Abstract
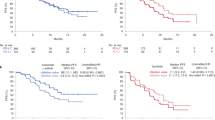
We describe results from IMmotion150, a randomized phase 2 study of atezolizumab (anti-PD-L1) alone or combined with bevacizumab (anti-VEGF) versus sunitinib in 305 patients with treatment-naive metastatic renal cell carcinoma. Co-primary endpoints were progression-free survival (PFS) in intent-to-treat and PD-L1+ populations. Intent-to-treat PFS hazard ratios for atezolizumab + bevacizumab or atezolizumab monotherapy versus sunitinib were 1.0 (95% confidence interval (CI), 0.69–1.45) and 1.19 (95% CI, 0.82–1.71), respectively; PD-L1+ PFS hazard ratios were 0.64 (95% CI, 0.38–1.08) and 1.03 (95% CI, 0.63–1.67), respectively. Exploratory biomarker analyses indicated that tumor mutation and neoantigen burden were not associated with PFS. Angiogenesis, T-effector/IFN-γ response, and myeloid inflammatory gene expression signatures were strongly and differentially associated with PFS within and across the treatments. These molecular profiles suggest that prediction of outcomes with anti-VEGF and immunotherapy may be possible and offer mechanistic insights into how blocking VEGF may overcome resistance to immune checkpoint blockade.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Change history
05 October 2018
In the version of this article originally published, there was an error in Fig. 2n. The top line of the HR comparison chart originally was Atezo + bev vs sun. It should have been Atezo + bev vs atezo. The error has been corrected in the HTML and PDF versions of this article.
References
Kaelin, W. G. Jr. The von Hippel-Lindau gene, kidney cancer, and oxygen sensing. J. Am. Soc. Nephrol. 14, 2703–2711 (2003).
George, D. J. & Kaelin, W. G. Jr. The von Hippel-Lindau protein, vascular endothelial growth factor, and kidney cancer. N. Engl. J. Med. 349, 419–421 (2003).
Motzer, R. J. et al. Tivozanib versus sorafenib as initial targeted therapy for patients with metastatic renal cell carcinoma: results from a phase III trial. J. Clin. Oncol. 31, 3791–3799 (2013).
Clark, J. I. et al. Impact of sequencing targeted therapies with high-dose interleukin-2 immunotherapy: an analysis of outcome and survival of patients with metastatic renal cell carcinoma from an on-going observational Il-2 clinical trial: PROCLAIMSM. Clin. Genitourin. Cancer 15, 31–41.e4 (2017).
Herbst, R. S. et al. Predictive correlates of response to the anti-PD-L1 antibody MPDL3280A in cancer patients. Nature 515, 563–567 (2014).
Choueiri, T. K. et al. PD-L1 expression in nonclear-cell renal cell carcinoma. Ann. Oncol. 25, 2178–2184 (2014).
Thompson, R. H. et al. PD-1 is expressed by tumor-infiltrating immune cells and is associated with poor outcome for patients with renal cell carcinoma. Clin. Cancer Res. 13, 1757–1761 (2007).
Motzer, R. J. et al. Nivolumab versus everolimus in advanced renal-cell carcinoma. N. Engl. J. Med. 373, 1803–1813 (2015).
Thompson, R. H. et al. Costimulatory B7-H1 in renal cell carcinoma patients: indicator of tumor aggressiveness and potential therapeutic target. Proc. Natl. Acad. Sci. USA 101, 17174–17179 (2004).
Thompson, R. H. et al. Tumor B7-H1 is associated with poor prognosis in renal cell carcinoma patients with long-term follow-up. Cancer Res. 66, 3381–3385 (2006).
Keir, M. E., Butte, M. J., Freeman, G. J. & Sharpe, A. H. PD-1 and its ligands in tolerance and immunity. Annu. Rev. Immunol. 26, 677–704 (2008).
Latchman, Y. E. et al. PD-L1-deficient mice show that PD-L1 on T cells, antigen-presenting cells, and host tissues negatively regulates T cells. Proc. Natl. Acad. Sci. USA 101, 10691–10696 (2004).
Yang, J. et al. The novel costimulatory programmed death ligand 1/B7.1 pathway is functional in inhibiting alloimmune responses in vivo. J. Immunol. 187, 1113–1119 (2011).
McDermott, D. F. et al. Atezolizumab, an anti-programmed death-ligand 1 antibody, in metastatic renal cell carcinoma: long-term safety, clinical activity, and immune correlates from a phase Ia study. J. Clin. Oncol. 34, 833–842 (2016).
