Abstract
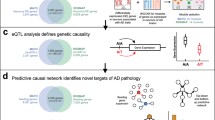
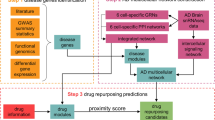
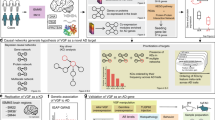
Identifying the mechanisms through which genetic risk causes dementia is an imperative for new therapeutic development. Here, we apply a multistage, systems biology approach to elucidate the disease mechanisms in frontotemporal dementia. We identify two gene coexpression modules that are preserved in mice harboring mutations in MAPT, GRN and other dementia mutations on diverse genetic backgrounds. We bridge the species divide via integration with proteomic and transcriptomic data from the human brain to identify evolutionarily conserved, disease-relevant networks. We find that overexpression of miR-203, a hub of a putative regulatory microRNA (miRNA) module, recapitulates mRNA coexpression patterns associated with disease state and induces neuronal cell death, establishing this miRNA as a regulator of neurodegeneration. Using a database of drug-mediated gene expression changes, we identify small molecules that can normalize the disease-associated modules and validate this experimentally. Our results highlight the utility of an integrative, cross-species network approach to drug discovery.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
miRNA-seq and mRNA-seq data from TPR50 tau mice, microarray data on PS19 hippocampus, microarray data on overexpression of miR-203 in vitro, RNA-seq on sorted mouse neurons and RNA-seq data with SAHA are available from the NCBI Gene Expression Omnibus database under Gene Expression Omnibus accession number GSE90696. Human FTD miRNA-seq and mRNA-seq data are available from https://www.synapse.org/#!Synapse:syn7818788. Human UPenn FTD Proteomics data are available from https://www.synapse.org/#!Synapse:syn9884357. The custom code used for the analysis can be accessed using this link in github: https://github.com/dhglab/Identification-of-evolutionarily-conserved-gene-networks-mediating-neurodegenerative-dementia
References
Hinz, F. I. & Geschwind, D. H. Molecular genetics of neurodegenerative dementias. Cold Spring Harb. Perspect. Biol. 9, a023705 (2017).
Iqbal, K., Liu, F. & Gong, C. X. Tau and neurodegenerative disease: the story so far. Nat. Rev. Neurol. 12, 15–27 (2016).
Masters, C. L. et al. Alzheimer’s disease. Nat. Rev. Dis. Primers 1, 15056 (2015).
Kovacs, G. G. Invited review: neuropathology of tauopathies: principles and practice. Neuropathol. Appl. Neurobiol. 41, 3–23 (2015).
Mullane, K. & Williams, M. Alzheimer’s therapeutics: continued clinical failures question the validity of the amyloid hypothesis—but what lies beyond? Biochem. Pharmacol. 85, 289–305 (2013).
Institute of Medicine. Improving the Utility and Translation of Animal Models for Nervous System Disorders: Workshop Summary (The National Academies Press, Washington DC, 2013).
Miller, J. A., Horvath, S. & Geschwind, D. H. Divergence of human and mouse brain transcriptome highlights Alzheimer disease pathways. Proc. Natl. Acad. Sci. USA 107, 12698–12703 (2010).
Qosa, H. & Kaddoumi, A. Effect of mouse strain as a background for Alzheimer’s disease models on the clearance of amyloid-β. J. Syst. Integr. Neurosci. 2, 135–140 (2016).
Weitzner, D. S., Engler-Chiurazzi, E. B., Kotilinek, L. A., Ashe, K. H. & Reed, M. N. Morris Water Maze Test: optimization for mouse strain and testing environment. J. Vis. Exp. e52706 (2015).
LaFerla, F. M. & Green, K. N. Animal models of Alzheimer disease. Cold Spring Harb. Perspect. Med. 2, a0066320 (2012).
Webster, S. J., Bachstetter, A. D., Nelson, P. T., Schmitt, F. A. & Van Eldik, L. J. Using mice to model Alzheimer’s dementia: an overview of the clinical disease and the preclinical behavioral changes in 10 mouse models. Front. Genet. 5, 88 (2014).
