunotherapy using checkpoint blockade. Science 359, 1350–1355 (2018).
Motzer, R. J. et al. Nivolumab versus everolimus in advanced renal-cell carcinoma. N. Engl. J. Med. 373, 1803–1813 (2015).
Motzer, R. J. et al. Avelumab plus axitinib versus sunitinib for advanced renal-cell carcinoma. N. Engl. J. Med. 380, 1103–1115 (2019).
Motzer, R. J. et al. Nivolumab plus ipilimumab versus sunitinib in advanced renal-cell carcinoma. N. Engl. J. Med. 378, 1277–1290 (2018).
Rini, B. I. et al. Pembrolizumab plus axitinib versus sunitinib for advanced renal-cell carcinoma. N. Engl. J. Med. 380, 1116–1127 (2019).
Choueiri, T. K. & Motzer, R. J. Systemic therapy for metastatic renal-cell carcinoma. N. Engl. J. Med. 376, 354–366 (2017).
Yarchoan, M., Hopkins, A. & Jaffee, E. M. Tumor mutational burden and response rate to PD-1 inhibition. N. Engl. J. Med. 377, 2500–2501 (2017).
Rizvi, N. A. et al. Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348, 124–128 (2015).
Critescu, R. et al. Pan-tumor genomic biomarkers for PD-1 checkpoint blockade-based immunotherapy. Science 362, eaar3593 (2018).
Mariathasan, S. et al. TGFβ attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature 554, 544–548 (2018).
Lawrence, M. S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013).
Fridman, W. H., Zitvogel, L., Sautes-Fridman, C. & Kroemer, G. The immune contexture in cancer prognosis and treatment. Nat. Rev. Clin. Oncol. 14, 717–734 (2017).
Cancer Genome Atlas Research Network. Comprehensive molecular characterization of clear cell renal cell carcinoma. Nature 499, 43–49 (2013).
Turajlic, S. et al. Tracking cancer evolution reveals constrained routes to metastases: TRACERx renal. Cell 173, 581–594 (2018).
Binnewies, M. et al. Understanding the tumor immune microenvironment (TIME) for effective therapy. Nat. Med. 24, 541–550 (2018).
Chen, D. S. & Mellman, I. Elements of cancer immunity and the cancer-immune set point. Nature 541, 321–330 (2017).
Choueiri, T. K. et al. Immunomodulatory activity of nivolumab in metastatic renal cell carcinoma. Clin. Cancer Res. 22, 5461–5471 (2016).
Motzer, R. J. et al. Nivolumab for metastatic renal cell carcinoma: results of a randomized phase II trial. J Clin Oncol 33, 1430–1437 (2015).
Miao, D. et al. Genomic correlates of response to immune checkpoint therapies in clear cell renal cell carcinoma. Science 359, 801–806 (2018).
Lawrence, M. S. et al. Discovery and saturation analysis of cancer genes across 21 tumour types. Nature 505, 495–501 (2014).
Mermel, C. H. et al. GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. 12, R41 (2011).
Mitchell, T. J. et al. Timing the landmark events in the evolution of clear cell renal cell cancer: TRACERx renal. Cell 173, 611–623 (2018).
Mashtalir, N. et al. Modular organization and assembly of SWI/SNF family chromatin remodeling complexes. Cell 175, 1272–1288 (2018).
Chen, Y. B. et al. Molecular analysis of aggressive renal cell carcinoma with unclassified histology reveals distinct subsets. Nat. Commun. 7, 13131 (2016).
Pal, S. K. et al. Characterization of clinical cases of collecting duct carcinoma of the kidney assessed by comprehensive genomic profiling. Eur. Urol. 70, 516–521 (2016).
Malouf, G. G. et al. Genomic characterization of renal cell carcinoma with sarcomatoid dedifferentiation pinpoints recurrent genomic alterations. Eur. Urol. 70, 348–357 (2016).
Pal, S. K. et al. Characterization of clinical cases of advanced papillary renal cell carcinoma via comprehensive genomic profiling. Eur. Urol. 73, 71–78 (2018).
van Slegtenhorst, M. et al. Identification of the tuberous sclerosis gene TSC1 on chromosome 9q34. Science 277, 805–808 (1997).