Elamin, Y. Y., Rafee, S., Toomey, S. & Hennessy, B. T. Immune effects of bevacizumab: killing two birds with one stone. Cancer Microenviron. 8, 15–21 (2015).
Kusmartsev, S. et al. Oxidative stress regulates expression of VEGFR1 in myeloid cells: link to tumor-induced immune suppression in renal cell carcinoma. J. Immunol. 181, 346–353 (2008).
Roland, C. L. et al. Inhibition of vascular endothelial growth factor reduces angiogenesis and modulates immune cell infiltration of orthotopic breast cancer xenografts. Mol. Cancer Ther. 8, 1761–1771 (2009).
Gabrilovich, D. I. & Nagaraj, S. Myeloid-derived suppressor cells as regulators of the immune system. Nat. Rev. Immunol. 9, 162–174 (2009).
Hodi, F. S. et al. Bevacizumab plus ipilimumab in patients with metastatic melanoma. Cancer Immunol. Res. 2, 632–642 (2014).
Roland, C. L. et al. Cytokine levels correlate with immune cell infiltration after anti-VEGF therapy in preclinical mouse models of breast cancer. PLoS One 4, e7669 (2009).
Wallin, J. J. et al. Atezolizumab in combination with bevacizumab enhances antigen-specific T-cell migration in metastatic renal cell carcinoma. Nat. Commun. 7, 12624 (2016).
Osada, T. et al. The effect of anti-VEGF therapy on immature myeloid cell and dendritic cells in cancer patients. Cancer Immunol. Immunother. 57, 1115–1124 (2008).
Brauer, M. J. et al. Identification and analysis of in vivo VEGF downstream markers link VEGF pathway activity with efficacy of anti-VEGF therapies. Clin. Cancer Res. 19, 3681–3692 (2013).
Fehrenbacher, L. et al. Atezolizumab versus docetaxel for patients with previously treated non-small-cell lung cancer (POPLAR): a multicentre, open-label, phase 2 randomised controlled trial. Lancet 387, 1837–1846 (2016).
Rosenberg, J. E. et al. Atezolizumab in patients with locally advanced and metastatic urothelial carcinoma who have progressed following treatment with platinum-based chemotherapy: a single-arm, multicentre, phase 2 trial. Lancet 387, 1909–1920 (2016).
Snyder, A. et al. Genetic basis for clinical response to CTLA-4 blockade in melanoma. N. Engl. J. Med. 371, 2189–2199 (2014).
Rizvi, N. A. et al. Activity and safety of nivolumab, an anti-PD-1 immune checkpoint inhibitor, for patients with advanced, refractory squamous non-small-cell lung cancer (CheckMate 063): a phase 2, single-arm trial. Lancet Oncol. 16, 257–265 (2015).
Balar, A. V. et al. Atezolizumab as first-line treatment in cisplatin-ineligible patients with locally advanced and metastatic urothelial carcinoma: a single-arm, multicentre, phase 2 trial. Lancet 389, 67–76 (2017).
Scheller, J., Chalaris, A., Schmidt-Arras, D. & Rose-John, S. The pro- and anti-inflammatory properties of the cytokine interleukin-6. Biochim. Biophys. Acta 1813, 878–888 (2011).
Russo, R. C., Garcia, C. C., Teixeira, M. M. & Amaral, F. A. The CXCL8/IL-8 chemokine family and its receptors in inflammatory diseases. Expert Rev. Clin. Immunol. 10, 593–619 (2014).
Ha, H., Debnath, B. & Neamati, N. Role of the CXCL8-CXCR1/2 axis in cancer and inflammatory diseases. Theranostics 7, 1543–1588 (2017).
Zelenay, S. et al. Cyclooxygenase-dependent tumor growth through evasion of immunity. Cell 162, 1257–1270 (2015).
Powles, T. et al. Immune biomarkers associated with clinical benefit from atezolizumab (MPDL3280a; anti-PD-L1) in advanced urothelial bladder cancer (UBC). J. Immunother. Cancer 3, 83 (2015).
Rizvi, N. A. et al. Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348, 124–128 (2015).
Carbone, D. P. et al. First-line nivolumab in stage IV or recurrent non-small-cell lung cancer. N. Engl. J. Med. 376, 2415–2426 (2017).
Turajlic, S. et al. Insertion-and-deletion-derived tumour-specific neoantigens and the immunogenic phenotype: a pan-cancer analysis. Lancet Oncol. 18, 1009–1021 (2017).