Karsten, S. L. et al. A genomic screen for modifiers of tauopathy identifies puromycin-sensitive aminopeptidase as an inhibitor of tau-induced neurodegeneration. Neuron 51, 549–560 (2006).
Onishi, T. et al. Early-onset cognitive deficits and axonal transport dysfunction in P301S mutant tau transgenic mice. Neurosci. Res. 80, 76–85 (2014).
Yoshiyama, Y. et al. Synapse loss and microglial activation precede tangles in a P301S tauopathy mouse model. Neuron 53, 337–351 (2007).
Zhang, B. & Horvath, S. A general framework for weighted gene co-expression network analysis. Stat. Appl. Genet. Mol. Biol. 4, Article17 (2005).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9, 559 (2008).
Spillantini, M. G. & Goedert, M. Tau protein pathology in neurodegenerative diseases. Trends Neurosci. 21, 428–433 (1998).
Braak, H. & Braak, E. Neuropathological stageing of Alzheimer-related changes. Acta Neuropathol. 82, 239–259 (1991).
Kuhn, A., Thu, D., Waldvogel, H. J., Faull, R. L. M. & Luthi-Carter, R. Population-specific expression analysis (PSEA) reveals molecular changes in diseased brain. Nat. Methods 8, 945–947 (2011).
Miller, J. A., Woltjer, R. L., Goodenbour, J. M., Horvath, S. & Geschwind, D. H. Genes and pathways underlying regional and cell type changes in Alzheimer’s disease. Genome Med. 5, 48 (2013).
Lage, K. et al. A human phenome–interactome network of protein complexes implicated in geneticdisorders.Nat. Biotechnol. 25, 309–316 (2007).
Stark, C. et al. BioGRID: a general repository for interaction datasets. Nucleic Acids Res. 34, D535–D539 (2006).
Parikshak, N. N. et al. Integrative functional genomic analyses implicate specific molecular pathways and circuits in autism. Cell 155, 1008–1021 (2013).
Parikshak, N. N., Gandal, M. J. & Geschwind, D. H. Systems biology and gene networks in neurodevelopmental and neurodegenerative disorders. Nat. Rev. Genet. 16, 441–458 (2015).
Murray, R. Z., Wylie, F. G., Khromykh, T., Hume, D. A. & Stow, J. L. Syntaxin 6 and Vti1b form a novel SNARE complex, which is up-regulated in activated macrophages to facilitate exocytosis of tumor necrosis Factor-alpha. J. Biol. Chem. 280, 10478–10483 (2005).
Gjørlund, M. D. et al. Neuroligin-1 induces neurite outgrowth through interaction with neurexin-1β and activation of fibroblast growth factor receptor-1. FASEB J. 26, 4174–4186 (2012).
Huang, K. P. et al. Neurogranin/RC3 enhances long-term potentiation and learning by promoting calcium-mediated signaling. J. Neurosci. 24, 10660–10669 (2004).
Jaworski, M. et al. Malt1 protease inactivation efficiently dampens immune responses but causes spontaneous autoimmunity. EMBO J. 33, 2765–2781 (2014).
Lessard, C. J. et al. Variants at multiple loci implicated in both innate and adaptive immune responses are associated with Sjögren’s syndrome. Nat. Genet. 45, 1284–1292 (2013).
Ng, A. S. L., Rademakers, R. & Miller, B. L. Frontotemporal dementia: a bridge between dementia and neuromuscular disease. Ann. N. Y. Acad. Sci. 1338, 71–93 (2015).
Maeda, S. et al. Expression of A152T human tau causes age-dependent neuronal dysfunction and loss in transgenic mice. EMBO Rep. 17, 530–551 (2016).
Parikshak, N. N. et al. Genome-wide changes in lncRNA, splicing, and regional gene expression patterns in autism. Nature 540, 423–427 (2016).
Gandal, M. J. et al. Shared molecular neuropathology across major psychiatric disorders parallels polygenic overlap. Science 359, 693–697 (2018).
Xie, J. et al. Long-term, efficient inhibition of microRNA function in mice using rAAV vectors. Nat. Methods 9, 403–409 (2012).
