Abstract
Cancer treatments have evolved from indiscriminate cytotoxic agents to selective genome- and immune-targeted drugs that have transformed the outcomes of some malignancies1. Tumor complexity and heterogeneity suggest that the ‘precision medicine’ paradigm of cancer therapy requires treatment to be personalized to the individual patient2,3,4,5,6. To date, precision oncology trials have been based on molecular matching with predetermined monotherapies7,8,9,10,11,12,13,14. Several of these trials have been hindered by very low matching rates, often in the 5–10% range15, and low response rates. Low matching rates may be due to the use of limited gene panels, restrictive molecular matching algorithms, lack of drug availability, or the deterioration and death of end-stage patients before therapy can be implemented. We hypothesized that personalized treatment with combination therapies would improve outcomes in patients with refractory malignancies. As a first test of this concept, we implemented a cross-institutional prospective study (I-PREDICT, NCT02534675) that used tumor DNA sequencing and timely recommendations for individualized treatment with combination therapies. We found that administration of customized multidrug regimens was feasible, with 49% of consented patients receiving personalized treatment. Targeting of a larger fraction of identified molecular alterations, yielding a higher ‘matching score’, was correlated with significantly improved disease control rates, as well as longer progression-free and overall survival rates, compared to targeting of fewer somatic alterations. Our findings suggest that the current clinical trial paradigm for precision oncology, which pairs one driver mutation with one drug, may be optimized by treating molecularly complex and heterogeneous cancers with combinations of customized agents.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Kato, S., Subbiah, V. & Kurzrock, R. Counterpoint: successes in the pursuit of precision medicine: biomarkers take credit. J. Natl Compr. Canc. Netw. 15, 863–866 (2017).
Gerlinger, M. et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 366, 883–892 (2012).
Hoadley, K. A. et al. Multiplatform analysis of 12 cancer types reveals molecular classification within and across tissues of origin. Cell 158, 929–944 (2014).
Ciriello, G. et al. Emerging landscape of oncogenic signatures across human cancers. Nat. Genet. 45, 1127–1133 (2013).
Kandoth, C. et al. Mutational landscape and significance across 12 major cancer types. Nature 502, 333–339 (2013).
Wheler, J., Lee, J. J. & Kurzrock, R. Unique molecular landscapes in cancer: implications for individualized, curated drug combinations. Cancer Res. 74, 7181–7184 (2014).
Von Hoff, D. D. et al. Pilot study using molecular profiling of patients’ tumors to find potential targets and select treatments for their refractory cancers. J. Clin. Oncol. 28, 4877–4883 (2010).
Tsimberidou, A. M. et al. Personalized medicine in a phase I clinical trials program: the MD Anderson Cancer Center initiative. Clin. Cancer Res. 18, 6373–6383 (2012).
Schwaederle, M. et al. Precision oncology: the UC San Diego Moores Cancer Center PREDICT experience. Mol. Cancer Ther. 15, 743–752 (2016).
Le Tourneau, C. et al. Molecularly targeted therapy based on tumour molecular profiling versus conventional therapy for advanced cancer (SHIVA): a multicentre, open-label, proof-of-concept, randomised, controlled phase 2 trial. Lancet Oncol. 16, 1324–1334 (2015).
Wheler, J. J. et al. Cancer therapy directed by comprehensive genomic profiling: a single center study. Cancer Res. 76, 3690–3701 (2016).
Chen, A. P. et al. Feasibility of molecular profiling based assignment of cancer treatment (MPACT): a randomized NCI precision medicine study. J. Clin. Oncol. 34, 2539 (2016).
Tsimberidou, A. M. et al. Initiative for molecular profiling and advanced cancer therapy (IMPACT): an MD Anderson precision medicine study. JCO Precis. Oncol. https://doi.org/10.1200/PO.17.00002 (2017).
Massard, C. et al. High-throughput genomics and clinical outcome in hard-to-treat advanced cancers: results of the MOSCATO 01 trial. Cancer Discov. 7, 586–595 (2017).
