Abstract
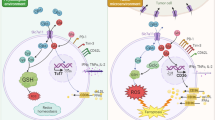
Cancer cells can evade immune surveillance through the expression of inhibitory ligands that bind their cognate receptors on immune effector cells. Expression of programmed death ligand 1 in tumor microenvironments is a major immune checkpoint for tumor-specific T cell responses as it binds to programmed cell death protein-1 on activated and dysfunctional T cells1. The activity of myeloid cells such as macrophages and neutrophils is likewise regulated by a balance between stimulatory and inhibitory signals. In particular, cell surface expression of the CD47 protein creates a ‘don’t eat me’ signal on tumor cells by binding to SIRPα expressed on myeloid cells2,3,4,5. Using a haploid genetic screen, we here identify glutaminyl-peptide cyclotransferase-like protein (QPCTL) as a major component of the CD47-SIRPα checkpoint. Biochemical analysis demonstrates that QPCTL is critical for pyroglutamate formation on CD47 at the SIRPα binding site shortly after biosynthesis. Genetic and pharmacological interference with QPCTL activity enhances antibody-dependent cellular phagocytosis and cellular cytotoxicity of tumor cells. Furthermore, interference with QPCTL expression leads to a major increase in neutrophil-mediated killing of tumor cells in vivo. These data identify QPCTL as a novel target to interfere with the CD47 pathway and thereby augment antibody therapy of cancer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All sequencing data sets have been deposited in the National Center for Biotechnology Information Sequence Read Archive under accession numbers SRP144590 and PRJNA505977. In addition, all processed screen results are accessible in an interactive database (https://phenosaurus.nki.nl/). All data presented in this manuscript are available from the corresponding authors upon reasonable request
References
Zou, W., Wolchok, J. D. & Chen, L. PD-L1 (B7-H1) and PD-1 pathway blockade for cancer therapy: Mechanisms, response biomarkers, and combinations. Sci. Transl. Med. 8, 328rv4–328rv4 (2016).
Jaiswal, S. et al. CD47 is upregulated on circulating hematopoietic stem cells and leukemia cells to avoid phagocytosis. Cell 138, 271–285 (2009).
Majeti, R. et al. CD47 is an adverse prognostic factor and therapeutic antibody target on human acute myeloid leukemia stem cells. Cell 138, 286–299 (2009).
Weiskopf, K. Cancer immunotherapy targeting the CD47/SIRPα axis. Eur. J. Cancer 76, 100–109 (2017).
Willingham, S. B. et al. The CD47-signal regulatory protein alpha (SIRPa) interaction is a therapeutic target for human solid tumors. Proc. Natl Acad. Sci. USA 109, 6662–6667 (2012).
Matlung, H. L., Szilagyi, K., Barclay, N. A. & van den Berg, T. K. The CD47-SIRPα signaling axis as an innate immune checkpoint in cancer. Immunol. Rev. 276, 145–164 (2017).
Zhao, X. W. et al. CD47-signal regulatory protein-α (SIRPα) interactions form a barrier for antibody-mediated tumor cell destruction. Proc. Natl Acad. Sci. USA 108, 18342–18347 (2011).
Chao, M. P. et al. Calreticulin is the dominant pro-phagocytic signal on multiple human cancers and is counterbalanced by CD47. Sci. Transl. Med. 2, 63ra94–63ra94 (2010).
Chen, J. et al. SLAMF7 is critical for phagocytosis of haematopoietic tumour cells via Mac-1 integrin. Nature 544, 493–497 (2017).
Chao, M. P. et al. Anti-CD47 antibody synergizes with rituximab to promote phagocytosis and eradicate non-Hodgkin lymphoma. Cell 142, 699–713 (2010).
Weiskopf, K. et al. Engineered SIRPα variants as immunotherapeutic adjuvants to anticancer antibodies. Science 341, 88–91 (2013).
Ring, N. G. et al. Anti-SIRPα antibody immunotherapy enhances neutrophil and macrophage antitumor activity. Proc. Natl Acad. Sci. USA 114, E10578–E10585 (2017).
Advani, R. et al. CD47 Blockade by Hu5F9-G4 and Rituximab in Non-Hodgkin’s Lymphoma. N. Engl. J. Med. 379, 1711–1721 (2018).
Casey, S. C. et al. MYC regulates the antitumor immune response through CD47 and PD-L1. Science 352, 227–231 (2016).
Seiffert, M. et al. Human signal-regulatory protein is expressed on normal, but not on subsets of leukemic myeloid cells and mediates cellular adhesion involving its counterreceptor CD47. Blood 94, 3633–3643 (1999).
Cynis, H. et al. Isolation of an isoenzyme of human glutaminyl cyclase: retention in the Golgi complex suggests involvement in the protein maturation machinery. J. Mol. Biol. 379, 966–980 (2008).
