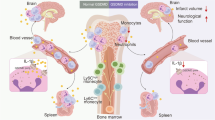
Abstract
Aging accounts for increased risk and dismal outcome of ischemic stroke. Here, we investigated the impact of age-related changes in the immune system on stroke. Upon experimental stroke, compared with young mice, aged mice had increased neutrophil clogging of the ischemic brain microcirculation, leading to worse no-reflow and outcomes. Aged mice showed an enhanced granulopoietic response to stroke that led to the accumulation of CD101+CD62Llo mature and CD177hiCD101loCD62Llo and CD177loCD101loCD62Lhi immature atypical neutrophils in the blood, endowed with increased oxidative stress, phagocytosis and procoagulant features. Production of CXCL3 by CD62Llo neutrophils of the aged had a key role in the development and pathogenicity of aging-associated neutrophils. Hematopoietic stem cell rejuvenation reverted aging-associated neutropoiesis and improved stroke outcome. In elderly patients with ischemic stroke, single-cell proteome profile of blood leukocytes identified CD62Llo neutrophil subsets associated with worse reperfusion and outcome. Our results unveil how stroke in aging leads to a dysregulated emergency granulopoiesis impacting neurological outcome.
Similar content being viewed by others
Main
Ischemic stroke often results in disability or mortality, despite recent advances in revascularization therapies. Age is the most prominent nonmodifiable risk factor for stroke1 and increased age is associated with worse clinical outcome2. Elements of the immune system are involved in every stage of the ischemic cascade3: stroke not only triggers a local immune response in the central nervous system (CNS)4, but also initiates systemic immune changes. Stroke directly activates hematopoietic stem cells (HSCs) through increased sympathetic tone and humoral signals5. Blood lymphocyte counts decline after stroke, while amounts of circulating neutrophils and monocytes increase in both mice and humans5,6. Myeloid cells that migrate to the lesioned brain tissue can contribute to reperfusion injury by plugging small capillaries, exacerbating the coagulation cascade locally or by direct endothelial damage7,8,9. Aging alters several features of the immune system. Several genetic and epigenetic factors associated with aging are known to reduce lymphopoiesis and increase myelopoiesis10,11. These changes are tolerated under homeostatic conditions but might lead to frailty in settings of overt inflammation, infections or neoplastic disease12. Increased occurrence of clonal hematopoiesis during aging was associated with greater vulnerability to cardiovascular disease13,14. However, the effects of age-related alterations in hematopoiesis and stroke outcome remain largely unexplored.
Here, we used experimental middle cerebral artery (MCA) occlusion in mice and blood samples from human stroke patients (SPs) to identify age-related alterations in stroke-induced granulopoiesis associated with worse outcome. Using several single-cell approaches, we found that early accumulation of age-specific mature and immature neutrophil subsets with prominent proinflammatory and prothrombotic features in the blood underlies the worse reperfusion and neurological outcomes seen in the elderly after stroke.
Results
Stroke in aged mice results in worse outcomes
To study ischemic stroke in aging, we subjected young (2–4 months) and aged (18–24 months) C57BL/6 male mice to transient MCA occlusion. Compared with young mice, aged mice suffered increased disability on the modified neurological stroke scale (mNSS) and a higher mortality rate after stroke (Fig. 1a,b and Extended Data Fig. 1a,b). Transcranial laser Doppler flowmetry (LDF) indicated a marked reduction in blood flow in the MCA area in mice during occlusion (Fig. 1c). Following recanalization (removal of the MCA occlusion), reperfusion was significantly impaired in aged compared with young mice (Fig. 1c). Of note, impairment of reperfusion correlated with mouse mNSS (Fig. 1d). At 6 h poststroke, aged mice displayed significantly more capillaries with entrapped erythrocytes (Fig. 1e) and with fibrinogen plugs (Fig. 1f) in the perilesional area compared with young mice. Intravenous injection of horseradish peroxidase (HRP) 6 h poststroke indicated significantly higher densities of capillary constrictions within the ischemic hemisphere of aged compared with young mice (Fig. 1g), suggesting increased microvascular occlusions and thrombosis leading to increase no-reflow in aged mice. We detected higher frequencies of TUNEL+NeuN+ apoptotic neurons at 6 h poststroke, and greater neuronal loss in the perilesional areas at 72 h poststroke in the brains of aged compared with young mice (Fig. 1h). Lesion volume in the cortex trended towards an increase in the aged compared with young mice, while the overall gross ischemic lesion volume was similar (Fig. 1i). Aged (18–24 months) female mice also suffered worse disability (mNSS) and mortality, increased perilesion neuronal loss and ischemic lesion volume upon stroke when compared with young (2–4 months) female mice (Extended Data Fig. 1c–e).
