Abstract
Immune checkpoint blockade (ICB) targeting PD-1 and CTLA-4 has revolutionized cancer treatment. However, many cancers do not respond to ICB, prompting the search for additional strategies to achieve durable responses. G-protein-coupled receptors (GPCRs) are the most intensively studied drug targets but are underexplored in immuno-oncology. Here, we cross-integrated large singe-cell RNA-sequencing datasets from CD8+ T cells covering 19 distinct cancer types and identified an enrichment of Gαs-coupled GPCRs on exhausted CD8+ T cells. These include EP2, EP4, A2AR, β1AR and β2AR, all of which promote T cell dysfunction. We also developed transgenic mice expressing a chemogenetic CD8-restricted Gαs–DREADD to activate CD8-restricted Gαs signaling and show that a Gαs–PKA signaling axis promotes CD8+ T cell dysfunction and immunotherapy failure. These data indicate that Gαs–GPCRs are druggable immune checkpoints that might be targeted to enhance the response to ICB immunotherapies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Gene Expression Omnibus accession numbers for the human datasets used in this study include GSE116256, GSE114727, GSE110686, GSE14018, GSE123813, GSE22898, GSE123139, GSE115978, GSE146771, GSE134520, GSE140228, GSE125449, GSE117570, GSE127465, GSE99254, GSE123813, GSE156728, GSE120575, GSE84820, GSE123235, GSE122969, GSE88987 and GSE141299. European Genome–Phenome Archive identifiers for the human datasets used in this study include EGAS00001002171, EGAS00001002486, EGAS00001002325 and EGAS00001002553. Additional information regarding these datasets can be found in Supplementary Table 1. The remaining data are available within the article and supplementary information. Source data are provided with this paper.
References
Okazaki, T. & Honjo, T. PD-1 and PD-1 ligands: from discovery to clinical application. Int. Immunol. 19, 813–824 (2007).
Leach, D. R., Krummel, M. F. & Allison, J. P. Enhancement of antitumor immunity by CTLA-4 blockade. Science 271, 1734–1736 (1996).
Mellman, I., Coukos, G. & Dranoff, G. Cancer immunotherapy comes of age. Nature 480, 480–489 (2011).
Larkin, J. et al. Combined nivolumab and ipilimumab or monotherapy in untreated melanoma. N. Engl. J. Med. 373, 23–34 (2015).
Haslam, A. & Prasad, V. Estimation of the percentage of US patients with cancer who are eligible for and respond to checkpoint inhibitor immunotherapy drugs. JAMA Netw. Open 2, e192535 (2019).
Hauser, A. S., Attwood, M. M., Rask-Andersen, M., Schioth, H. B. & Gloriam, D. E. Trends in GPCR drug discovery: new agents, targets and indications. Nat. Rev. Drug Discov. 16, 829–842 (2017).
O’Hayre, M., Degese, M. S. & Gutkind, J. S. Novel insights into G protein and G protein-coupled receptor signaling in cancer. Curr. Opin. Cell Biol. 27, 126–135 (2014).
Wu, V. et al. Illuminating the Onco-GPCRome: novel G protein-coupled receptor-driven oncocrine networks and targets for cancer immunotherapy. J. Biol. Chem. 294, 11062–11086 (2019).
Pierce, K. L., Premont, R. T. & Lefkowitz, R. J. Seven-transmembrane receptors. Nat. Rev. Mol. Cell Biol. 3, 639–650 (2002).
Tokunaga, R. et al. CXCL9, CXCL10, CXCL11/CXCR3 axis for immune activation—a target for novel cancer therapy. Cancer Treat. Rev. 63, 40–47 (2018).
Ayers, M. et al. IFN-γ-related mRNA profile predicts clinical response to PD-1 blockade. J. Clin. Invest. 127, 2930–2940 (2017).
Yao, C. et al. Prostaglandin E2–EP4 signaling promotes immune inflammation through TH1 cell differentiation and TH17 cell expansion. Nat. Med. 15, 633–640 (2009).