Menkes-Caspi, N. et al. Pathological tau disrupts ongoing network activity. Neuron 85, 959–966 (2015).
Zhang, B. et al. Integrated systems approach identifies genetic nodes and networks in late-onset Alzheimer’s disease. Cell 153, 707–720 (2013).
Jones, L. et al. Convergent genetic and expression data implicate immunity in Alzheimer’s disease. Alzheimers Dement. 11, 658–671 (2015).
Narayanan, M. et al. Common dysregulation network in the human prefrontal cortex underlies two neurodegenerative diseases. Mol. Syst. Biol. 10, 743 (2014).
Dolmetsch, R. & Geschwind, D. H. The human brain in a dish: the promise of iPSC-derived neurons. Cell 145, 831–834 (2011).
Hardy, J. Catastrophic cliffs: a partial suggestion for selective vulnerability in neurodegenerative diseases. Biochem. Soc. Trans. 44, 659–661 (2016).
Gupta, S., Verma, S., Mantri, S., Berman, N. E. & Sandhir, R. Targeting microRNAs in prevention and treatment of neurodegenerative disorders. Drug Dev. Res. 76, 397–418 (2015).
Janssen, H. L. A. et al. Treatment of HCV infection by targeting microRNA. N. Engl. J. Med. 368, 1685–1694 (2013).
Young, D. D., Connelly, C. M., Grohmann, C. & Deiters, A. Small molecule modifiers of microRNA miR-122 function for the treatment of hepatitis C virus infection and hepatocellular carcinoma. J. Am. Chem. Soc. 132, 7976–7981 (2010).
Allen, M. et al. Human whole genome genotype and transcriptome data for Alzheimer’s and other neurodegenerative diseases. Sci. Data 3, 160089 (2016).
Seyfried, N. T. et al. A multi-network approach identifies protein-specific co-expression in asymptomatic and symptomatic Alzheimer’s disease. Cell Syst. 4, 60–72.e4 (2017).
Prudencio, M. et al. Distinct brain transcriptome profiles in C9orf72-associated and sporadic ALS. Nat. Neurosci. 18, 1175–1182 (2015).
Chang, L.-C. et al. A conserved BDNF, glutamate- and GABA-enriched gene module related to human depression identified by coexpression meta-analysis and DNA variant genome-wide association studies. PLoS ONE 9, e90980 (2014).
Fromer, M. et al. Gene expression elucidates functional impact of polygenic risk for schizophrenia. Nat. Neurosci. 19, 1442–1453 (2016).
Ferrari, R. et al. Frontotemporal dementia and its subtypes: a genome-wide association study. Lancet Neurol. 13, 686–699 (2014).
Höglinger, G. U. et al. Identification of common variants influencing risk of the tauopathy progressive supranuclear palsy. Nat. Genet. 43, 699–705 (2011).
Lambert, J.-C. et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer's disease. Nat. Genet. 45, 1452–1458 (2013).
Schroeder, A. et al. The RIN: an RNA integrity number for assigning integrity values to RNA measurements. BMC Mol. Biol. 7, 3 (2006).
Wes, P. D. et al. Tau overexpression impacts a neuroinflammation gene expression network perturbed in Alzheimer’s disease. PLoS ONE 9, e106050 (2014).
Srinivasan, K. et al. Untangling the brain’s neuroinflammatory and neurodegenerative transcriptional responses. Nat. Commun. 7, 11295 (2016).
Chishti, M. A. et al. Early-onset amyloid deposition and cognitive deficits in transgenic mice expressing a double mutant form of amyloid precursor protein 695. J. Biol. Chem. 276, 21562–21570 (2001).
Matarin, M. et al. A genome-wide gene-expression analysis and database in transgenic mice during development of amyloid or tau pathology. Cell Rep. 10, 633–644 (2015).