a, mNSS in young (2–4 months) and aged (18–24 months) C57Bl/6 male mice up to 3 days poststroke (two-way ANOVA, Sidak’s post hoc test; n = 15 mice per group; pool of three independent experiments, ****P ≤ 0.0001). b, Kaplan–Meier survival curve of young and aged stroke mice (as in a). (Mantel–Cox test, P = 0.045; n = 38 mice per group; pool of seven independent experiments). c, LDF recording after stroke onset (red arrow) and during recanalization (red box) (n = 32 young, n = 27 aged; pool of six independent experiments, mixed-effect model (REML), Benjamini–Hochberg post hoc test, P ≤ 0.0001). d, Integrated reperfusion slope and correlation with mNSS at day 3 poststroke (two-tailed t-test, ****P ≤ 0.0001; Spearman correlation; groups as in c). e, IF images of entrapped erythrocytes (NaBH4, red) in capillaries (tomato lectin+ vessels, green) of the ischemic cortex, at 6 h poststroke and quantification (n = 3 mice per group, **P = 0.0044). f, IF images of thrombosed capillaries (CD31+ red, fibrinogen+ white) in the ischemic cortex and quantification (n = 5 young, n = 4 aged mice, **P = 0.0053). g, Representative images of HRP staining of cortex capillaries at 6 h poststroke and quantification of node-like constrictions (arrowheads) (n = 6 young, n = 4 aged mice, *P = 0.0447). h, IF images of TUNEL+ cells (red) and neurons (NeuN+ green) in the ischemic cortex of young and aged stroke mice and quantification of apoptotic neurons (TUNEL+NeuN+) at 6 h (n = 3 per group; P = 0.0070) and surviving neurons (NeuN+) at 72 h poststroke in the perilesional area (n = 7 young, n = 6 aged, **P = 0.0041). i, Representative 3D reconstructions of the ischemic brain: cortical (dark gray) and striatal (green) ischemic lesion, healthy striatum (yellow) in young and aged mice, 72 h poststroke. Cortical, striatal and total ischemic volumes 72 h poststroke in young and aged mice (n = 10–15 mice per group; pool of three independent experiments). j, DHE+ cells in the perilesional area at 24 h poststroke (n = 6 young, n = 5 aged mice, **P = 0.003). k,l, DHE+ cell subtypes (n = 3 and 4 mice per group, *P = 0.046 and P = 0.030) (k) and representative IF for DHE+ cells (red) and CD45+ leukocytes (green) and quantification (n = 4 mice per group, **P = 0.0029) (l), 24 h poststroke in the perilesional area. In the scatterplots each dot corresponds to a single mouse; mean ± s.e.m. is shown. Two-tailed t-test if not otherwise specified; *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001, ****P ≤ 0.0001; scale bars, 50 µm (e–h,l). Nuclei stained by 4,6-diamidino-2-phenylindole (DAPI) (blue).
Nitrative and oxidative stress are among the leading causes of neuronal damage in ischemic brain tissue and can contribute to no-reflow15,16. Injection of dihydroethidium (DHE) to measure reactive oxygen species (ROS) indicated significantly higher frequencies of DHE+ cells in the perilesional area at 24 h poststroke in aged compared with young mice (Fig. 1j). The density of DHE+ leukocytes (DHE+CD45+) poststroke was increased in aged compared with young mice (60.6 versus 13.2 cells per mm2, respectively) between different cell types (Fig. 1k,l; Extended Data Fig. 1f–i). Staining of brain sections with 3-nitrotyrosine (3NT) found greater densities of 3NT+ cells in the ischemic hemisphere of aged compared with young mice at 72 h poststroke (Extended Data Fig. 1j). Overall, ischemic stroke in aged mice resulted in severe no-reflow, high levels of oxidative and nitrative stress, extensive neuronal loss and a poor outcome.