Vigano, S. et al. Targeting adenosine in cancer immunotherapy to enhance T-cell function. Front. Immunol. 10, 925 (2019).
Valero, C. et al. Response rates to anti-PD-1 immunotherapy in microsatellite-stable solid tumors with 10 or more mutations per megabase. JAMA Oncol. 7, 739–743 (2021).
Andreatta, M. et al. Interpretation of T cell states from single-cell transcriptomics data using reference atlases. Nat. Commun. 12, 2965 (2021).
Nieto, P. et al. A single-cell tumor immune atlas for precision oncology. Genome Res. 31, 1913–1926 (2021).
Zheng, L. et al. Pan-cancer single-cell landscape of tumor-infiltrating T cells. Science 374, abe6474 (2021).
Li, H. et al. Dysfunctional CD8 T cells form a proliferative, dynamically regulated compartment within human melanoma. Cell 176, 775–789 (2019).
Zander, R. et al. CD4+ T cell help is required for the formation of a cytolytic CD8+ T cell subset that protects against chronic infection and cancer. Immunity 51, 1028–1042 (2019).
Wherry, E. J. et al. Molecular signature of CD8+ T cell exhaustion during chronic viral infection. Immunity 27, 670–684 (2007).
Wherry, E. J. & Kurachi, M. Molecular and cellular insights into T cell exhaustion. Nat. Rev. Immunol. 15, 486–499 (2015).
Inoue, A. et al. Illuminating G-protein-coupling selectivity of GPCRs. Cell 177, 1933–1947 (2019).
Liu, C. et al. Neuropilin-1 is a T cell memory checkpoint limiting long-term antitumor immunity. Nat. Immunol. 21, 1010–1021 (2020).
Andrews, L. P. et al. A Cre-driven allele-conditioning line to interrogate CD4. Immunity 54, 2209–2217 (2021).
Iglesias-Bartolome, R. et al. Inactivation of a Gαs–PKA tumour suppressor pathway in skin stem cells initiates basal-cell carcinogenesis. Nat. Cell Biol. 17, 793–803 (2015).
Roth, B. L. DREADDs for neuroscientists. Neuron 89, 683–694 (2016).
Nagai, Y. et al. Deschloroclozapine, a potent and selective chemogenetic actuator enables rapid neuronal and behavioral modulations in mice and monkeys. Nat. Neurosci. 23, 1157–1167 (2020).
Zhang, Y. et al. MKP-1 is necessary for T cell activation and function. J. Biol. Chem. 284, 30815–30824 (2009).
Lang, R. & Raffi, F. A. M. Dual-specificity phosphatases in immunity and infection: an update. Int. J. Mol. Sci. 20, 2710 (2019).
Seo, H. et al. TOX and TOX2 transcription factors cooperate with NR4A transcription factors to impose CD8. Proc. Natl Acad. Sci. USA 116, 12410–12415 (2019).
Thommen, D. S. & Schumacher, T. N. T cell dysfunction in cancer. Cancer Cell 33, 547–562 (2018).
Wang, Z. et al. Syngeneic animal models of tobacco-associated oral cancer reveal the activity of in situ anti-CTLA-4. Nat. Commun. 10, 5546 (2019).
Sade-Feldman, M. et al. Defining T cell states associated with response to checkpoint immunotherapy in melanoma. Cell 175, 998–1013 (2018).
Samstein, R. M. et al. Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat. Genet. 51, 202–206 (2019).
Sinha, N. et al. Immune Determinants of the Association between Tumor Mutational Burden and Immunotherapy Response across Cancer Types. Cancer Res. 82, 2076–2083 (2022).
Miller, B. C. et al. Subsets of exhausted CD8+ T cells differentially mediate tumor control and respond to checkpoint blockade. Nat. Immunol. 20, 326–336 (2019).
Zelenay, S. et al. Cyclooxygenase-dependent tumor growth through evasion of immunity. Cell 162, 1257–1270 (2015).
