Abstract
Viruses are implicated in autoimmune destruction of pancreatic islet β cells, which results in insulin deficiency and type 1 diabetes (T1D)1,2,3,4. Certain enteroviruses can infect β cells in vitro5, have been detected in the pancreatic islets of patients with T1D6 and have shown an association with T1D in meta-analyses4. However, establishing consistency in findings across studies has proven difficult. Obstacles to convincingly linking RNA viruses to islet autoimmunity may be attributed to rapid viral mutation rates, the cyclical periodicity of viruses7 and the selection of variants with altered pathogenicity and ability to spread in populations. β cells strongly express cell-surface coxsackie and adenovirus receptor (CXADR) genes, which can facilitate enterovirus infection8. Studies of human pancreata and cultured islets have shown significant variation in enteroviral virulence to β cells between serotypes and within the same serotype9,10. In this large-scale study of known eukaryotic DNA and RNA viruses in stools from children, we evaluated fecally shed viruses in relation to islet autoimmunity and T1D. This study showed that prolonged enterovirus B rather than independent, short-duration enterovirus B infections may be involved in the development of islet autoimmunity, but not T1D, in some young children. Furthermore, we found that fewer early-life human mastadenovirus C infections, as well as CXADR rs6517774, independently correlated with islet autoimmunity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
TEDDY virome sequencing data that support the findings of this study have been deposited in the NCBI database of Genotypes and Phenotypes (dbGaP) with the primary accession code phs001442, in accordance with the dbGaP controlled-access authorization process. Clinical metadata and virome results data analyzed for the current study will be made available in the NIDDK Central Repository at https://www.niddkrepository.org/studies/teddy.
Code availability
VirMAP was used to generate the virome data and has been deposited in GitHub (https://github.com/cmmr/virmap). All of the software code and dependencies are listed on the GitHub site. SAS 9.4 (SAS Institute) was used for the statistical analysis, and GraphPad Prism 8.0 was used to create the figures.
References
Van der Werf, N., Kroese, F. G. M., Rozing, J. & Hillebrands, J. L. Viral infections as potential triggers of type 1 diabetes. Diabetes Metab. Res. 23, 169–183 (2007).
Stene, L. C. et al. Enterovirus infection and progression from islet autoimmunity to type 1 diabetes: the Diabetes and Autoimmunity Study in the Young (DAISY). Diabetes 59, 3174–3180 (2010).
Laitinen, O. H. et al. Coxsackievirus B1 is associated with induction of β-cell autoimmunity that portends type 1 diabetes. Diabetes 63, 446–455 (2014).
Yeung, W. C., Rawlinson, W. D. & Craig, M. E. Enterovirus infection and type 1 diabetes mellitus: systematic review and meta-analysis of observational molecular studies. Br. Med. J. 342, d35 (2011).
Anagandula, M. et al. Infection of human islets of Langerhans with two strains of coxsackie B virus serotype 1: assessment of virus replication, degree of cell death and induction of genes involved in the innate immunity pathway. J. Med. Virol. 86, 1402–1411 (2014).
Krogvold, L. et al. Detection of a low-grade enteroviral infection in the islets of Langerhans of living patients newly diagnosed with type 1 diabetes. Diabetes 64, 1682–1687 (2015).
Abedi, G. R. et al. Enterovirus and human parechovirus surveillance—United States, 2009–2013. Morb. Mortal. Weekly Rep. 64, 940–943 (2015).
Ifie, E. et al. Unexpected subcellular distribution of a specific isoform of the coxsackie and adenovirus receptor, CAR-SIV, in human pancreatic beta cells. Diabetologia 61, 2344–2355 (2018).
Roivainen, M. et al. Mechanisms of coxsackievirus-induced damage to human pancreatic beta-cells. J. Clin. Endocrinol. Metab. 85, 432–440 (2000).
Roivainen, M. et al. Functional impairment and killing of human beta cells by enteroviruses: the capacity is shared by a wide range of serotypes, but the extent is a characteristic of individual virus strains. Diabetologia 45, 693–702 (2002).
Ajami, N. J., Wong, M. C., Ross, M. C., Lloyd, R. E. & Petrosino, J. F. Maximal viral information recovery from sequence data using VirMAP. Nat. Commun. 9, 3205 (2018).
Laassri, M. et al. Evolution of echovirus 11 in a chronically infected immunodeficient patient. PLoS Pathog. 14, e1006943 (2018).
Schneider, V. A. et al. Evaluation of GRCh38 and de novo haploid genome assemblies demonstrates the enduring quality of the reference assembly. Genome Res. 27, 849–864 (2017).
Tapia, G. et al. Human enterovirus RNA in monthly fecal samples and islet autoimmunity in Norwegian children with high genetic risk for type 1 diabetes: the MIDIA study. Diabetes Care 34, 151–155 (2011).