Neutrophil capillary plugging increases in aged stroke
Brain-infiltrating inflammatory cells can produce ROS and induce vessel and tissue damage9,15. High parametric multicolor flow cytometry (MCFC) showed that, at 24 h poststroke, aged mice had 30.8% neutrophils among infiltrating brain leukocytes, compared with 17.8% in young mice (Fig. 2a–c and Extended Data Fig. 2a,b), coupled with lower frequencies of natural killer (NK) cells and Ly6Clo monocytes (5.2% versus 12.9% and 3.6% versus 9.2%, respectively) (Fig. 2c). Immunofluorescence (IF) staining indicated the density of Ly6G+ neutrophils was greater in the ischemic brain of aged mice at both 6 h and 24 h poststroke (Fig. 2d). Ly6G+ neutrophils in the ischemic hemisphere were predominantly found within blood vessels in both age groups, with a significant increase in the proportion of extravasated Ly6G+ neutrophils in the aged at 24 h poststroke (Fig. 2e). In the aged ischemic brain, 7/4+ neutrophils were present in plugged vascular segments in association with fibrinogen/fibrin+ deposits and jammed Ter119+ erythrocytes (Fig. 2f,g). The release of citH3+ neutrophil extracellular traps (NETs) and the number of MPO+ neutrophils were increased at 48 h poststroke in aged mice. The NETosis ratio per neutrophil (NETs/MPO+cell) (Fig. 2h,i) and SiglecF+ eosinophils (Extended Data Fig. 2c) were not significantly different between groups. No marked immune cell infiltration was found in nonlesion hemisphere or other organs of stroke mice (Supplementary Table 1).
a, UMAP plot displaying nonmicroglial leukocyte clusters in the ischemic brains of young and aged mice at 24 h poststroke analyzed by 19-color flow cytometry. b, Heatmap showing the median antigen expression (MAE) values for each cell cluster (shown in a). c, Frequencies of nonmicroglial leukocytes (as in a, n = 5 mice per group, the boxes extend from the 25th to 75th percentiles, the midline denotes the median while whiskers plot minimum and maximum; *P = 0.038; **P = 0.0083; **P = 0.002, left to right). d, Ly6G+ neutrophils in the ischemic lesion and perilesional area at 0 h, 6 h and 24 h poststroke in young and aged mice (two-way ANOVA, Sidak’s post hoc test; n = 5 mice per group at 0 h; n = 4 per group at 6 h; n = 5 young mice, n = 4 aged mice at 24 h, from left to right: *P = 0.0458, *P = 0.0483, *P = 0.0134, ****P ≤ 0.0001, **P = 0.0098). e, Confocal microscopy images showing Ly6G+ neutrophils (green) in the ischemic tissue of young and aged mice at 24 h poststroke and within blood vessels (tomato lectin+, red,); donut plots showing the relative frequencies of neutrophils (Ly6G+ cells) inside (light green) and outside (dark green) blood vessels (BV) at 6 h and 24 h poststroke (n = 5 young, n = 4 aged mice, *P=0.0005). f, IF images in the ischemic tissue and quantification of vascular segments (CD31+, white) occluded by neutrophils (7/4+, green) at 6 h poststroke (n = 4 young, n = 4 aged mice; P = 0.0116). Yellow arrows indicate blood vessels occluded by neutrophils and fibrinogen (red). g, Representative images of neutrophils (Ly6G+, in green) in the ischemic tissue at 6 h poststroke in young and aged mice occluding blood vessels (tomato lectin+, white) that are also clogged by RBCs (TER119+ cells, in red). Yellow arrows indicate RBC/neutrophil occlusions. h,i, Representative IF images for NETs (citH3+, red) released by neutrophils (MPO+, green) in capillaries (tomato lectin+, white) of the ischemic brain of young and aged mice at 48 h poststroke (h) and quantification (i) (n = 8 young, n = 7 aged mice; MPO P = 0.0125; NET P = 0.0164). Nuclei in e, f, g and h are stained by DAPI (blue). Scale bars, 25 µm (e–h). In the scatterplots each dot corresponds to a single mouse; mean ± s.e.m. is shown. Two-tailed t-test if not otherwise specified, *P ≤ 0.05, **P ≤ 0.01, ****P ≤ 0.0001.