Schilling, S. et al. Identification of human glutaminyl cyclase as a metalloenzyme. Potent inhibition by imidazole derivatives and heterocyclic chelators. J. Biol. Chem. 278, 49773–49779 (2003).
Stephan, A. et al. Mammalian glutaminyl cyclases and their isoenzymes have identical enzymatic characteristics. FEBS. J. 276, 6522–6536 (2009).
Jimenez-Sanchez, M. et al. siRNA screen identifies QPCT as a druggable target for Huntington’s disease. Nat. Chem. Biol. 11, 347–354 (2015).
Saido, T. C. et al. Dominant and differential deposition of distinct β-amyloid peptide species, AβN3(pE), in senile plaques. Neuron 14, 457–466 (1995).
Tekirian, T. L., Yang, A. Y., Glabe, C. & Geddes, J. W. Toxicity of pyroglutaminated amyloid β-peptides 3(pE)-40 and -42 is similar to that of Aβ1-40 and -42. J. Neurochem 73, 1584–1589 (2002).
Russo, C. et al. Pyroglutamate-modified amyloid β-peptides - AβN3(pE) - strongly affect cultured neuron and astrocyte survival. J. Neurochem. 82, 1480–1489 (2002).
Hatherley, D. et al. Paired receptor specificity explained by structures of signal regulatory proteins alone and complexed with CD47. Mol. Cell 31, 266–277 (2008).
Ho, C. C. M. et al. ‘Velcro’ engineering of high affinity CD47 ectodomain as signal regulatory protein α (SIRPα) antagonists that enhance antibody-dependent cellular phagocytosis. J. Biol. Chem. 290, 12650–12663 (2015).
Brooke, G., Holbrook, J. D., Brown, M. H. & Barclay, A. N. Human lymphocytes interact directly with CD47 through a novel member of the signal regulatory protein (SIRP) family. J. Immunol. 173, 2562–2570 (2004).
Huang, K.-F., Wang, Y.-R., Chang, E.-C., Chou, T.-L. & Wang, A. H.-J. A conserved hydrogen-bond network in the catalytic centre of animal glutaminyl cyclases is critical for catalysis. Biochem. J. 411, 181–190 (2008).
Grizenkova, J. et al. Overexpression of the Hspa13 (Stch) gene reduces prion disease incubation time in mice. Proc. Natl Acad. Sci. USA 109, 13722–13727 (2012).
Schilling, S. et al. Glutaminyl cyclase inhibition attenuates pyroglutamate Abeta and Alzheimer’s disease-like pathology. Nat. Med. 14, 1106–1111 (2008).
Hoffmann, T. et al. Glutaminyl cyclase inhibitor PQ912 improves cognition in mouse models of Alzheimer’s disease-studies on relation to effective target occupancy. J. Pharmacol. Exp. Ther. 362, 119–130 (2017).
Lues, I. et al. A phase 1 study to evaluate the safety and pharmacokinetics of PQ912, a glutaminyl cyclase inhibitor, in healthy subjects. Alzheimers Dement. (NY) 1, 182–195 (2015).
Huang, K. F., Liu, Y. L., Cheng, W. J., Ko, T. P. & Wang, A. H. J. Crystal structures of human glutaminyl cyclase, an enzyme responsible for protein N-terminal pyroglutamate formation. Proc. Natl Acad. Sci. USA 102, 13117–13122 (2005).
Boross, P. et al. IgA EGFR antibodies mediate tumour killing in vivo. EMBO Mol. Med. 5, 1213–1226 (2013).
Liu, X. et al. CD47 blockade triggers T cell-mediated destruction of immunogenic tumors. Nat. Med. 21, 1209–1215 (2015).
Ingram, J. R. et al. Localized CD47 blockade enhances immunotherapy for murine melanoma. Proc. Natl Acad. Sci. USA 114, 10184–10189 (2017).
Manguso, R. T. et al. In vivo CRISPR screening identifies Ptpn2 as a cancer immunotherapy target. Nature 547, 413–418 (2017).
Sockolosky, J. T. et al. Durable antitumor responses to CD47 blockade require adaptive immune stimulation. Proc. Natl Acad. Sci. USA 113, E2646–E2654 (2016).
Ansell, S. et al. A phase 1 study of TTI-621, a novel immune checkpoint inhibitor targeting CD47, in patients with relapsed or refractory hematologic malignancies. Blood 128, 1812 (2016).
Scheltens, P. et al. Safety, tolerability and efficacy of the glutaminyl cyclase inhibitor PQ912 in Alzheimer’s disease: results of a randomized, double-blind, placebo-controlled phase 2a study. Alzheimers Res. Ther. 10, 107 (2018).
Morty, R. E., Bulau, P., Pellé, R., Wilk, S. & Abe, K. Pyroglutamyl peptidase type I from Trypanosoma brucei: a new virulence factor from African trypanosomes that de-blocks regulatory peptides in the plasma of infected hosts. Biochem. J. 394, 635–645 (2006).