Prasad, V. Perspective: the precision-oncology illusion. Nature 537, S63 (2016).
Patel, S. P. & Kurzrock, R. PD-L1 expression as a predictive biomarker in cancer immunotherapy. Mol. Cancer Ther. 14, 847–856 (2015).
Goodman, A. M. et al. Tumor mutational burden as an independent predictor of response to immunotherapy in diverse cancers. Mol. Cancer Ther. 16, 2598–2608 (2017).
Le, D. T. et al. PD-1 blockade in tumors with mismatch-repair deficiency. N. Engl. J. Med. 372, 2509–2520 (2015).
Mazumdar, M. & Glassman, J. R. Categorizing a prognostic variable: review of methods, code for easy implementation and applications to decision-making about cancer treatments. Stat. Med. 19, 113–132 (2000).
Goel, M. K., Khanna, P. & Kishore, J. Understanding survival analysis: Kaplan–Meier estimate. Int. J. Ayurveda Res. 1, 274–278 (2010).
Nikanjam, M., Patel, H. & Kurzrock, R. Dosing immunotherapy combinations: analysis of 3,526 patients for toxicity and response patterns. Oncoimmunology 6, e1338997 (2017).
Liu, S., Nikanjam, M. & Kurzrock, R. Dosing de novo combinations of two targeted drugs: towards a customized precision medicine approach to advanced cancers. Oncotarget 7, 11310–11320 (2016).
Nikanjam, M., Liu, S., Yang, J. & Kurzrock, R. Dosing three-drug combinations that include targeted anti-cancer agents: analysis of 37,763 pPatients. Oncologist 22, 576–584 (2017).
Nikanjam, M., Liu, S. & Kurzrock, R. Dosing targeted and cytotoxic two-drug combinations: lessons learned from analysis of 24,326 patients reported 2010 through 2013. Int. J. Cancer 139, 2135–2141 (2016).
Wood, K., Hensing, T., Malik, R. & Salgia, R. Prognostic and predictive value in KRAS in non-small-cell lung cancer: a review. JAMA Oncol. 2, 805–812 (2016).
Jacobsen, E., Shanmugam, V. & Jagannathan, J. Rosai–Dorfman disease with activating KRAS mutation: response to cobimetinib. N. Engl. J. Med. 377, 2398–2399 (2017).
Wheler, J. J. et al. TP53 alterations correlate with response to VEGF/VEGFR inhibitors: implications for targeted therapeutics. Mol. Cancer Ther. 15, 2475–2485 (2016).
Koehler, K., Liebner, D. & Chen, J. L. TP53 mutational status is predictive of pazopanib response in advanced sarcomas. Ann. Oncol. 27, 539–543 (2016).
Sicklick, J. K. et al. Personalized, molecularly matched combination therapies for treatment-naïve, lethal malignancies: the I-PREDICT study. J. Clin. Oncol. 35, 2512 (2017).
Parker, B. A. et al. Breast cancer experience of the molecular tumor board at the University of California, San Diego Moores Cancer Center. J. Oncol. Pract. 11, 442–449 (2015).
Schwaederle, M. et al. Molecular tumor board: the University of California-San Diego Moores Cancer Center experience. Oncologist 19, 631–636 (2014).
Frampton, G. M. et al. Development and validation of a clinical cancer genomic profiling test based on massively parallel DNA sequencing. Nat. Biotechnol. 31, 1023–1031 (2013).
Stephens, P. et al. Analytic validation of a clinical circulating tumor DNA assay for patients with solid tumors. J. Clin. Oncol. 34, https://doi.org/10.1200/JCO.2016.34.15_suppl.e23049 (2017).
Hartmaier, R. J. et al. High-throughput genomic profiling of adult solid tumors reveals novel insights into cancer pathogenesis. Cancer Res. 77, 2464–2475 (2017).
Chalmers, Z. R. et al. Analysis of 100,000 human cancer genomes reveals the landscape of tumor mutational burden. Genome Med. 9, 34 (2017).