Lui, H. et al. Progranulin deficiency promotes circuit-specific synaptic pruning by microglia via complement activation. Cell 165, 921–935 (2016).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Anders, S., Pyl, P. T. & Huber, W. HTSeq: a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Conesa, A. et al. A survey of best practices for RNA-seq data analysis. Genome Biol. 17, 13 (2016).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Risso, D., Schwartz, K., Sherlock, G. & Dudoit, S. GC-content normalization for RNA-seq data.BMC Bioinformatics 12, 480 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Hansen, K. D., Irizarry, R. A. & Wu, Z. Removing technical variability in RNA-seq data using conditional quantile normalization. Biostatistics 13, 204–216 (2012).
Friedländer, M. R., Mackowiak, S. D., Li, N., Chen, W. & Rajewsky, N. miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Res. 40, 37–52 (2012).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Li, A. & Horvath, S. Network neighborhood analysis with the multi-node topological overlap measure. Bioinformatics 23, 222–231 (2007).
Hawrylycz, M. et al. Canonical genetic signatures of the adult human brain. Nat. Neurosci. 18, 1832–1844 (2015).
Zambon, A. C. et al. GO-Elite: a flexible solution for pathway and ontology over-representation. Bioinformatics 28, 2209–2210 (2012).
Csardi, G. & Nepusz, T. The igraph software package for complex network research. nterJournal,Complex Systems 1695, 1–9 (2006).
Zhang, Y. et al. An RNA-sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J. Neurosci. 34, 11929–11947 (2014).
Langfelder, P., Luo, R., Oldham, M. C. & Horvath, S. Is my network module preserved and reproducible? PLoS Comput. Biol. 7, e1001057 (2011).
Rivals, I., Personnaz, L., Taing, L. & Potier, M. C. Enrichment or depletion of a GO category within a class of genes: which test? Bioinformatics 23, 401–407 (2007).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Series B Methodol. 57, 289–300 (1995).
Grimson, A. et al. MicroRNA targeting specificity in mammals: determinants beyond seed pairing. Mol. Cell 27, 91–105 (2007).
Rossin, E. J. et al. Proteins encoded in genomic regions associated with immune-mediated disease physically interact and suggest underlying biology. PLoS Genet. 7, e1001273 (2011).
Voineagu, I. et al. Transcriptomic analysis of autistic brain reveals convergent molecular pathology. Nature 474, 380–384 (2011).
Cox, J. et al. Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ. Mol. Cell Proteomics 13, 2513–2526 (2014).
Lamb, J. et al. The Connectivity Map: using gene-expression signatures to connect small molecules, genes, and disease. Science 313, 1929–1935 (2006).
Cheng, C., Fass, D. M., Folz-Donahue, K., Macdonald, M. E. & Haggarty, S. J. Highly expandable human iPS cell–derived neural progenitor cells (NPC) and neurons for central nervous system disease modeling andhigh-throughput screening. Curr. Protoc. Hum. Genet. 92, 21.8.1–21.8.21 (2017).
Almeida, S. et al. Induced pluripotent stem cell models of progranulin-deficient frontotemporal dementia uncover specific reversible neuronal defects. Cell Rep. 2, 789–798 (2012).
Biswas, M. H. U. et al. MMP-9 and MMP-2 contribute to neuronal cell death in iPSC models of frontotemporal dementia with MAPT mutations. Stem Cell Reports 7, 316–324 (2016).