Motzer, R. J. et al. IMmotion151: a randomized phase III study of atezolizumab plus bevacizumab vs. sunitinib in untreated metastatic renal cell carcinoma (mRCC). J. Clin. Oncol. 36, abstr, 578 (2018).
Choueiri, T. K. et al. First-line avelumab + axitinib therapy in patients (pts) with advanced renal cell carcinoma (aRCC): Results from a phase Ib trial. J. Clin. Oncol. 35, 4504 (2017).
Motzer, R. Nivolumab plus ipilimumab versus aunitinib in advanced renal-cell carcinoma. N. Engl. J. Med. 378, 1277–1290 (2018).
Voss, M. H. et al. Integrated biomarker analysis for 412 renal cell cancer (RCC) patients (pts) treated on the phase 3 COMPARZ trial: correlating common mutation events in PBRM1 and BAP1 with angiogenesis expression signatures and outcomes on tyrosine kinase inhibitor (TKI) therapy. J. Clin. Oncol. 35, 4523 (2017).
Prima, V., Kaliberova, L. N., Kaliberov, S., Curiel, D. T. & Kusmartsev, S. COX2/mPGES1/PGE2 pathway regulates PD-L1 expression in tumor-associated macrophages and myeloid-derived suppressor cells. Proc. Natl. Acad. Sci. USA 114, 1117–1122 (2017).
Coussens, L. M. & Werb, Z. Inflammation and cancer. Nature 420, 860–867 (2002).
Liu, Q. et al. The CXCL8-CXCR1/2 pathways in cancer. Cytokine Growth Factor Rev. 31, 61–71 (2016).
Yuan, M. et al. Tumor-derived CXCL1 promotes lung cancer growth via recruitment of tumor-associated neutrophils. J. Immunol. Res. https://doi.org/10.1155/2016/6530410 (2016).
Sumida, K. et al. Anti-IL-6 receptor mAb eliminates myeloid-derived suppressor cells and inhibits tumor growth by enhancing T-cell responses. Eur. J. Immunol. 42, 2060–2072 (2012).
Najjar, Y. G. et al. Myeloid-derived suppressor cell subset accumulation in renal cell carcinoma parenchyma is associated with intratumoral expression of IL1β, IL8, CXCL5, and Mip-1α. Clin. Cancer Res. 23, 2346–2355 (2017).
Draghiciu, O., Nijman, H. W., Hoogeboom, B. N., Meijerhof, T. & Daemen, T. Sunitinib depletes myeloid-derived suppressor cells and synergizes with a cancer vaccine to enhance antigen-specific immune responses and tumor eradication. OncoImmunology 4, e989764 (2015).
Reck, M. et al. Primary PFS and safety analyses of a randomized phase III study of carboplatin + paclitaxel +/− bevacizumab, with or without atezolizumab in 1 L non-squamous metastatic NSCLC (IMpower150). Ann. Oncol. https://doi.org/10.1093/annonc/mdx760.002 (2017).
Rooney, M. S., Shukla, S. A., Wu, C. J., Getz, G. & Hacohen, N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 160, 48–61 (2015).
Miao, D. et al. Genomic correlates of response to immune checkpoint therapies in clear cell renal cell carcinoma. Science 359, 801–806 (2018).
Hsieh, J. J. et al. Genomic biomarkers of a randomized trial comparing first-line everolimus and sunitinib in patients with metastatic renal cell carcinoma. Eur. Urol. 71, 405–414 (2017).
Motzer, R. J. et al. Survival and prognostic stratification of 670 patients with advanced renal cell carcinoma. J. Clin. Oncol. 17, 2530–2540 (1999).
Wu, T. D. & Nacu, S. Fast and SNP-tolerant detection of complex variants and splicing in short reads. Bioinformatics 26, 873–881 (2010).
Wu, T. D., Reeder, J., Lawrence, M., Becker, G. & Brauer, M. J. GMAP and GSNAP for genomic sequence alignment: enhancements to speed, accuracy, and functionality. Methods Mol. Biol. 1418, 283–334 (2016).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLOS Comput. Biol. 9, e1003118 (2013).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Ritchie, M. E. et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Wilm, A. et al. LoFreq: a sequence-quality aware, ultra-sensitive variant caller for uncovering cell-population heterogeneity from high-throughput sequencing datasets. Nucleic Acids Res. 40, 11189–11201 (2012).
Saunders, C. T. et al. Strelka: accurate somatic small-variant calling from sequenced tumor-normal sample pairs. Bioinformatics 28, 1811–1817 (2012).
McLaren, W. et al. The Ensembl variant effect predictor. Genome Biol. 17, 122 (2016).