Gong, Q. et al. A ultrasensitive near-infrared fluorescent probe reveals pyroglutamate aminopeptidase 1 can be a new inflammatory cytokine. Adv. Sci. (Weinh) 5, 1700664 (2018).
Blomen, V. A. et al. Gene essentiality and synthetic lethality in haploid human cells. Science 350, 1092–1096 (2015).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Carette, J. E. et al. Ebola virus entry requires the cholesterol transporter Niemann-Pick C1. Nature 477, 340–343 (2011).
Bracke, M., Lammers, J. W., Coffer, P. J. & Koenderman, L. Cytokine-induced inside-out activation of FcalphaR (CD89) is mediated by a single serine residue (S263) in the intracellular domain of the receptor. Blood 97, 3478–3483 (2001).
Berkovits, B. D. & Mayr, C. Alternative 3’ UTRs act as scaffolds to regulate membrane protein localization. Nature 522, 363–367 (2015).
Lackner, D. H. et al. A generic strategy for CRISPR-Cas9-mediated gene tagging. Nat. Commun. 6, 10237 (2015).
Brinkman, E. K., Chen, T., Amendola, M. & van Steensel, B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Res. 42, e168–e168 (2014).
Neefjes, J. J., Breur-Vriesendorp, B. S., van Seventer, G. A., Iványi, P. & Ploegh, H. L. An improved biochemical method for the analysis of HLA-class I antigens. Definition of new HLA-class I subtypes. Hum. Immunol. 16, 169–181 (1986).
Kuijpers, T. W. et al. Membrane surface antigen expression on neutrophils: a reappraisal of the use of surface markers for neutrophil activation. Blood 78, 1105–1111 (1991).
Brandsma, A. M. et al. Simultaneous targeting of FcγRs and FcαRI enhances tumor cell killing. Cancer Immunol. Res. 3, 1316–1324 (2015).
Meyer, S. et al. Improved in vivo anti-tumor effects of IgA-Her2 antibodies through half-life extension and serum exposure enhancement by FcRn targeting. Mabs 8, 87–98 (2016).
de Haij, S. et al. In vivo cytotoxicity of type I CD20 antibodies critically depends on Fc receptor ITAM signaling. Cancer Res. 70, 3209–3217 (2010).
Gee, M. H. et al. Antigen identification for orphan T cell receptors expressed on tumor-infiltrating lymphocytes. Cell 172, 549–563.e16 (2018).
Hadrup, S. R. et al. Parallel detection of antigen-specific T-cell responses by multidimensional encoding of MHC multimers. Nat. Methods 6, 520–526 (2009).
Acknowledgements
We thank R. Mezzadra, M. Wellenstein, C. Sun and members of the Schumacher, Brummelkamp, Scheeren, Leusen, van den Berg and Haanen laboratories for discussions, and O. van Tellingen, the Netherlands Cancer Institute–Antoni van Leeuwenhoek (NKI-AVL) Preclinical Intervention Unit and the NKI-AVL flow facility for technical support and input. This work was supported by European Research Council (ERC) advanced grant SENSIT (to T.N.S.), the Institute for Chemical Immunology (to T.N.S., J.N. and M.V.), ERC starting grant CHEMCHECK (to M.V.), Dutch Cancer Society/Koningin Wilhelmina Fonds kankerbestrijding (KWF) grant 10300 (to T.K.v.d.B.), KWF grant 11537 (to H.L.M.), a Leiden University Medical Center fellowship and the Errol McDowell Cancer Foundation (to F.A.S.), Netherlands Organization for Scientific Research (NWO) Vici Grant 016.Vici.170.033, KWF grant NKI 2015-7609, the Cancer Genomics Center (CGC.nl) and the Ammodo KNAW Award 2015 for Biomedical Sciences (to T.R.B.), and KWF grant UU 2015-7650 (to J.H.W.L.).