Yan, L. et al. Genetic alteration of histone lysine methyltransferases and their significance in renal cell carcinoma. PeerJ 7, e6396 (2019).
Heng, D. Y. et al. Prognostic factors for overall survival in patients with metastatic renal cell carcinoma treated with vascular endothelial growth factor-targeted agents: results from a large, multicenter study. J. Clin. Oncol. 27, 5794–5799 (2009).
Motzer, R. J. et al. Prognostic factors for survival in previously treated patients with metastatic renal cell carcinoma. J. Clin. Oncol. 22, 454–463 (2004).
Snyder, A. et al. Genetic basis for clinical response to CTLA-4 blockade in melanoma. N. Engl. J. Med. 371, 2189–2199 (2014).
Van Allen, E. M. et al. Genomic correlates of response to CTLA-4 blockade in metastatic melanoma. Science 350, 207–211 (2015).
Turajlic, S. et al. Insertion-and-deletion-derived tumour-specific neoantigens and the immunogenic phenotype: a pan-cancer analysis. Lancet Oncol. 18, 1009–1021 (2017).
Braun, D. A. et al. Clinical validation of PBRM1 alterations as a marker of immune checkpoint inhibitor response in renal cell carcinoma. JAMA Oncol. 5, 1631–1633 (2019).
McDermott, D. F. et al. Clinical activity and molecular correlates of response to atezolizumab alone or in combination with bevacizumab versus sunitinib in renal cell carcinoma. Nat. Med. 24, 749–757 (2018).
George, S. et al. Loss of PTEN is associated with resistance to anti-PD-1 checkpoint blockade therapy in metastatic uterine leiomyosarcoma. Immunity 46, 197–204 (2017).
Peng, W. et al. Loss of PTEN promotes resistance to T cell-mediated immunotherapy. Cancer Discov. 6, 202–216 (2016).
Hanzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinf. 14, 7 (2013).
Liberzon, A. et al. The molecular signatures database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Taylor, M. D. et al. Mutations in SUFU predispose to medulloblastoma. Nat. Genet. 31, 306–310 (2002).
Rooney, M. S., Shukla, S. A., Wu, C. J., Getz, G. & Hacohen, N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 160, 48–61 (2015).
Smith, C. C. et al. Endogenous retroviral signatures predict immunotherapy response in clear cell renal cell carcinoma. J. Clin. Invest. 128, 4804–4820 (2018).
Panda, A. et al. Endogenous retrovirus expression is associated with response to immune checkpoint blockade in clear cell renal cell carcinoma. JCI Insight 3, 121522 (2018).
Takahashi, Y. et al. Regression of human kidney cancer following allogeneic stem cell transplantation is associated with recognition of an HERV-E antigen by T cells. J. Clin. Invest. 118, 1099–1109 (2008).
Newman, A. M. et al. Determining cell type abundance and expression from bulk tissues with digital cytometry. Nat. Biotechnol. 37, 773–782 (2019).
Chevrier, S. et al. An immune atlas of clear cell renal cell carcinoma. Cell 169, 736–749 (2017).
Choueiri, T. K. et al. Biomarker analyses from JAVELIN Renal 101: avelumab + axitinib (A+Ax) versus sunitinib (S) in advanced renal cell carcinoma (aRCC). J. Clin. Oncol. 37, 101–101 (2019).
Danaher, P. et al. Pan-cancer adaptive immune resistance as defined by the tumor inflammation signature (TIS): results from The Cancer Genome Atlas (TCGA). J. Immunother. Cancer 6, 63 (2018).
Sato, Y. et al. Integrated molecular analysis of clear-cell renal cell carcinoma. Nat. Genet. 45, 860–867 (2013).
Gerlinger, M. et al. Genomic architecture and evolution of clear cell renal cell carcinomas defined by multiregion sequencing. Nat. Genet. 46, 225–233 (2014).
Gerlinger, M. et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 366, 883–892 (2012).