Bamford, S. et al. The COSMIC (Catalogue of Somatic Mutations in Cancer) database and website. Br. J. Cancer 91, 355–358 (2004).
Sun, J. X. et al. A computational approach to distinguish somatic vs. germline origin of genomic alterations from deep sequencing of cancer specimens without a matched normal. PLoS Comput. Biol. 14, e1005965 (2018).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Helsten, T. et al. Cell-cycle gene alterations in 4,864 tumors analyzed by next-generation sequencing: implications for targeted therapeutics. Mol. Cancer Ther. 15, 1682–1690 (2016).
Kato, S. et al. Cyclin-dependent kinase pathway aberrations in diverse malignancies: clinical and molecular characteristics. Cell Cycle 14, 1252–1259 (2015).
Said, R. et al. P53 mutations in advanced cancers: clinical characteristics, outcomes, and correlation between progression-free survival and bevacizumab-containing therapy. Oncotarget 4, 705–714 (2013).
Schwaederle, M. et al. VEGF-A expression correlates with TP53 mutations in non-small cell lung cancer: implications for antiangiogenesis therapy. Cancer Res. 75, 1187–1190 (2015).
Goodman, A. M. et al. Prevalence of PDL1 amplification and preliminary response to immune checkpoint blockade in solid tumors. JAMA Oncol. 4, 1237–1244 (2018).
Jia, J. et al. Correlation of tumor mutational burden and predicted functional impact of mutations across cancer types. J. Clin. Oncol. 36, https://doi.org/10.1200/JCO.2018.36.15_suppl.e24296 (2018).
Eisenhauer, E. A. et al. New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). Eur. J. Cancer 45, 228–247 (2009).
Dhani, N., Tu, D., Sargent, D. J., Seymour, L. & Moore, M. J. Alternate endpoints for screening phase II studies. Clin. Cancer Res. 15, 1873–1882 (2009).
Acknowledgements
We thank J. Wrolstad, M. Scur, and J. Porter for their critical roles in clinical trial and data management, L. Marquez for medication acquisition, D. Sandler, C. M. Tang, H. Yoon, M. Yerba and S. Silverman for assistance with manuscript preparation, T. Reya for her critical review of the manuscript, and most importantly, the patients and their families. This work was supported in part by Foundation Medicine, Inc. (J.K.S., R.O., B.L.-J., R.K.), as well as the Joan and Irwin Jacobs Philanthropic Fund (R.K.), the Jon Schneider Memorial Cancer Research Fund (J.K.S., P.T.F.), and National Institutes of Health (NIH) grant nos. P30CA023100 (J.K.S., S.M.L., R.K.) and P30CA016672 (J.J.L.). The authors also acknowledge the support of NIH grant nos. K08CA168999 and R21CA192072, as well as Pedal the Cause, The David Foundation, and Kristen Ann Carr Fund (J.K.S.).
Author information
Authors and Affiliations
Contributions
J.K.S. contributed to the conception/design of the work, the acquisition, analysis, and interpretation of the data, and the drafting and substantive revision of the work. S.K. and R.O. contributed to the acquisition, analysis, and interpretation of the data, and the drafting and substantive revision of the work. M.S. contributed to the acquisition, analysis, and interpretation of the data, and the drafting of the work. M.E.H., C.B.W., P.D., and P.T.F. contributed to the acquisition and analysis of data. A.K. and D.E.P. contributed to the acquisition of the data. V.A.M. contributed to the design of the work, the drafting of the work, and the substantive revision of the work. J.S.R. contributed to the conception/design of the work, the acquisition, analysis, and interpretation of the data, and the drafting of the work. A.B., J.W., and P.J.S. contributed to the acquisition, analysis, and interpretation of the data. J.J.L. contributed to the design of the work, the analysis and interpretation of the data, and the drafting of the work. S.M.L. contributed to the conception/design of the work. B.L.-J. contributed to the acquisition, analysis, and interpretation of data, and the drafting of the work. R.K. contributed to the conception/design of the work, the acquisition, analysis, and interpretation of the data, and the drafting and substantive revision of the work. All authors approved the submitted version (and any substantially modified version that involves the authors’ contribution to the study). All authors have agreed to be personally accountable for their own contributions and to ensure that questions related to the accuracy or integrity of any part of the work, even ones where the author was not personally involved, are appropriately investigated, resolved, and the resolution documented in the literature.