Author information
Authors and Affiliations
Contributions
M.E.W.L. conceived the project, designed and performed experiments, interpreted data and co-wrote the manuscript. M.R., A.F. and T.R.B. designed, performed and interpreted the haploid genetic screens. M.T. and J.N. designed, performed and interpreted biochemical data. J.H.M.J., A.M.B. and J.H.W.L. designed, performed and interpreted anti-Her2 in vitro and in vivo data, and J.H.W.L. co-wrote the manuscript. K.F., H.L.M. and T.K.v.d.B. designed, performed and interpreted in vitro data with human effector cells. S.v.d.S. supported and performed flow cytometry analyses. R.G.-E. and N.A.M.B. designed, performed and interpreted in vitro studies with human T cells. J.H.v.d.B. and J.B.A.G.H. supervised analyses of T cell reactivity. K.A.M. performed and interpreted experiments. M.V. designed experiments and provided reagents. F.A.S. and T.N.S. conceived the project, designed experiments, interpreted data and co-wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
M.E.W.L., M.R., T.R.B., F.A.S., J.H.W.L. and T.N.S. are inventors on a patent application that covers manipulation of the CD47-SIRPα axis via QPCTL. T.N.S. is advisor for Adaptive Biotechnologies, AIMM Therapeutics, Allogene Therapeutics, Amgen, Merus and Neon Therapeutics; is a recipient of grant or research support from MSD, Bristol-Myers Squibb and Merck KgaA; is a stockholder in AIMM Therapeutics, Allogene Therapeutics, Merus, Neogene Therapeutics and Neon Therapeutics; and is venture partner at Third Rock Ventures. T.R.B. is cofounder and Scientific Advisory Board member of Haplogen GmbH and cofounder and managing director of Scenic Biotech. J.H.W.L. is founder, advisor and shareholder of TigaTx. J.H.B. is recipient of grant or research support from Bristol-Myers Squibb, Medimmune and Neon Therapeutics. K.F., H.L.M. and T.K.v.d.B are recipients of research support from Synthon Biopharmaceuticals BV. J.B.A.G.H. is advisor to Bristol-Myers Squibb, MSD, Novartis, Roche/Genentech, Pfizer, IPSEN, AZ/MedImmune, Bayer, Seattle Genetics, Immunocore, Gadeta, Neon Therapeutics, and Celsius Therapeutics, and is a recipient of grant or research support from Bristol-Myers Squibb, MSD, Novartis and Neon Therapeutics.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 QPCTL deletion leads to reduced binding of recombinant SIRPα and SIRPγ.
a, Cell surface binding of αhCD47-2D3, αhCD47-B6H12, αhCD47-CC2C6 and hSIRPα-Fc to WT, CD47 and QPCTL KO lung cancer (A549) (upper left panel), colorectal cancer (DLD1) (upper right panel) and rectal carcinoma (RKO) (lower left panel) cells, as determined by flow cytometry. b, Cell surface binding of human SIRPα-Fc (hSIRPα-Fc) to lung cancer (A549) WT, CD47 KO and QPCTL KO cells alone or in combination with blocking antibody αhCD47-B6H12, as determined by flow cytometry. c, Cell surface binding of indicated concentrations of human SIRPα-His (hSIRPα-His) to WT, CD47 KO and QPCTL KO lung cancer (A549) cells, as determined by flow cytometry. d, Cell surface binding of indicated concentrations of hSIRPα-Fc and human SIRPγ-Fc (SIRPγ-Fc) to WT, CD47 KO and QPCTL KO lung cancer (A549) cells, as determined by flow cytometry. e, Representative plot (see d) of hSIRPγ-Fc binding (36.0 μg/ml) to WT, CD47 KO and QPCTL KO lung cancer (A549) cells. Values in a–d indicate MFI relative to WT cells stained with the same reagent (a), MFI relative to WT cells without blocking antibody (b) or MFI values (c,d). Data represent n = 3 biological replicates (a–e) and mean ± s.d. of triplicates (a–d). ***P≤0.0001 by one-way ANOVA with multiple comparison correction (a–c) or unpaired two-sided t-test (d); n.s., not significant. Data are representative of at least two (a–e) independent experiments. MFI, mean fluorescence intensity; WT, wild-type; KO, knock-out.
Extended Data Fig. 2 QPCTL regulates binding of αhCD47-CC2C6.
a, Cell surface binding of αhCD47-2D3 and αhCD47-CC2C6 to WT, QPCTL KO and QPCTL KO cells reconstituted with FLAG-tagged cDNA of QPCTL isoform 1 (OE var.1) or QPCTL isoform 2 (OE var.2) melanoma (A375) (a) epidermoid carcinoma (A431) (b) and lung cancer (A549) cells (c), as determined by flow cytometry. d, Western blot analysis of WT, QPCTL KO and QPCTL KO melanoma (A375) cells reconstituted with FLAG-tagged cDNA of QPCTL isoform 1 (OE var.1) or QPCTL isoform 2 (OE var.2). Blot image has been cropped to show the relevant bands, and molecular mass markers are indicated (in kD). See Source Data for the uncropped western blot. e, Cell surface binding of αhCD47-CC2C6 and αhCD47-2D3 to HAP1 QPCTL KO cells reconstituted with QPCTL var.1 or a catalytically inactive QPCTL variant (QPCLT var.1 D326E), as determined by flow cytometry. f, Cell surface binding of αhCD47-CC2C6 and αhCD47-2D3 to QPCTL KO melanoma (A375) cells reconstituted with QPCTL var.1 or QPCTL var.1 (D326E), as determined by flow cytometry. Values in a-c, e, f indicate MFI relative to WT cells stained with the same reagent. Data represent n = 3 biological replicates and mean ± s.d. of triplicates (a–c,e–f). Data are representative of at least two (a–c,d,e–f) independent experiments. OE, over-expression.