Rosenthal, R. et al. Neoantigen-directed immune escape in lung cancer evolution. Nature 567, 479–485 (2019).
Hakimi, A. A. et al. Transcriptomic profiling of the tumor microenvironment reveals distinct subgroups of clear cell renal cell cancer: data from a randomized phase III trial. Cancer Discov. 9, 510–525 (2019).
Nargund, A. M. et al. The SWI/SNF protein PBRM1 restrains VHL-loss-driven clear cell renal cell carcinoma. Cell Rep. 18, 2893–2906 (2017).
Gao, W., Li, W., Xiao, T., Liu, X. S. & Kaelin, W. G.Jr. Inactivation of the PBRM1 tumor suppressor gene amplifies the HIF-response in VHL-/- clear cell renal carcinoma. Proc. Natl Acad. Sci. USA 114, 1027–1032 (2017).
Palmer, A. C. & Sorger, P. K. Combination cancer therapy can confer benefit via patient-to-patient variability without drug additivity or synergy. Cell 171, 1678–1691.e1613 (2017).
Cibulskis, K. et al. ContEst: estimating cross-contamination of human samples in next-generation sequencing data. Bioinformatics 27, 2601–2602 (2011).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
Saunders, C. T. et al. Strelka: accurate somatic small-variant calling from sequenced tumor-normal sample pairs. Bioinformatics 28, 1811–1817 (2012).
Ramos, A. H. et al. Oncotator: cancer variant annotation tool. Hum. Mutat. 36, E2423–E2429 (2015).
Costello, M. et al. Discovery and characterization of artifactual mutations in deep coverage targeted capture sequencing data due to oxidative DNA damage during sample preparation. Nucleic Acids Res. 41, e67 (2013).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Carter, S. L. et al. Absolute quantification of somatic DNA alterations in human cancer. Nat. Biotechnol. 30, 413–421 (2012).
Zack, T. I. et al. Pan-cancer patterns of somatic copy number alteration. Nat. Genet. 45, 1134–1140 (2013).
Shukla, S. A. et al. Comprehensive analysis of cancer-associated somatic mutations in class I HLA genes. Nat. Biotechnol. 33, 1152–1158 (2015).
Jurtz, V. et al. NetMHCpan-4.0: improved peptide-MHC class I interaction predictions integrating eluted ligand and peptide binding affinity data. J. Immunol. 199, 3360–3368 (2017).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinf. 12, 323 (2011).
DeLuca, D. S. et al. RNA-SeQC: RNA-seq metrics for quality control and process optimization. Bioinformatics 28, 1530–1532 (2012).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Vargiu, L. et al. Classification and characterization of human endogenous retroviruses; mosaic forms are common. Retrovirology 13, 7 (2016).
Anders, S., Pyl, P. T. & Huber, W. HTSeq-a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Specht, K. et al. Quantitative gene expression analysis in microdissected archival formalin-fixed and paraffin-embedded tumor tissue. Am. J. Pathol. 158, 419–429 (2001).
Giraldo, N. A. et al. Orchestration and prognostic significance of immune checkpoints in the microenvironment of primary and metastatic renal cell cancer. Clin. Cancer Res. 21, 3031–3040 (2015).
Granier, C. et al. Tim-3 expression on tumor-infiltrating PD-1+ CD8+ T cells correlates with poor clinical outcome in renal cell carcinoma. Cancer Res. 77, 1075–1082 (2017).
Acknowledgements
We are grateful to G. Getz, I. Leshchiner, S. Moreno, R. Beroukhim, M. Sticco-Ivins, S. Gohil, P. Bachireddy, M. Atkins, K. Mahoney, R. Bhatt, G. Bouchard and M. Lee for discussions and input. We also appreciate the efforts of all study nurses and clinical staff that made this study feasible, and patients who generously provided their samples for this research. This works was supported in part by Dana-Farber/Harvard Cancer Center Kidney Cancer SPORE (P50-CA101942-12), DOD CDMRP (W81XWH-18-1-0480), and in part by Bristol-Myers Squibb. D.A.B. is supported by the John. R. Svenson Fellowship. C.J.W. acknowledges support from NIH: NCI-1RO1CA155010 and NIH/NCI U24 CA224331. This work was supported in part by The G. Harold and Leila Y. Mathers Foundation. C.J.W. is a scholar of the Leukemia and Lymphoma Society, and is supported in part by the Parker Institute for Cancer Immunotherapy. S.A.S acknowledges support by the NCI (R50RCA211482). T.K.C. is supported in part by the Dana-Farber/Harvard Cancer Center Kidney SPORE and Program, the Kohlberg Chair at Harvard Medical School and the Trust Family, Michael Brigham, and Loker Pinard Funds for Kidney Cancer Research at DFCI.