Bottcher, J. P. et al. NK cells stimulate recruitment of cDC1 into the tumor microenvironment promoting cancer immune control. Cell 172, 1022–1037 (2018).
Pelly, V. S. et al. Anti-inflammatory drugs remodel the tumor immune environment to enhance immune checkpoint blockade efficacy. Cancer Discov. 11, 2602–2619 (2021).
Daher, C. et al. Blockade of β-adrenergic receptors improves CD8+ T-cell priming and cancer vaccine efficacy. Cancer Immunol. Res. 7, 1849–1863 (2019).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Saddawi-Konefka, R. et al. Lymphatic-preserving treatment sequencing with immune checkpoint inhibition unleashes cDC1-dependent antitumor immunity in HNSCC. Nat. Commun. 13, 4298 (2022).
Belteki, G. et al. Conditional and inducible transgene expression in mice through the combinatorial use of Cre-mediated recombination and tetracycline induction. Nucleic Acids Res. 33, e51 (2005).
Akhmedov, D. et al. Gs–DREADD knock-in mice for tissue-specific, temporal stimulation of cyclic AMP signaling. Mol. Cell. Biol. 37, e00584-16 (2017).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Singh, G., Inoue, A., Gutkind, J. S., Russell, R. B. & Raimondi, F. PRECOG: predicting coupling probabilities of G-protein coupled receptors. Nucleic Acids Res. 47, W395–W401 (2019).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Valero, C. et al. Pretreatment neutrophil-to-lymphocyte ratio and mutational burden as biomarkers of tumor response to immune checkpoint inhibitors. Nat. Commun. 12, 729 (2021).
Acknowledgements
This project was supported by grants from the National Cancer Institute (R01CA247551 and U54CA209891) and National Institute of Dental and Craniofacial Research (NIH/NIDCR, R01DE026870 and U01DE028227). V.H.W. was supported by an NRSA Training Award (NIH/NCI 1F31CA250488-01). R.S.-K. was supported by an NRSA Training Award (NIH/NIDCR F32DE029990-01). B.S.Y. was supported by the Pharmacological Sciences Training Program (5T32GM007752-40) and an NRSA Training Award (NIH/NIDCR1F31DE031961-01). J.P.M. was supported by grants from the National Cancer Institute (U24CA248457 and U24CA220341). A.T.W. was supported by grants from the National Cancer Institute (F31CA257344 and U54CA209891) and the National Library of Medicine (T15LM011271). R.B. was supported by grants from the National Institutes of Health NIDDK (R01-DK092590) and NIAMS (R01-AR-072368). We acknowledge A. Sharabi (University of California, San Diego) for gifting the MC38-OVA cell line, D. Vignali (University of Pittsburg) for gifting the E8iCreErt2 mouse model, R. Iglesias-Bartolome (National Cancer Institute) for gifting the Gnas-exon 1fl/fl and the Tet-GFP-PKI mouse models and R. Berdeaux (The University of Texas) for gifting the ROSA26LSLGsDREADD mice. We acknowledge A. Goldrath for expertise and guidance in the study design and direction of the project.
Author information
Authors and Affiliations
Contributions
V.H.W. and B.S.Y. designed and conceived the studies, conducted most experiments described in this study and interpreted the data. F.F., Z.W. and R.S.-K. performed experiments described in the study. A.T.W., M.J.S., M.S.P., L.M.C., J.C., S.S., M.M., F.R., H.C., E.R. and J.P.M. all contributed to the bioinformatics analysis of this project. V.H.W., B.S.Y., F.F., R.S.-K., T.S.H., D.A.A.V., R.I.-B., R.B. and J.S.G. contributed to the study design and writing of the manuscript. J.S.G. provided oversight and direction of the entire project and study design, provided financial support for the study, interpreted the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
J.S.G. reports consulting fees from Domain Pharmaceuticals, Pangea Therapeutics and io9 and is founder of Kadima Pharmaceuticals, unrelated to the current study. R.B. is an employee and shareholder of CellChorus, Inc. J.C. is an employee and shareholder of Pfizer, Inc. V.H.W. is an employee and shareholder of Septerna, Inc. The other authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks William Murphy, Carla Rothlin, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: N. Bernard, in collaboration with the Nature Immunology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
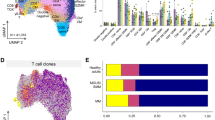
Extended Data Fig. 1 Patient and cancer information for integration analysis of CD8 T cells.