Extended Data Fig. 3 Two-dimensional genetic screen to reveal selective modifiers of αhCD47-CC2C6 binding.
a, Schematic overview of FACS-based haploid genetic screen on αhCD47-BH612 (‘B6H12’) and αhCD47-CC2C6 (‘CC2C6’) double-stained cells, employing a 4-way sorting strategy to distinguish regulators that affect CD47 levels or that affect the αhCD47-CC2C6/SIRPα binding site of CD47. b,c, Results of the genetic screen for general CD47 modulators (b) and αhCD47-CC2C6/SIRPα binding site regulators (c). Dots represent individual genes. The relative mutation frequency (MI) in the B6H12hi/CC2C6hi versus B6H12lo/CC2C6lo (b) and B6H12lo/CC2C6hi versus B6H12hi/CC2C6lo cell population (c) is plotted against the total number of insertions mapped per gene. Significantly enriched genes (FDR-corrected P < 0.05) are colored according to channel (B6H12hi/CC2Chi, green; B6H12lo/CC2Clo, dark blue; B6H12lo/CC2Chi, light blue; B6H12hi/CC2C6lo, orange) and selected regulators are labeled. n = 3,253,240 (b) and n = 3,209,992 insertions (c) were identified and data were analyzed by two-sided Fisher’s Exact test with multiple comparison correction. d, Combined results for 4-way-sort screen. Dots represent individual genes. x-axis shows relative mutation frequency (MI) for the B6H12hi/CC2C6hi versus B6H12lo/CC2C6lo cell population, y-axis shows relative mutation frequency for the B6H12lo/CC2C6hi versus B6H12hi/CC2C6lo cell population. Selected regulators are highlighted. e, Ratio of αhCD47-CC2C6 and αhCD47-BH612 cell surface binding to HAP1 WT, CD47 KO, QPCTL KO and HSPA13 KO cells, as determined by flow cytometry. Values indicate ratio of MFI of αhCD47-CC2C6/αhCD47-BH612 for each cell population. Data represent n = 3 biological replicates and mean ± s.d. of triplicates. ***P = 0.0001 by one-way ANOVA with multiple comparison correction. Data are representative of one (a–d) or two (e) independent experiments. MI, mutation index; WT, wild-type; KO, knockout; MFI, mean fluorescence intensity.
Extended Data Fig. 4 Glutaminyl cyclase inhibition leads to reduced binding of recombinant hSIRPα.
a, Cell surface binding of αhCD47-2D3, αhCD47-CC2C6 and hSIRPα-Fc to control (DMSO)-treated (-) or SEN177-treated (+) lung cancer (A549), colorectal (DLD1), HAP1, rectal carcinoma (RKO) and breast cancer (SKBR3) cells, as determined by flow cytometry. Data represent n = 3 biological replicates and mean ± s.d. of triplicates. ***P≤0.000695 by unpaired two-sided t-test. b, Cell surface binding of αhCD47-2D3, αhCD47-CC2C6 and hSIRPα-Fc to control (DMSO)-treated (-), SEN177-treated, and PQ912-treated melanoma (A375) cells, as determined by flow cytometry. Data represent n = 3 biological replicates and mean ± s.d. of triplicates. ***P = 0.0001 by one-way ANOVA with multiple comparison correction. c, Flow cytometry plot of surface binding of αhCD47-B6H12 and αhCD47-CC2C6 to control-treated and PQ912-treated melanoma (A375) cells. Data are representative of two independent experiments with similar results (n = 3 biological replicates per experiment). d, Cell surface binding of αhCD47-2D3, αhCD47-CC2C6 and hSIRPα-Fc to control (DMSO)-treated (-) and SEN177-treated (+) wild-type and QPCTL-knockout epidermoid carcinoma (A431) and lung cancer (A549) cells, as determined by flow cytometry. Data represent n = 3 biological replicates and mean ± s.d. of triplicates. e, Cell surface binding of secondary antibody alone, hSIRPα-Fc (followed by secondary antibody) or hSIRPα-Fc in the presence of the CD47 blocking antibody αhCD47-B6H12 (followed by secondary antibody) to control (DMSO)-treated (-) or SEN177-treated lung cancer (A549) cells, as determined by flow cytometry. Data represent n = 3 biological replicates and mean ± s.d. of triplicates. Values indicate MFI relative to WT cells stained with the same reagent (a,b,d) or MFI (e). Data are representative of one (e) or at least two independent experiments (a–d).
Extended Data Fig. 5 Analysis of phagocytosis.
a,b Representative images of gating strategy and phagocytosis (a) and examples of staining (b), as determined by ImageStream analysis. Data are representative of four independent experiments with similar results.