Author information
Authors and Affiliations
Contributions
D.A.B., D.F.M., S.S., C.J.W., S.A.S. and T.K.C. designed the study, analyzed and interpreted the data. D.A.B., Y.H., Z.B., J.F., A.C.B., L.E., A.D.C., L.L. and S.A.S. performed the genomic and transcriptomic analysis, including data filtering, identification of recurrent mutations and copy number alterations, and association with immune phenotypes and clinical outcomes. M.F., M.S.A., J.-C.P. and S.S. performed the immunofluorescence analysis. O.A.J. and P.C. performed statistical analysis. J.S., M.S., P.R.-M. and M.W.-R. contributed to the collection and assembly of data, and study design. M.W.-R, D.N., G.J.F. and A.H.S. contributed to the interpretation of the data. D.A.B., E.M.V.A., S.S., C.J.W., S.A.S. and T.K.C wrote the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
D.A.B. reported non-financial support from Bristol-Myers Squibb, honoraria from LM Education/Exchange Services, and personal fees from Octane Global, Defined Health, Dedham Group, Adept Field Solutions, Slingshot Insights, Blueprint Partnerships, Charles River Associates, Trinity Group, and Insight Strategy, outside of the submitted work. M.S. reported grants from Exelixis and Pfizer. G.J.F. and A.H.S. have patents/pending royalties on the PD-1 pathway from Roche, Merck, Bristol-Myers-Squibb, EMD-Serono, Boehringer-Ingelheim, AstraZeneca, Dako and Novartis. G.J.F. has equity in Nextpoint, Triursus, and Xios. G.J.F. has served on advisory boards for Roche, Bristol-Myers-Squibb, Xios, Origimed, NextPoint, and IgM. G.J.F. has received research funding from Bristol-Myers Squibb, outside of the submitted work. A.H.S. has served on advisory boards for Novartis, Surface Oncology, Elstar, SQZ Biotechnologies, Adaptimmune, Elpiscience, Monopteros. A.H.S. has received research funding from Novartis, Roche, UCB, Ipsen, Quark and Merck. D.F.M. reported personal fees from Bristol-Myers Squibb, Pfizer, Merck, Novartis, Exelixis, Array BioPharm, Genentech, Alkermes, Jounce Therapeutics, X4 Pharma, Peloton, EMD Serono, and Eli Lilly; research support from Bristol-Myers Squibb, Prometheus Laboratories, Merck, Genentech, Pfizer, Exelixis, Novartis, X4 Pharma, Alkermes, and Peloton. E.M.V.A. had a patent to the association of mutations in PBAF genes and response to cancer immunotherapy pending. E.M.V.A. reported personal fees from Tango Therapeutics, Genome Medical, Invitae, Illumina, and Dynamo; grants from Novartis, Bristol-Myers Squibb-IION, nonfinancial support from Genentech, personal fees from Synapse and Microsoft outside the submitted work. S.S. reported personal fees from Merck, AstraZeneca, Bristol-Myers Squibb, AACR, and NCI, grants from Bristol-Myers Squibb, AstraZeneca, Novartis, and Exelixis, and royalties from Biogenex. C.J.W. is a founder and equity holder of Neon Therapeutics and a member of its scientific advisory board. S.A.S. reported nonfinancial support from Bristol-Myers Squibb outside the submitted work. S.A.S. previously advised and has received consulting fees from Neon Therapeutics. S.A.S. reported nonfinancial support from Bristol-Myers Squibb, and equity in Agenus Inc., Agios Pharmaceuticals, Breakbio Corp., Bristol-Myers Squibb and NewLink Genetics, outside the submitted work. T.K.C. reported grants and personal fees from AstraZeneca, personal fees from Bayer, grants and personal fees from Bristol-Myers Squibb, personal fees from Cerulean, grants and personal fees from Eisai, personal fees from Foundation Medicine Inc, grants and personal fees from Exelixis, grants and personal fees from Genentech, personal fees from Roche, grants and personal fees from GlaxoSmithKline, grants and personal fees from Merck, from Novartis, Peloton, and Pfizer, personal fees from Prometheus Labs, grants and personal fees from Corvus, personal fees from Ipsen, grants from Tracon, grants from Astellas outside the submitted work. The other authors declare no conflicts of interest.