Honkanen, H. et al. Detection of enteroviruses in stools precedes islet autoimmunity by several months: possible evidence for slowly operating mechanisms in virus-induced autoimmunity. Diabetologia 60, 424–431 (2017).
Simonen-Tikka, M. L. et al. Human enterovirus infections in children at increased risk for type 1 diabetes: the Babydiet study. Diabetologia 54, 2995–3002 (2011).
Oikarinen, S. et al. Enterovirus RNA in blood is linked to the development of type 1 diabetes. Diabetes 60, 276–279 (2011).
Salminen, K. et al. Enterovirus infections are associated with the induction of beta-cell autoimmunity in a prospective birth cohort study. J. Med. Virol. 69, 91–98 (2003).
Cinek, O. et al. Enterovirus RNA in longitudinal blood samples and risk of islet autoimmunity in children with a high genetic risk of type 1 diabetes: the MIDIA study. Diabetologia 57, 2193–2200 (2014).
Lee, H. S. et al. Next-generation sequencing for viruses in children with rapid-onset type 1 diabetes. Diabetologia 56, 1705–1711 (2013).
Sioofy-Khojine, A. B. et al. Coxsackievirus B1 infections are associated with the initiation of insulin-driven autoimmunity that progresses to type 1 diabetes. Diabetologia 61, 1193–1202 (2018).
Oikarinen, S. et al. Virus antibody survey in different European populations indicates risk association between coxsackievirus B1 and type 1 diabetes. Diabetes 63, 655–662 (2014).
Bessaud, M., Joffret, M. L., Blondel, B. & Delpeyroux, F. Exchanges of genomic domains between poliovirus and other cocirculating species C enteroviruses reveal a high degree of plasticity. Sci. Rep. 6, 38831 (2016).
Hamalainen, S. et al. Coxsackievirus B1 reveals strain specific differences in plasmacytoid dendritic cell mediated immunogenicity. J. Med. Virol. 86, 1412–1420 (2014).
Leveque, N. et al. Functional consequences of RNA 5′-terminal deletions on coxsackievirus B3 RNA replication and ribonucleoprotein complex formation. J. Virol. 91, e00423-17 (2017).
Chung, P. W., Huang, Y. C., Chang, L. Y., Lin, T. Y. & Ning, H. C. Duration of enterovirus shedding in stool. J. Microbiol. Immunol. Infect. 34, 167–170 (2001).
Alexander, J. P. Jr., Gary, H. E. Jr. & Pallansch, M. A. Duration of poliovirus excretion and its implications for acute flaccid paralysis surveillance: a review of the literature. J. Infect. Dis. 175, S176–S182 (1997).
Melnick, J. L. & Rennick, V. Infectivity titers of enterovirus as found in human stools. J. Med. Virol. 5, 205–220 (1980).
Ylipaasto, P. et al. Enterovirus infection in human pancreatic islet cells, islet tropism in vivo and receptor involvement in cultured islet beta cells. Diabetologia 47, 225–239 (2004).
Richardson, S. J., Leete, P., Bone, A. J., Foulis, A. K. & Morgan, N. G. Expression of the enteroviral capsid protein VP1 in the islet cells of patients with type 1 diabetes is associated with induction of protein kinase R and downregulation of Mcl-1. Diabetologia 56, 185–193 (2013).
Busse, N. et al. Detection and localization of viral infection in the pancreas of patients with type 1 diabetes using short fluorescently-labelled oligonucleotide probes. Oncotarget 8, 12620–12636 (2017).
Hodik, M. et al. Coxsackie–adenovirus receptor expression is enhanced in pancreas from patients with type 1 diabetes. BMJ Open Diabetes Res. Care 4, e000219 (2016).
Shafren, D. R. et al. Coxsackieviruses B1, B3, and B5 use decay accelerating factor as a receptor for cell attachment. J. Virol. 69, 3873–3877 (1995).
Ito, M. et al. Expression of coxsackievirus and adenovirus receptor in hearts of rats with experimental autoimmune myocarditis. Circ. Res. 86, 275–280 (2000).
Garnett, C. T. et al. Latent species C adenoviruses in human tonsil tissues. J. Virol. 83, 2417–2428 (2009).
Wang, Z. et al. Broad spectrum respiratory pathogen analysis of throat swabs from military recruits reveals interference between rhinoviruses and adenoviruses. Microb. Ecol. 59, 623–634 (2010).
Ingle, H. et al. Viral complementation of immunodeficiency confers protection against enteric pathogens via interferon-λ. Nat. Microbiol. 4, 1120–1128 (2019).
Messacar, K., Abzug, M. J. & Dominguez, S. R. 2014 outbreak of enterovirus D68 in North America. J. Med. Virol. 88, 739–745 (2016).