Extended Data Fig. 6 Synergy between blockade of CD47 pyroglutamate formation and tumors opsonization in tumor cell killing by macrophages and neutrophils.
a, MFI of lamin B-Turquoise of the total CD11b + macrophage population in samples incubated with control (DMSO)-treated (-) or SEN177-treated (+) Turquoise-expressing Burkitt’s lymphoma (Raji) cells in the presence or absence of the anti-human CD20 antibody Rituximab, CD47-blocking F(ab’)2 fragment B6H12, or SIRPα blocking antibody 12C4, as determined by ImageStream analysis. Symbols represent individual donors. Data represent mean ± s.d. of independent donors. ***P < 0.0001; **P = 0.0016; *P = 0.0256 by one-way ANOVA with multiple comparison correction. b, Specific lysis of control (DMSO)-treated (-) or SEN177-treated (+) WT, QPCTL KO or CD47 KO epidermoid carcinoma (A431) cells by human neutrophils in the presence or absence of the anti-human EGFR antibody cetuximab in a 4 h 51Cr-release assay. Data represent mean ± s.d. of independent donors. ***P < 0.0001; 0.0325≥*P≥0.0207 by one-way ANOVA with multiple comparison correction. n.s.; not significant. c, Flow cytometry plot of cell surface binding of anti-human CD20 antibody to Burkitt’s lymphoma (Raji) cells (left panel) and anti-human EGFR antibody to epidermoid carcinoma (A431) cells (right panel) treated with control (DMSO) or SEN177 for 4 days. Data are representative of one (c) or at least three independent experiments (a,b) representing 4 donors (for B6H12(Fab’)2 conditions), 8 donors (all other conditions) (a), and 8 donors (b).
Extended Data Fig. 7 QPCTL deficiency and QPCTL inhibition enhances tumor-specific antibody-induced killing of mouse tumor cells by mouse effector cells.
a, Cell surface binding of anti-mouse CD47 antibody MIAP301 (αmCD47-MIAP301) and mouse SIRPα-Fc (mSIRPα-Fc) to WT, CD47 KO and QPCTL bulk KO (KO#1 and KO#2) murine melanoma (B16F10) cells, and WT, CD47 KO and QPCTL KO (cl8 and cl30) Her2-expressing mouse pro-B (Ba/F3−Her2) cells, as determined by flow cytometry. b, Cell surface binding of αmCD47-MIAP301 and mSIRPα-Fc to WT, QPCTL KO or QPCTL KO murine melanoma (B16F10) cells reconstituted with the murine QPCTL cDNA (OE), as determined by flow cytometry. c, Cell surface binding of αmCD47-MIAP301 and mSIRPα-Fc to control (DMSO)-treated (-) or SEN177-treated ( + ) murine melanoma (B16F10) or Her2-expressing murine pro-B (Ba/F3-Her2) cells, as determined by flow cytometry. (a–c) Data represent n = 3 biological replicates and mean ± s.d. of triplicates. ***P≤0.0001 by one-way ANOVA with multiple comparison correction (a) or unpaired two-sided t-test (c). d, Flow cytometry plots of cell surface binding of anti-human Her2 antibody to WT, CD47 KO or QPCTL KO Ba/F3-Her2 cells (left), or control (DMSO)-treated or SEN177-treated Ba/F3-Her2 cells (right), as determined by flow cytometry. Data are representative of two independent experiments with similar results (n = 3 biological replicates per experiment) (left graph) or one experiment with two biological replicates (right graph). e, Specific lysis of control (DMSO)-treated (-) and SEN177-treated ( + ) CD47 KO and QPCTL KO murine pro-B cells (Ba/F3-Her2) by human neutrophils in the presence of anti-Her2 (IgA1) in a 4 h 51Cr-release assay. Data are representative of n = 3 biological replicates and represent ± s.d. of triplicates of one representative donor. f, Specific lysis of WT, CD47 KO and QPCTL KO murine pro-B cells (Ba/F3-Her2) by murine immune cells isolated from whole blood in the presence or absence of anti-Her2 (IgA1) in a 4 h 51Cr-release assay. Data are representative of n = 3 biological replicates and represent mean ± s.d. of triplicates of one representative donor. ***P≤0.0007 by one-way ANOVA with multiple comparison correction. g, Specific lysis of control (DMSO)-treated (-) or SEN177-treated ( + ) murine pro-B cells (Ba/F3-Her2) by murine immune cells isolated from whole blood in the presence or absence of anti-Her2 (IgA1) in a 4 h 51Cr-release assay. Data represent n = 3 biological replicates and represent mean ± s.d. of triplicates of one representative donor. ***P < 0.0001 by unpaired two-sided t-test. Values in a–c indicate MFI relative to WT cells stained with the same reagent. Data are representative of at least two independent experiments (a–g).