Additional information
Peer review information Saheli Sadanand was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
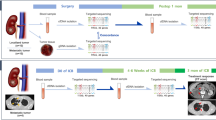
Extended Data Fig. 1 Sample inclusion and exclusion criteria, including quality control filtering, for CM-010.
a, WES and (b) RNA-seq, and CM-025 (c) WES and (d) RNA-seq analyses.
Extended Data Fig. 2 Response and survival for sequenced vs. non-sequenced patients in each cohort.
Comparison of PFS (a,c,e) and OS (b,d,f) included and excluded patients for (a-b) WES, (c-d) RNA-seq, and (e-f) immunofluorescence analysis. Left panels, CM-010; middle panels, CM-025 patients treated with PD-1 blockade; right panels, CM-025 patients treated with mTOR inhibition. CM-025 patients treated with PD-1 blockade who were included in WES analysis had shorter OS than patients not included (p < 0.05). There were no other significant differences between included and excluded patients (two-sided log-rank test). g, There were no significant differences in PFS or OS among anti-PD-1 treated patients in the three trial cohorts.
Extended Data Fig. 3 Genomic and immune features associated with MSKCC risk groups.
a-b, Total number of nonsynonymous mutations and neoantigen load were not significantly different between risk groups (two-sided Wilcoxon rank-sum test). c-d, Poor risk tumors were associated with a lower number of frameshift indels (compared to intermediate risk), and a higher copy number burden / chromosomal instability (measured by wGII, compared to intermediate and favorable risk; two-sided Wilcoxon rank-sum test). e, Total CD8 + T cell infiltration (by immunofluorescence) did not significantly differ between risk groups (two-sided Wilcoxon rank-sum test). Boxplot hinges represent 25th to 75th percentiles, central lines represent the medians, the whiskers extend to highest and lowest values no greater than 1.5× interquartile range and the dots indicate outliers; the violin component refers to the kernel probability density and encompasses all cells. f, Individual somatic copy number alterations (sCNAs) that different in frequency between MSKCC risk groups (two-sided chi-squared test; p < 0.05). No mutations (sSNVs and sIndels) significantly differed in frequency between risk groups.
Extended Data Fig. 4 Survival of patients with high versus low somatic alteration burden.
There were no significant differences in response (complete or partial response [CR/PR] vs progressive disease [PD], two-sided Wilcoxon ranksum test) or PFS or OS (two-sided log-rank test) in anti-PD-1 and mTOR inhibitor treated patients with respect to (a) total nonsynonymous mutation burden, (b) neoantigen load, (c) number of frameshift insertions and deletions, or (d) copy number burden / chromosomal instability (wGII). Boxplot hinges represent 25th to 75th percentiles, central lines represent the medians, the whiskers extend to highest and lowest values no greater than 1.5× interquartile range and the dots indicate outliers; the violin component refers to the kernel probability density and encompasses all cells. e, There was no significant difference in PFS or OS (two-sided log-rank test) between patients with high or low intratumor heterogeneity (ITH; ratio of subclonal mutations to clonal mutations) or wGII. Assignment of ‘high’ or ‘low’ for each feature was based on a median cutoff (for each somatic feature, tumors were assigned as ‘high’ if they were at or above the median cohort value for that specific somatic feature). f, Response to PD-1 blockade was not significantly different between patients who were heterozygous for HLA class I at all alleles, and patients who were homozygous at all alleles (two-sided Fisher’s exact test between CR/PR and PD. Error bars are SEM and measure of center is mean).