Lawrence, M., Degenhardt, J. & Gentleman, R. VariantTools: tools for working with genetic variants. version 1.12.0 Bioconductor https://bioconductor.org/packages/release/bioc/html/VariantTools.html (2018).
Karosiene, E., Lundegaard, C., Lund, O. & Nielsen, M. NetMHCcons: a consensus method for the major histocompatibility complex class I predictions. Immunogenetics 64, 177–186 (2012).
Shukla, S. A. et al. Comprehensive analysis of cancer-associated somatic mutations in class I HLA genes. Nat. Biotechnol. 33, 1152–1158 (2015).
Clopper, C. J. & Pearson, E. S. The use of confidence or fiducial limits illustrated in the case of the binomial. Biometrika 26, 404–413 (1934).
Acknowledgements
The authors thank A. Bailey for her contributions to development of the protocol and Z. Boyd for his contributions to the development of the PD-L1 IHC assay and its implementation in this study. Support for third-party writing assistance for this manuscript—by P.S. Davies of Health Interactions, Inc.—was provided by F. Hoffmann-La Roche, AG. This study was sponsored by F. Hoffmann-La Roche, AG. Authors were funded by NCI grants P50 CA101942-13 to D.F.M, M.B.A., and T.K.C.; P30 CA008748 to R.J.M.; and P30 CA14599 to W.M.S.
Author information
Authors and Affiliations
Contributions
D.F.M., M.B.A., R.J.M., B.I.R., B.E., and T.P. contributed to the conception, trial design, and data acquisition, analysis, and interpretation; T.P. was the principal investigator of the study; M.A.H., S.J., and D.N. performed biomarker analyses and interpretation; J.Q. supervised the analysis of the clinical data; M.A.H. and P.S.H. supervised the analysis of biomarker data; L.F., R.W.J., S.K.P., J.A.R., M.S., J.H., W.K.R., W.M.S., T.H., M.E.G., A.R., S.B., C. Suárez, V.G., T.K.C., D.N., A.T., C. Schiff, E.P.-L., R.D., and G.D.F. made substantial contributions to the acquisition of data and data analysis and interpretation; P.S.H., M.A.H., and D.S.C. had overall biomarker oversight; C. Schiff, G.D.F., and D.S.C. had overall medical oversight.
Corresponding author
Ethics declarations
Competing interests
D.F.M. reports a consulting/advisory role for Bristol-Myers Squibb, Merck, Roche/Genentech, Pfizer, Exelixis, Novartis, Eisai, X4 Pharmaceuticals, and Array BioPharma; and reports that his home institution receives research funding from Prometheus Laboratories. M.B.A. has been a paid consultant to Roche/Genentech, Bristol-Myers Squibb, Merck, Pfizer, Novartis, Exelixis, and Eisai. R.J.M. reports consulting fees from Roche/Genentech, Novartis, Pfizer, Eisai, and Exelixis, and research funds from Roche/Genentech, Bristol Myers Squibb, Pfizer, Novartis, Eisai, and Exelixis to the hospital for which he is employed. B.I.R. reports research funding to his institution from Roche/Genentech during the conduct of the study and grants/fees from Pfizer and Merck outside the submitted work. B.E. reports honoraria and research funding from Bristol-Myers Squibb, Novartis, Pfizer, and Ipsen; honoraria from Eusa Pharma, Roche, and Eisai; and research funding from Aveo. L.F. reports research funding to his institution from Roche/Genentech outside the submitted work. R.W.J. reports a consulting/advisory role with Bristol-Myers Squibb, Nektar, Genoptix, Eisai, Novartis, and Exilixis and research funding from Merck and Bristol-Myers Squibb; his home institution is in a consulting/advisory role with Merck and receives research funding from Roche/Genentech, X4 Pharmaceuticals, and Amgen. S.K.P. reports honoraria and a consulting/advisory role with Novartis, Astellas Pharma, Pfizer, Aveo, Myriad Pharmaceuticals, Roche/Genentech, Exelixis, Bristol-Myers Squibb, Ipsen, and Eisai and honoraria and research funding from Medivation. M.S. reports stock option interest in Amphivena Therapeutics, Intensity Therapeutics, and Adaptive Biotechnologies; a consulting role with Bristol-Myers Squibb, Roche/Genentech, AstraZeneca/MedImmune, Pfizer, Nektar, Lilly, Merck, Alexion Pharmaceuticals, Theravance, Baxalta/Shire, Seattle Genetics, Ignyta, Pierre Fabre, Incyte, Newlink Genetics, Celldex, Gritstone, and Innate Pharma; and