Extended Data Fig. 8 QPCTL deficiency leads to enhanced tumor cell control by tumor specific antibodies.
a, Schematic representation of in vivo set-up. b, Absolute number (see Fig. 4c) of recovered tumor cells from mice injected with 1:1 mixtures of WT and QPCTL KO Ba/F3-Her2 cells that were then treated with control (PBS) (-) or anti-Her2 (IgA1) ( + ). n = 6 control-treated animals; n = 6 anti-Her2-treated animals. ***P≤0.0003 by unpaired two-sided t-test. c, Ratio of in vivo killing of target cells in mice injected with a 1:1 mixture of WT and CD47-KO cells, or a 1:1 mixture of WT and QPCTL KO Ba/F3-Her2 cells, and that were either treated with control (PBS) (-) or anti-Her2 (IgA1) antibody ( + ).n = 6 control-treated animals (left graph); n = 5 anti-Her2-treated animals (left graph); n = 6 control-treated animals (right graph); n = 6 anti-Her2-treated animals (right graph). ***P < 0.0001 unpaired two-sided t-test. d, Absolute number (see Extended Data Fig. 8c) of recovered tumor cells in mice injected with a 1:1 mixture of WT and CD47 KO cells, or a 1:1 mixture of WT and QPCTL KO Ba/F3-Her2 cells, and that were either treated with control (PBS) (-) or anti-Her2 (IgA1) antibody ( + ). n = 6 control-treated animals (left graph); n = 5 anti-Her2-treated animals (left graph); n = 6 control-treated animals (right graph); n = 6 anti-Her2-treated animals (right graph). ***P < 0.0001 by one-way ANOVA with multiple comparison correction; n.s., not significant. Dots represent mice treated with control (PBS), squares represent mice treated with anti-Her2 (IgA1) (b-d) and represent mean ± s.d. of individual mice (b–d). Data are representative of two independent experiments (b) or one experiment (c,d).
Extended Data Fig. 9 QPCTL deficiency in combination with tumor specific antibodies leads to an enhanced neutrophil influx.
a, Absolute number (see Fig. 4c,d,f) of CD8+ T (CD3+ CD8+), CD4+ T (CD3+ CD4+) or B (B220+ MHCII+) cells present in mice that received a 1:1 mixture of WT and QPCTL KO Ba/F3-Her2 cells, and that were either control (PBS)-treated (-) or treated with anti-Her2 (IgA1) ( + ). n = 6 control-treated animals; n = 6 anti-Her2-treated animals. **P = 0.0050. by unpaired two-sided t-test; n.s., not significant. b, Absolute number of peritoneal neutrophils (Ly-6G+/CD11b+), macrophages (F4/80+ CD11b+), CD8+ T (CD3+/CD8+), CD4+ T (CD3+/CD4+) and B (B220+/MHCII+) cells present in recipients of a 1:1 mixture of WT and QPCTL KO Ba/F3-Her2 cells that were control (PBS)-treated (-) or treated with anti-Her2 (IgA1) ( + )(see Extended Data Fig. 8c). n = 6 control-treated animals injected with mixture of WT/CD47 KO cells; n = 6 anti-Her2-treated animals injected with mixture of WT/CD47 KO cells; n = 6 control-treated animals injected with mixture of WT/QPCTL KO cells; n = 5 anti-Her2-treated animals injected with mixture of WT/QPCTL KO cells. ***P < 0.0001; 0.0015≤**P≤0.0022; *P = 0.0226 by one-way ANOVA with multiple comparison correction; n.s., not significant. c, Absolute number of recovered tumor cells in recipients of WT (blue) or QPCTL KO (green) Ba/F3-Her2 cells that were treated with anti-Her2 (IgA1) antibody. n = 5 animals injected with WT cells. n = 5 animals injected with QPCTL KO cells. *P = 0.0161 by unpaired two-sided t-test; n.s., not significant. d, Absolute number of peritoneal neutrophils (Ly-6G+/CD11b+), macrophages (F4/80+ CD11b+) and CD3+ T cells, CD4 effector (CD4+/FOXP3-) and CD4 regulatory (CD4+/FOXP3+) T cells present in recipients of WT (blue) or QPCTL KO (green) Ba/F3-Her2 cells that were treated with anti-Her2 (IgA1) (see Extended Data Fig. 7g). n = 5 animals injected with WT cells. n = 5 animals injected with QPCTL KO cells. n.s., not significant. Dots represent mice treated with control (PBS), squares represent mice treated with anti-Her2 (IgA1) (a–d) and represent mean ± s.d. of individual mice (a–d). Data are representative of two independent experiments (a) or one experiment (b–d).
Extended Data Fig. 10 Expansion, differentiation, cytokine production and killing capacity of human T cells are unaltered by glutaminyl cyclase inhibition.