Extended Data Fig. 5 Genomic correlates of survival following anti-PD-1 or mTOR treatment.
a, PBRM1 truncating mutations are recurrent (MutSig2CV q < 0.05) and associated with better PFS (two-sided univariable Cox regression with truncating mutation status as a categorical covariate, p < 0.05) with anti-PD-1 therapy (n = 249 patients). b, Deletions in 10q23.31 are recurrent (GISTIC2 q < 0.1) and associated with improved PFS (two-sided univariable Cox regression with copy number deletion status as a categorical covariate, p < 0.05) following anti-PD-1 therapy (n = 249 patients). (c-d) No recurrent amplifications (GISTIC2 q < 0.1) are significantly associated with improved or worsened PFS and OS (two-sided univariable Cox regression with copy number amplification status as a categorical covariate, n = 249 patients). e, RNA-seq—based inference of ERV expression correlates with experimental RT-qPCR expression measurements, with the exception of ERV-3-2 (Spearman correlation, p values obtained from two-sided t-tests, n = 36 RT-qPCR samples). f, As continuous variables, expression of ERV2282 and ERV3382 are associated with altered response (two-sided Fisher’s exact test for clinical benefit vs. no clinical benefit, p < 0.05, n = 57 clinical benefit patients and n = 67 no clinical benefit patients), PFS, and OS (two-sided univariable Cox regression with ERV expression as a continuous covariate, p < 0.05, n = 181 patients with anti-PD-1 therapy). g, However, when samples were dichotomized into ERV expression high vs. low (based on median expression of each ERV), ERV2282 and ERV3382 were no long significant for both altered PFS and OS (two-sided log-rank test).
Extended Data Fig. 6 Characterization of immune infiltration and its association with clinical outcome.
a, Estimated CD8 + T cell infiltration by CIBERSORTx deconvolution of RNA-seq data is highly correlated with experimental CD8 + T cell measurements by immunofluorescence (Spearman correlation, p values obtained from two-sided t-tests, n = 144 samples with both mutation calls and IF data). b, RNA-seq—based inference of T cell infiltration results in some misclassifications of infiltrated and non-infiltrated tumors. Left panel, immunophenotype by CD8 IF, with each sample colored by RNA-seq—based T effector cell infiltration score (IMmotion 150 Teff signature)36. Right panels, examples of misclassifications using the RNA-seq—based T effector cell infiltration score, including (top) a tumor with low Teff signature but high CD8 + T cell infiltration by IF, and (bottom) a tumor with high Teff signature but low CD8 + T cell infiltration by IF. c, No association was observed between immune infiltration phenotype and PFS (left panel) or OS (right panel) with MTOR inhibition (two-sided log-rank test).
Extended Data Fig. 7 Immune-related gene signature expression is not associated with improved response or survival with anti-PD-1 therapy.
For established immune signatures of (a-b) T effector cell infiltration (IMmotion 150 Teff)36, (c-d) myeloid cell infiltration (IMmotion150 Myeloid)36, (e-f) immune infiltration (JAVELIN)48, and (g-h) tumor inflammation (TIS)49, high signature scores were not associated with improved (a,c,e,g) clinical benefit (two-sided Wilcoxon rank-sum test) or (b,d,f,h) survival (two-sided log-rank test). A high IMmotion150 Teff signature was associated with significantly worse OS with PD-1 blockade (p = 0.018, two-sided log-rank test), but was not significantly associated with altered clinical benefit or PFS. Assignment of ‘high’ or ‘low’ for each signature was based on a median cutoff (for each signature, tumors were assigned as ‘high’ if that tumor’s signature score was at or above the median value). Boxplot hinges represent 25th to 75th percentiles, central lines represent the medians, the whiskers extend to highest and lowest values no greater than 1.5× interquartile range and the dots indicate outliers; the violin component refers to the kernel probability density and encompasses all cells.