Acknowledgements
Funding for this work was provided by Takeda Pharmaceuticals (D.H.G.), Rainwater Charitable Foundation/Tau consortium (D.H.G., S.J.H.), NIH grants to D.H.G., S.J.H., A.L. (5U01AG046161) and J.R. (5R25 NS065723) and Larry L. Hillblom Foundation Postdoctoral Fellowship to V.S. K.N., H.T., A.O., K.H. and S.K. are employees of Takeda Pharmaceuticals. The authors thank M. Hattori, Y. Obayashi and K. Nakamura for their contribution to the TPR50 mouse sample preparation and analysis. The authors thank M. Gearing at the Emory Alzheimer’s Disease Research Center brain bank for providing human FTD samples. The authors also thank N. Parikshak for help with network analysis and critical reading of the manuscript. We thank Eli Lilly and Company scientists for generating the Tg4510 microglia RNA-seq data and providing access to them. For the PSP and FTD temporal cortex RNA-seq dataset, study data were provided by the following sources: Mayo Clinic Alzheimer’s Disease Genetic Studies, led by N. Taner and S. G. Younkin, Mayo Clinic Jacksonville, using samples from the Mayo Clinic Study of Aging, Mayo Clinic Alzheimer’s Disease Research Center, and Mayo Clinic Brain Bank. Data collection was supported through funding by National Institute on Aging (NIA) grants P50 AG016574, R01 AG032990, U01 AG046139, R01 AG018023, U01 AG006576, U01 AG006786, R01 AG025711, R01 AG017216, R01 AG003949, National Institute of Neurological Disorders and Stroke (NINDS) grant R01 NS080820, the CurePSP Foundation, and support from the Mayo Foundation. Study data include samples collected through the Sun Health Research Institute Brain and Body Donation Program of Sun City, Arizona. The Brain and Body Donation Program is supported by the NINDS (U24 NS072026 National Brain and Tissue Resource for Parkinson’s Disease and Related Disorders), the NIA (P30 AG19610 Arizona Alzheimer’s Disease Core Center), the Arizona Department of Health Services (contract 211002, Arizona Alzheimer’s Research Center), the Arizona Biomedical Research Commission (contracts 4001, 0011, 05-901 and 1001 to the Arizona Parkinson’s Disease Consortium) and the Michael J. Fox Foundation for Parkinson’s Research. Tg4510 replication and CRND8 RNA-seq data were provided by the NIH U01 AG046139. We thank J. Lewis, K. Duff, D. Westaway and D. Borchelt for generating these lines of transgenic mice and providing access to them.
Author information
Authors and Affiliations
Consortia
Contributions
V.S. and D.H.G. planned and directed the experiments, guided the analysis and wrote the manuscript in conjunction with F.I.H. and J.E.R. All authors revised and edited the final version of the manuscript. F.I.H. performed all the experiments in mouse cortical cultures. V.S. performed all the bioinformatic analyses and performed dissections on human postmortem samples and isolated RNA. K.N., H.T., A.O., K.H. and S.K. bred the TPR50 mouse, characterized the F1 hybrids and collected the tissue samples. J.E.R. performed bioinformatic analysis using purified glial cells. J.E.R. and A.S. stained for and quantified inflammation in mouse brain samples. IFGC consortia members collected and analyzed FTD GWAS data. N.T.S., J.J.L. and A.I.L. performed mass spectrometry–based quantitative proteomics on human FTD samples obtained from M.G., V.M.V.D. and J.Q.T. The SAHA experiments were performed by C.C. and S.J.H. on human iPSC-derived neurons from tau and control patients.
Corresponding author
Ethics declarations
Competing interests
D.H.G. has received research funding from Takeda Pharmaceutical Company Limited. K.N., H.T., A.O., K.H. and S.K. are employees of Takeda Pharmaceutical Company Limited.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10
Supplementary Table 1
TPR50 mRNA-seq Analysis
Supplementary Table 2
Human transcriptomics and proteomics data
Supplementary Table 3
TPR50 miRNA-seq analysis
Supplementary Table 4
Module preservation and connectivity map
Rights and permissions
About this article
Cite this article
Swarup, V., Hinz, F.I., Rexach, J.E. et al. Identification of evolutionarily conserved gene networks mediating neurodegenerative dementia. Nat Med 25, 152–164 (2019). https://doi.org/10.1038/s41591-018-0223-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-018-0223-3
This article is cited by
-
Whole-genome sequencing analysis reveals new susceptibility loci and structural variants associated with progressive supranuclear palsy
Molecular Neurodegeneration (2024)
-
Investigation of early molecular alterations in tauopathy with generative adversarial networks
Scientific Reports (2023)
-
Frontotemporal lobar degeneration
Nature Reviews Disease Primers (2023)
-
Tau activation of microglial cGAS–IFN reduces MEF2C-mediated cognitive resilience
Nature Neuroscience (2023)
-
miR-203, fine-tunning neuroinflammation by juggling different components of NF‐κB signaling
Journal of Neuroinflammation (2022)