an advisory role with Symphogen, Adaptimmune, Omniox, Lycera, and Molecular Partners. W.K.R. reports research funding to her home institution from Pfizer, Novartis, Tracon Pharmaceuticals, Bristol-Myers Squibb, Calithera Biosciences, and Peloton Therapeutics and research funding to an immediate family member from Incyte and Merck. W.M.S. reports honoraria and a consulting/advisory role with CVS Caremark, AstraZeneca, Bristol-Myers Squibb, Roche/Genentech, and Pfizer; research funding to his home institution from AstraZeneca, Bayer, Bristol-Myers Squibb, Boehringer Ingelheim, Exelixis, Novartis, Roche/Genentech, Pfizer, Merck, Janssen, and X4 Pharmaceuticals; and other relationships with UpToDate and American Cancer Society. M.E.G. acknowledges NHS funding to the NIHR Biomedical Research Centre at the Royal Marsden Hospital and Institute of Cancer Research, London UK. A.R. reports honoraria, accommodations, and a consulting/advisory role with Pfizer, Novartis, and Bristol-Myers Squibb; a consulting/advisory role with Ipsen and Roche; and research funding to his home institution from Pfizer and Novartis. S.B. reports personal fees and nonfinancial support for advisory boards from Pfizer, Astellas, Bristol-Myers Squibb, and Novartis; nonfinancial support for advisory boards from Bayer and Roche/Genentech; and nonfinancial support from Exelixis. V.G. reports grants from Bristol-Myers Squibb, Merck, Pfizer, and AstraZeneca, personal fees and nonfinancial support from Bristol-Myers Squibb, Merck, Roche, Novartis, Ipsen, Pfizer, AstraZeneca, Eisai, Eusa Pharma, and Cerulean outside the submitted work. T.K.C. reports consulting/advisory fees from AstraZeneca, Bayer, Bristol-Myers Squibb, Cerulean, Eisai, Foundation Medicine, Exelixis, Roche/Genentech, GlaxoSmithKline, Merck, Novartis, Peloton, Pfizer, Prometheus Laboratories, and Corvus; and research funding to his home institution from AstraZeneca, Bristol-Myers Squibb, Exelixis, Genentech, GlaxoSmithKline, Merck, Novartis, Peloton, Pfizer, Roche, Tracon, and Eisai. C. Schiff and P.S.H. report employment, including stock, with Genentech, Inc. G.D.F. reports employment, including stock, with Genentech, Inc. and stock with Foundation Medicine. T.P. reports honoraria and a consulting/advisory role with Roche/Genentech, Bristol-Myers Squibb, and Merck; a consulting/advisory role with AstraZeneca and Novartis; research funding from AstraZeneca/MedImmune and Roche/Genentech; and other relationships with Ipsen and Bristol-Myers Squibb (ASCO). M.A.H, D.N., S.J., E.P-L., J.Q., and D.S.C. are employees of Genentech, Inc. A.T. is an employee of Roche Products Ltd. J.A.R., J.H., T.H., C. Suárez, and R.D. have nothing to disclose.
Supplementary information
Supplementary Figures and Tables
Supplementary Figures 1–8 and Supplementary Tables 1–3
Rights and permissions
About this article
Cite this article
McDermott, D.F., Huseni, M.A., Atkins, M.B. et al. Clinical activity and molecular correlates of response to atezolizumab alone or in combination with bevacizumab versus sunitinib in renal cell carcinoma. Nat Med 24, 749–757 (2018). https://doi.org/10.1038/s41591-018-0053-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41591-018-0053-3
This article is cited by
-
Translational research on drug development and biomarker discovery for hepatocellular carcinoma
Journal of Biomedical Science (2024)
-
Multi-omics and immunogenomics analysis revealed PFKFB3 as a targetable hallmark and mediates sunitinib resistance in papillary renal cell carcinoma: in silico study with laboratory verification
European Journal of Medical Research (2024)
-
Five years of safety profile of bevacizumab: an analysis of real-world pharmacovigilance and randomized clinical trials
Journal of Pharmaceutical Health Care and Sciences (2024)
-
Antiangiogenic–immune-checkpoint inhibitor combinations: lessons from phase III clinical trials
Nature Reviews Clinical Oncology (2024)
-
Dual-loss of PBRM1 and RAD51 identifies hyper-sensitive subset patients to immunotherapy in clear cell renal cell carcinoma
Cancer Immunology, Immunotherapy (2024)