a, Information about patients, cancer type, and dataset used in the singe-cell RNA-seq integration. b, Visualization of 217,953 CD8 T cells after integration from 30 singe-cell RNA-seq datasets. c, Statistical comparison of calculated dysfunction score from tumor-infiltrating populations of CD8s characterized from Fig. 1b. Each dot represents one cell from groups listed in Supplementary Table 1a. Naïve: lower bound=0.147, middle bound=0.229, upper bound=0.337, 25th percentile=0.228, 75th percentile=0.231. Proliferating: lower bound=0.372, middle bound=0.564, upper bound=0.790, 25th percentile=0.556, 75th percentile=0.571. Cytotoxic: lower bound=0.184, middle bound=0.290, upper bound=0.447, 25th percentile=0.288, 75th percentile=0.292. Effector Memory: lower bound=0.209, middle bound=0.304, upper bound=0.431, 25th percentile=0.303, 75th percentile=0.305. Exhausted: lower bound=0.361, middle bound=0.555, upper bound=0.800, 25th percentile=0.551, 75th percentile=0.559.
Extended Data Fig. 2 Effect of targeting the Gαs/PKA signaling pathway in CD8 T cells.
a, Upregulation of inhibitory receptors and decrease of IFNγ and TNF in chronically versus acutely simulated CD8 T cells. The average relative expression and s.e.m. are shown (n = 6 biologically independent samples). b, Significant decrease of IFNγ or TNF with Gαs agonists in chronically stimulated CD8 T cells. The average relative expression and s.e.m. are shown (n = 6 biologically independent samples). c, Significant decrease of Ki-67 and viability with Gαs agonists in chronically stimulated CD8 T cells. The average relative expression and s.e.m. are shown (n = 6 biologically independent samples). d, Representative flow cytometry plots showing expression of Tim-3 and PD-1 in chronically stimulated CD8 T cells after treatment with 1 µM PGE2 (P), 5 µM Dobutamine (D), or 5 µM CGS-21860 (C). e, Effect of CXCL10 on PGE2-mediated decrease in IFNγ and TNFα The average frequency and s.e.m. are shown (n = 3 per group). Statistical significance was determined by two-way ANOVA. Unless indicated otherwise, statistical significance was determined by two-tailed unpaired Student’s t-test.
Extended Data Fig. 3 Development of a CD8-restricted PKI transgenic mouse model.
a, Scheme illustrating the generation of CD8-PKI mice. b, Genotyping information for CD8-PKI mice. c, Confirmation of PKI expression in CD4 or CD8 T cells isolated from splenocytes of CD8-PKI mice and littermate controls after induction by doxycycline and tamoxifen. The average relative expression and s.e.m. are shown (n = 6 mice per group). d, Quantification of IFNγ and TNF inhibition by PGE2 in chronically stimulated CD8 T cells from CD8-PKI mice. The average relative expression and s.e.m. are shown (n = 3 biologically independent samples). Statistical significance was determined by two-way ANOVA.
Extended Data Fig. 4 Development of a CD8-restricted Gαs-DREADD transgenic mouse model.
a, Genotyping confirmation for CD8-GsD mice. Primers detecting the Gαs-DREADD, ROSA26, and E8i-Cre were used to confirm recombination by the Cre-recombinase. Information about primers and genotyping is listed in Supplementary Table 4. b, Effect of DCZ on circulating CD8, CD4, NK cells, and CD11b myeloid cells in the peripheral blood of CD8-GsD mice treated with tamoxifen and 5 doses of DCZ (n = 5 mice for -DCZ; n = 6 mice for +DCZ). c, Effect of DCZ on non-tamoxifen-treated CD8-GsD mice. Quantification of IFNγ and TNF and PD-1 and Tim-3 in non-tamoxifen-treated CD8 T cells treated with or without DCZ. The average frequency and s.e.m. are shown (n = 3 biologically independent samples). Statistical significance was determined by two-tailed unpaired Student’s t-test.