Greninger, A. L. et al. A novel outbreak enterovirus D68 strain associated with acute flaccid myelitis cases in the USA (2012–14): a retrospective cohort study. Lancet Infect. Dis. 15, 671–682 (2015).
Krischer, J. P. et al. Genetic and environmental interactions modify the risk of diabetes-related autoimmunity by 6 years of age: the TEDDY study. Diabetes Care 40, 1194–1202 (2017).
Hagopian, W. A. et al. The Environmental Determinants of Diabetes in the Young (TEDDY): genetic criteria and international diabetes risk screening of 421 000 infants. Pediatr. Diabetes 12, 733–743 (2011).
Parkes, M., Cortes, A., van Heel, D. A. & Brown, M. A. Genetic insights into common pathways and complex relationships among immune-mediated diseases. Nat. Rev. Genet. 14, 661–673 (2013).
Stewart, C. J. et al. Temporal development of the gut microbiome in early childhood from the TEDDY study. Nature 562, 583–588 (2018).
Vatanen, T. et al. The human gut microbiome in early-onset type 1 diabetes from the TEDDY study. Nature 562, 589–594 (2018).
Bonifacio, E. et al. Harmonization of glutamic acid decarboxylase and islet antigen-2 autoantibody assays for National Institute of Diabetes and Digestive and Kidney Diseases consortia. J. Clin. Endocrinol. Metab. 95, 3360–3367 (2010).
American Diabetes Association. Diagnosis and classification of diabetes mellitus. Diabetes Care 37, S81–S90 (2014).
Lee, H. S. et al. Biomarker discovery study design for type 1 diabetes in the Environmental Determinants of Diabetes in the Young (TEDDY) study. Diabetes Metab. Res. Rev. 30, 424–434 (2014).
TEDDY Study Group. The Environmental Determinants of Diabetes in the Young (TEDDY) study. Ann. NY Acad. Sci. 1150, 1–13.
TEDDY Study Group. The Environmental Determinants of Diabetes in the Young (TEDDY) study: study design. Pediatr. Diabetes 8, 286–298 (2007).
Vehik, K. et al. Methods, quality control and specimen management in an international multicentre investigation of type 1 diabetes: TEDDY. Diabetes Metab. Res. Rev. 29, 557–567 (2013).
Clem, A. L., Sims, J., Telang, S., Eaton, J. W. & Chesney, J. Virus detection and identification using random multiplex (RT)-PCR with 3′-locked random primers. Virol. J. 4, 65 (2007).
Bushnell, B. BBMap Short Read Aligner (Univ. California, 2016).
Sioofy-Khojine, A. B. et al. Molecular epidemiology of enteroviruses in young children at increased risk of type 1 diabetes. PLoS ONE 13, e0201959 (2018).
Krischer, J. P. et al. The influence of type 1 diabetes genetic susceptibility regions, age, sex, and family history on the progression from multiple autoantibodies to type 1 diabetes: a TEDDY study report. Diabetes 66, 3122–3129 (2017).
Handbook of Modern Statistical Methods (Taylor & Francis Group, 2018).
Uusitalo, U. et al. Association of early exposure of probiotics and islet autoimmunity in the TEDDY study. JAMA Pediatr. 170, 20–28 (2016).
Acknowledgements
The TEDDY Study is funded by U01 DK63829, U01 DK63861, U01 DK63821, U01 DK63865, U01 DK63863, U01 DK63836, U01 DK63790, UC4 DK63829, UC4 DK63861, UC4 DK63821, UC4 DK63865, UC4 DK63863, UC4 DK63836, UC4 DK95300, UC4 DK100238, UC4 DK106955, UC4 DK112243, UC4 DK117483 and contract number HHSN267200700014C from the NIDDK, National Institute of Allergy and Infectious Diseases, National Institute of Child Health and Human Development, National Institute of Environmental Health Sciences, Centers for Disease Control and Prevention and JDRF. This work was supported in part by the NIH/NCATS Clinical and Translational Science Awards to the University of Florida (UL1 TR000064) and the University of Colorado (UL1 TR001082).
Author information
Authors and Affiliations
Consortia
Contributions
K.V., K.F.L., M.R., J.T., A.G.Z., J.-X.S., A.L., B.A., W.A.H., D.A.S., J.P.K., H.H. and R.E.L. designed the study. M.R., W.A.H., J.T., A.G.Z., J.-X.S., B.A., A.L., H.H., K.V., K.F.L. and J.P.K. participated in patient recruitment and diagnosis, sample collection, and generation of the metadata. J.F.P., R.E.L., N.J.A., M.C.W., M.C.R., X.T. and R.A.G. generated and processed the raw sequencing data. K.F.L., K.V., H.H. and R.E.L. performed the data analysis, data interpretation and figure generation. K.V., K.F.L., H.H. and R.E.L. wrote the paper. All authors contributed to critical revisions and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
H.H. is a shareholder and chairman of the board of Vactech, and a member of the Scientific Advisory Board of Provention Bio, which develops vaccines against picornaviruses and CVB. The authors have no other relevant affiliations, nor financial involvement with any organization or entity with a financial interest in or financial conflict with the subject matter or materials discussed in the manuscript apart from those disclosed.