Corresponding authors
Ethics declarations
Competing interests
J.K.S. receives research funds from Foundation Medicine, Inc., Novartis Pharmaceuticals, Blueprint Medicines, and Amgen, as well as consultant fees from Loxo Oncology, Biotheranostics, and Grand Rounds. M.E.H. receives research funds from General Electric and is an equity holder in Illumina, Inc. C.W. receives research support from Takeda, TESARO, and Pfizer, as well as consultant fees from Takeda. V.A.M., J.S.R., and J.W. are employees and equity holders in Foundation Medicine, Inc. V.A.M. is also on the Board of Directors for Revolution Medicines (equity and compensation >$10,000) and has patent royalties for EGFR T790M testing issued to the Memorial Sloan Kettering Cancer Center. A.B. is formerly an employee and is an equity holder in Foundation Medicine, Inc. He is now an employee of Scipher Medicine. P.J.S. is formerly an employee and is formerly an equity holder in Foundation Medicine, Inc. He is now an employee of GRAIL, Inc. J.J.L. served on the Statistical Advisory Board of AbbVie. R.K. has research funding from Incyte, Genentech, Merck Serono, Pfizer, Sequenom, Grifols, OmniSeq, Foundation Medicine, Inc., Guardant Health, and Konica Minolta, as well as consultant fees from Loxo Oncology, Actuate Therapeutics, Roche, XBiotech, and NeoMed. She serves as an advisor to Soluventis. She receives speaker fees from Roche, and has an equity interest in IDbyDNA, CureMatch, Inc., and Soluventis. All other authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
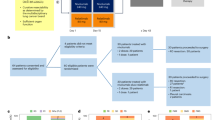
Extended Data Fig. 1 Consolidated Standards of Reporting Trials (CONSORT) diagram, which includes the 149 patients that consented to I-PREDICT.
*Treated evaluable patients includes patients who received >10 d of treatment for drugs given on a daily basis (generally drugs given by mouth) or at least two doses of a drug normally given every two weeks or more frequently (the latter generally being intravenous drugs). Only patients whose treatment was reviewed and validated by data analysis lockdown are included. **One patient had inadequate tissue for NGS and declined biopsy; he was later reenrolled after he agreed to undergo biopsy. One treated patient who initially was believed to have prior therapy was found, after data lockdown analysis, to have not received the prior regimen.
Supplementary information
Supplementary Information
Supplementary Text, Supplementary Tables 3–10, and Supplementary References
Supplementary Tables
Supplementary Tables 1 and 2
Rights and permissions
About this article
Cite this article
Sicklick, J.K., Kato, S., Okamura, R. et al. Molecular profiling of cancer patients enables personalized combination therapy: the I-PREDICT study. Nat Med 25, 744–750 (2019). https://doi.org/10.1038/s41591-019-0407-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-019-0407-5
This article is cited by
-
Heterogeneity and molecular landscape of melanoma: implications for targeted therapy
Molecular Biomedicine (2024)
-
The molecular interaction pattern of lenvatinib enables inhibition of wild-type or kinase-mutated FGFR2-driven cholangiocarcinoma
Nature Communications (2024)
-
Patient derived tumoroids of high grade neuroendocrine neoplasms for more personalized therapies
npj Precision Oncology (2024)
-
Adaptive Darwinian off-target resistance mechanisms to selective RET inhibition in RET driven cancer
npj Precision Oncology (2024)
-
Engineering biomimetic nanosystem targeting multiple tumor radioresistance hallmarks for enhanced radiotherapy
Science China Life Sciences (2024)