Extended Data Fig. 5 Effect of tamoxifen and DCZ on tumor growth.
Tumor growth curve of CD8-GsD littermate control mice implanted with 4MOSC1 tumors treated with or without tamoxifen or DCZ. Mice were given 3 doses of tamoxifen, and 5 × 105 4MOSC1 cells were implanted into the tongue. Where indicated, 0.01 mg/kg DCZ was administered daily starting one day after tumor implantation.
Extended Data Fig. 6 Anti-CTLA-4 response in CD8-GsD mice bearing 4MOSC1 tumors.
Tumor growth curve (left panel) and survival plot (right panel) of CD8-GsD mice implanted with 4MOSC1 tumors treated with anti-CTLA-4 with or without DCZ (n = 7 mice per group). Mice were given three doses of tamoxifen before orthotopic tumor implantation and treated with checkpoint inhibitors and DCZ as previously described. Statistical significance was determined by two-way ANOVA. Statistical significance of survival data was calculated by the log-rank test.
Extended Data Fig. 7 Effect of Gnas deletion CD8 T cells in mice bearing 4MOSC1 tumors.
a, Tumor growth curve (left panel) and quantification of endpoint tumor volume (right panel) of CD8-Gnas+/+ (n = 14 mice) and CD8-Gnas−/− mice ( = 12 mice) implanted with 4MOSC1 tumors. Mice were given 3 doses of tamoxifen prior to orthotopic tumor implantation. The average tumor volume and s.e.m. are shown. Statistical significance was determined by two-way ANOVA. b, Quantification of PD-1+TIGIT+ CD8 T cells in 4MOSC1 tumors and draining lymph nodes at endpoint. The average frequency and s.e.m. are shown (n = 5 mice per group). Statistical significance was determined by two-tailed unpaired Student’s t-test. c, Frequency of CD8+ T cells in 4MOSC1 tumors in CD8-Gnas KO mice versus littermate controls. The average frequency and s.e.m. are shown (n = 5 mice per group). Statistical significance was determined by two-tailed unpaired Student’s t-test.
Extended Data Fig. 8 Gating strategy for chronic stimulation and in vivo experiments.
a, For chronic stimulation experiments, lymphocytes were gated from forward scatter area (FSC-A) and side scatter area (SSC-A). Single cells were distinguished from doublet cells in forward scatter height (FSC-H) and forward scatter width (FSC-W), and then side scatter height (SSC-H) and side scatter width (SSC-W). Live CD8 cells were then gated. b, For in vivo experiments, lymphocytes were gated from forward scatter area (FSC-A) and side scatter area (SSC-A). Single cells were distinguished from doublet cells in forward scatter height (FSC-H) and forward scatter width (FSC-W), and then side scatter height (SSC-H) and side scatter width (SSC-W). Live CD45 cells were then gated. T cells were distinguished by NK1.1 negative, CD19 negative, and CD3 positive. CD8 T cells were then gated as CD4 negative.
Supplementary information
Supplementary Table
Supplementary Tables 1–6.
Source data
Source Data Figs. 2–4 and Extended Data Figs. 1–7
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wu, V.H., Yung, B.S., Faraji, F. et al. The GPCR–Gαs–PKA signaling axis promotes T cell dysfunction and cancer immunotherapy failure. Nat Immunol 24, 1318–1330 (2023). https://doi.org/10.1038/s41590-023-01529-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-023-01529-7
This article is cited by
-
An adrenaline kick to exhaust T cells
Nature Immunology (2023)