Additional information
Peer review information Jennifer Sargent was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Percentage of stool samples at age of first appearance of Enterovirus B.
Panel a shows sample positivity and Panel b sample consecutive positivity. Panels c and d show months prior to autoantibody seroconversion of Enterovirus B for sample positivity (c) and sample consecutive positivity (d) by autoantibody case status (n=383 matched pair children). Blue line represents control samples and red line represents case samples. The timing of the first appearance of an Enterovirus B infection from enrollment (3 months of age) or months prior to islet autoimmunity showed no obvious trend by age of child.
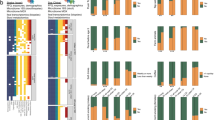
Extended Data Fig. 2 Common human viruses related to type 1 diabetes (T1D).
The three forest plots (a-c) show how common human viruses relate to the odds of children being diagnosed with T1D. The results were shown as odds ratios (OR, circle) and 95% confidence intervals (CI, bars) and were calculated using conditional logistic regression models with adjustment for HLA-DR-DQ genotype. OR>1 indicated a positive correlation between virus pattern and diagnosis with T1D, OR<1 indicated an inverse correlation. Plot (a) examined if an increase in the number of samples positive for virus was correlated with T1D (n=112 matched pair children). Plot (b) examined if children positive for the virus between 3 and 6 months of age were related to T1D (n=103 matched pair children). Plot (c) examined if children positive for the common virus in at least two consecutive samples (yes versus no) were related to T1D (n=112 matched pair children). Black circles and CI bars represent non-significant associations. Red circles and CI bars represent significant association with T1D. The number of positive stool samples for Enterovirus B was lower among T1D cases compared to matched controls. Human mastadenovirus C, similar to islet autoimmunity cases, was less likely to be detected in early stool samples (3–6 months of age) compared to the matched control for T1D cases. All p-values were two-sided.
Extended Data Fig. 3 Heatmaps of contig alignments of successive stools (n=6 children).
Heatmaps showing percent homology of alignments of enterovirus contigs isolated from successive stools from the same child. Stool collection date (successive days in the study) are shown, the serotype for the enterovirus aligned, all are aligned to an enterovirus genome map with scale of nucleotides at the bottom. Heatmap color is assigned on ~7 nt/pixel, heatmap color scale of percent homology is shown at the top.
Extended Data Fig. 4 Children consecutive positive for Enterovirus B with prolonged shedding of same serotype.
Categorical months of shedding by number of children for islet autoimmunity cases (n=45) and controls (n=25). Red bars denote cases and blue bars denote controls. Length of prolonged shedding period (duration) was not associated with case status in the children with consecutive positive Enterovirus B. Conditional logistic regression was used to evaluate significance; test was two-sided.
Supplementary information
Supplementary Information
Supplementary Note
Supplementary Tables
Supplementary Tables 1–9.
Source data
Source Data Fig. 1
Statistical Source Data
Source Data Fig. 2
Statistical Source Data
Source Data Fig. 3
Statistical Source Data
Source Data Fig. 4
Statistical Source Data
Source Data Extended Data Fig. 1
Statistical Source Data
Source Data Extended Data Fig. 2
Statistical Source Data
Source Data Extended Data Fig. 4
Statistical Source Data
Rights and permissions
About this article
Cite this article
Vehik, K., Lynch, K.F., Wong, M.C. et al. Prospective virome analyses in young children at increased genetic risk for type 1 diabetes. Nat Med 25, 1865–1872 (2019). https://doi.org/10.1038/s41591-019-0667-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-019-0667-0
This article is cited by
-
imply: improving cell-type deconvolution accuracy using personalized reference profiles
Genome Medicine (2024)
-
Dietary emulsifier consumption accelerates type 1 diabetes development in NOD mice
npj Biofilms and Microbiomes (2024)
-
Exploring the Human Virome: Composition, Dynamics, and Implications for Health and Disease
Current Microbiology (2024)
-
Safety, tolerability and immunogenicity of PRV-101, a multivalent vaccine targeting coxsackie B viruses (CVBs) associated with type 1 diabetes: a double-blind randomised placebo-controlled Phase I trial
Diabetologia (2024)
-
Pathobionts from chemically disrupted gut microbiota induce insulin-dependent diabetes in mice
Microbiome (2023)