Extended Data Fig. 8 Enrichment of individual mutations and chromosomal instability in infiltrated versus noninfiltrated tumors.
a, PBRM1 mutations were the only significantly enriched recurrent mutation in non-infiltrated tumors (p = 0.0126, two-sided Fisher’s exact test for infiltrated vs. non-infiltrated tumors, n = 79 infiltrated and n = 24 noninfiltrated). There were no recurrent significantly enriched mutations in infiltrated tumors. Dotted lines indicate p-values of 0.25 (dark) and 0.05 (light). Chromosomal instability (that is copy number burden, as measured by wGII) is (b) increased in infiltrated tumors (p = 0.012, two-sided Fisher’s exact test), but is not associated with significantly altered (c) clinical benefit (two-sided Wilcoxon rank-sum test) or (d) PFS or OS (two-sided log-rank test) with PD-1 blockade or mTOR inhibition. Boxplot hinges represent 25th to 75th percentiles, central lines represent the medians, the whiskers extend to highest and lowest values no greater than 1.5× interquartile range and the dots indicate outliers; the violin component refers to the kernel probability density and encompasses all cells.
Extended Data Fig. 9 Association of focal amplifications and deletions with T cell infiltration and survival with PD1 blockade.
a, 9p21.3 loss is enriched in immune infiltrated samples (two-sided Fisher’s exact test and BenjaminiHochberg method for FDR correction, q < 0.05) and associated with worse PFS (two-sided log-rank test, p < 0.05) following anti-PD-1 therapy (n = 57 infiltrated patients). b, Within infiltrated tumors, del(9p21.3) is associated with worse response to PD-1 blockade (two-sided Fisher’s exact test for clinical benefit vs. no clinical benefit, p < 0.05. Error bars are SEM and measure of center is mean). c, del(9p21.3) is associated with worse PFS following PD-1 blockade (left panel) but not mTOR inhibition (right panel) (two-sided log-rank test). d, There were no chromosomal amplifications that were enriched in infiltrated tumors (two-sided Fisher’s exact test and Benjamini Hochberg method for FDR correction, vertical dotted line at q = 0.05) and significantly associated with improved or worsened PFS or OS (two-sided log-rank test, horizontal dotted line at p = 0.05) (n = 57 infiltrated patients).
Extended Data Fig. 10 GSEA of 9p21.3 deleted tumors versus wildtype using the Hallmark gene sets.
Several pathways including MTORC signaling, angiogenesis, hypoxia, and glycolysis were enriched in del(9p21.3) tumors (blue dots. Dotted lines indicate GSEA FDR q-values of 0.25 (dark) and 0.05 (light) (two-sided weighted KolmogorovSmirnov-like statistic test, n = 259 del(9p21.3) tumors and n = 158 WT tumors).
Supplementary information
Supplementary Tables
Supplementary Tables 1–4
Rights and permissions
About this article
Cite this article
Braun, D.A., Hou, Y., Bakouny, Z. et al. Interplay of somatic alterations and immune infiltration modulates response to PD-1 blockade in advanced clear cell renal cell carcinoma. Nat Med 26, 909–918 (2020). https://doi.org/10.1038/s41591-020-0839-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-020-0839-y
This article is cited by
-
Tumor immune dysfunction and exclusion subtypes in bladder cancer and pan-cancer: a novel molecular subtyping strategy and immunotherapeutic prediction model
Journal of Translational Medicine (2024)
-
Cancer immunometabolism: advent, challenges, and perspective
Molecular Cancer (2024)
-
Quantified pathway mutations associate epithelial-mesenchymal transition and immune escape with poor prognosis and immunotherapy resistance of head and neck squamous cell carcinoma
BMC Medical Genomics (2024)
-
A promising natural killer cell-based model and a nomogram for the prognostic prediction of clear-cell renal cell carcinoma
European Journal of Medical Research (2024)
-
Development and validation of a disulfidptosis and disulfide metabolism-related risk index for predicting prognosis in lung adenocarcinoma
Cancer Cell International (2024)