To determine the effect of glutaminyl cyclase inhibition on different aspects of T cell function, in vitro T cell cytokine production and secretion, T cell phenotype, T cell killing capacity, and T cell induction by autologous dendritic cells (DCs) was assessed in the presence of control (DMSO) or SEN177. a and b, CDK4 T cell receptor-transduced T cells (CDK4−specific T cells) from two donors were cultured for 4 days in the presence of control (DMSO) or SEN177 a, Intracellular cytokine production of control (DMSO)-treated or SEN177-treated CDK4-specific T cells upon co-culture with JY cells pulsed with the indicated concentrations of CDK4 or MART-1 peptide, as analyzed by flow cytometry. Values indicate percentage of IL-2+ (left), IFNγ+ (middle) and TNFα+ (right) CD8+ cells of total CD8+ cells. Data represent n = 3 biological replicates and mean ± s.d. of triplicates. b, Specific lysis of CDK4+ (NKIRTIL006) and CDK4- (A375) tumor cells by control (DMSO)-treated or SEN177-treated CDK4-specific T cells, as determined in a 4 h 51Cr-release assay. Data represent n = 3 (Donor 1), n = 2 (Donor 2 – NKIRTIL006 cocultures) or n = 1 (donor 2 - A375 cocultures) biological replicates and mean ± s.d of triplicates (Donor 1). c–e, Peripheral blood lymphocytes (PBLs) from two donors were isolated, stimulated with anti-CD3/CD28 beads (2:1 bead to T cell ratio), and cultured for two weeks in the presence of control (DMSO) or SEN177. Data represent n = 3 biological replicates and mean ± s.d. of triplicates. c, Fold expansion of control (DMSO)-treated or SEN177-treated CDK4-specific T cells over a period of 11 days. d, Surface expression of CCR7, CD4, CD8, CD27, CD28, CD45RA, CD45RO, CD62L and PD-1 on CD3+ T cells after 14 days of control (DMSO) treatment or SEN177 treatment, as determined by flow cytometry. Values indicate percentage of CD3+ cells positive for the indicated marker. e, IFNγ, IL-2, IL-4, TNFα, IL-6, IL-17A and IL-10 secretion of T cells after 14 days of control (DMSO) treatment or SEN177 treatment and subsequent re-stimulation with anti-CD3/CD28 beads, as determined by CBA bead array. Values indicate MFI. f, Differentiation of control (DMSO)-treated (-) or SEN177-treated ( + ) CD4+ T cells into Th17 cells after 7 days (t = 7d) or 10 days (t = 10d) of culture, as determined by flow cytometry. Values indicate the percentage of IL17A+ cells of total lymphocytes. Data represent n = 3 biological replicates and mean ± s.d. of triplicates. n.s., not significant by unpaired two-sided t-test. h and i, T cells were induced by autologous dendritic cells (DCs) under three conditions; control (DMSO) was present during DC maturation and during T cell induction (‘control’), control (DMSO) was present during DC maturation and SEN177 was present during T cell induction (‘SEN177 induction)’, or SEN177 was present during both DC maturation and T cell induction (‘SEN177 maturation and induction’). h, Surface expression of CCR7, CD4, CD8, CD27, CD28, CD45RA, CD62L and PD-1 on CD3+ T cells 12 days after start of induction by autologous DCs. Values reflect percentage of CD3+ cells positive for the indicated marker. g, Percentage of MHC multimer positive CD8+ T cells before induction (t = 0) and 11 days after start of induction by autologous DCs, as determined by flow cytometry. Values indicate the percentage of MHC multimer (HLA-A*02:01 multimers loaded with the MART-1-ELA, CMV-NLV, EBV-GLC or EBV-YVL epitopes) CD8+ cells of total CD8+ cells. i, Intracellular cytokine production of CD8+ T cells 6 days after start of induction and subsequent re-stimulation with unloaded DCs (–) or peptide-loaded DCs (+). Values indicate percentage of IL-2+ (left), IFNγ+ (middle) and TNFα+ (right) CD8+ cells of total CD8+ cells. Data are representative of one (a–i) independent experiment.
Supplementary information
Supplementary Information
Supplementary Table 1
Source data
Source Data Fig. 2
Unprocessed Western Blots
Source Data Fig. 3
Unprocessed gel and Western Blot
Source Data Extended Data Fig. 2
Unprocessed Western Blot
Rights and permissions
About this article
Cite this article
Logtenberg, M.E.W., Jansen, J.H.M., Raaben, M. et al. Glutaminyl cyclase is an enzymatic modifier of the CD47- SIRPα axis and a target for cancer immunotherapy. Nat Med 25, 612–619 (2019). https://doi.org/10.1038/s41591-019-0356-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-019-0356-z
This article is cited by
-
Gentulizumab, a novel anti-CD47 antibody with potent antitumor activity and demonstrates a favorable safety profile
Journal of Translational Medicine (2024)
-
Unsynchronized butyrophilin molecules dictate cancer cell evasion of Vγ9Vδ2 T-cell killing
Cellular & Molecular Immunology (2024)
-
Pan-cancer analysis for the prognostic and immunological role of CD47: interact with TNFRSF9 inducing CD8 + T cell exhaustion
Discover Oncology (2024)
-
Emerging phagocytosis checkpoints in cancer immunotherapy
Signal Transduction and Targeted Therapy (2023)
-
Development of a potent benzonitrile-based inhibitor of glutaminyl-peptide cyclotransferase-like protein (QPCTL) with antitumor efficacy
Signal Transduction and Targeted Therapy (2023)