Quantification of blood leukocytes at 0 h, 6 h and 24 h poststroke using MCFC (Extended Data Fig. 2d–f) indicated that both the frequency and numbers of Ly6G+CXCR2+ neutrophils expanded poststroke in both aged and young mice, with higher frequencies and numbers at 6 h and 24 h in aged mice compared with young mice (Extended Data Fig. 2f and Supplementary Table 2), while the frequencies and numbers of CD4+ and CD8+ T cells, B cells, NK cells, plasmacytoid dendritic cells and Ly6Clo monocytes were lower in young mice (Extended Data Fig. 2f and Supplementary Table 2). Aged mice depleted for neutrophils (with the anti-Ly6G antibody, 1A8) at stroke onset (Extended Data Fig. 3a) showed significant improvement of LDF measured reperfusion, fewer NaBH4+ erythrocyte-plugged brain capillaries at 6 h and fewer brain-infiltrating 7/4+ neutrophils at 72 h poststroke, decreased disability on mNSS and significantly improved perilesional neuronal numbers compared with isotype-treated mice (Extended Data Fig. 3b–f). As such, stroke in aged mice resulted in blood neutrophilia and increased infiltration of neutrophils to the ischemic brain, which leads to the plugging of the ischemic microvasculature.
Stroke in aged mice associates with atypical neutrophils
We next used single-cell RNA sequencing (scRNA-seq) on sorted CD45+CD11b+ cells from the blood of young and aged mice 24 h poststroke to dissect the transcriptional heterogeneity of neutrophils (Supplementary Fig. 1a). After rigorous quality controls, we obtained 5,793 high-quality cells with an average of 999 genes per cell (total of 11,691 genes). Unbiased, graph-based clustering identified seven main distinct cell populations. A total of 3,852 cells clustered as neutrophils (Ly6g, Csfr3, Ngp) (Fig. 3a,b, Supplementary Table 3 and Supplementary Fig. 1b), which, based on gene expression patterns, clustered into six transcriptionally distinct clusters: scNeu1 (Mmp8, Cd177, Ly6c2, Lgals3) and scNeu2 (Cd14, Tagln2, Cxcl2, Wfdc17) were more represented in blood of the aged mice, while scNeu3 (Lst1, Gm2a, Il1b, Csf3r) was increased in the young mice (Fig. 3b,c and Extended Data Fig. 3g). The smaller clusters were represented by scNeu4 (Ifit1, Ifi27l2a, Isg15, Irf7), which expressed interferon-stimulated genes17,18,19, scNeu-Plt (Pf4, Ppbp, Ctla2a, Cd9), which probably represented neutrophil-platelet conjugates18, and scNeu-Ery (Gypa, Fech, Bpg, Hbb-bt), probably comprising neutrophils that phagocytosed erythrocytes (Fig. 3b–e). A data integration approach (INT)20 confirmed the three main neutrophil subsets (scNeu1-INT, scNeu2-INT and scNeu3-INT; Supplementary Fig. 1d–i).
a, UMAP of the 5,793 myeloid cells obtained by scRNA-seq of sorted blood CD45+CD11b+ cells (see Supplementary Fig.1a) at 24 h poststroke from young and aged mice featuring the expression of the neutrophil-specific genes Ly6G and Csf3r. b, UMAPs of extracted neutrophils (from a) colored by inferred cluster identity, combined and separated for young and aged mice. The boxed cluster (in a) of 3,852 neutrophils is extracted, reclustered and analyzed. c, Graphs showing the proportion of each neutrophil cluster among neutrophils as in b (chi-square test, ****P ≤ 0.0001). d, Heatmap showing the row-scaled relative expression level of the ten highest DEGs (P ≤ 0.05) per cluster as in b. e, Dot plot for scaled expression of the gene signature for each neutrophil cluster as in b. f, Heatmap showing the row-scaled relative expression level of the genes used to calculate the BM proximity score and violin plot (n = 1,670 scNeu1, n = 1,156, scNeu2, n = 812, scNeu3, n = 105, scNeu4, n = 87, scNeu-Plt, n = 22 cells, scNeu-Ery, one experiment). g, Dot plot showing the correlation score between neutrophil clusters (scNeu1, scNeu2, scNeu3, scNeu4 as in b) and scRNA-seq-defined neutrophil populations and progenitors reported by Xie et al.17 that is, HSPC, G0, G1, G2, G3, G4, G5a, G5b and G5c. h, Violin plot for neutrophil maturation score of neutrophil clusters (as in f). i, Heatmap showing the row-scaled expression of ‘neutrophil aging’-related genes as reported by Adrover et al.7 in neutrophil clusters (as in b). j, UMAP showing the weighted average expression of ‘neutrophil aging’-related genes in neutrophils (as in b) and violin plot of neutrophil aging score for neutrophil clusters (as in b). k, Proportion of aged neutrophil cells in scNeu1, scNeu2 and scNeu3 clusters (as in j). (****P ≤ 0.000, chi-square test). l–n, Violin plots of ROS production and NADPH oxidase score (l), NETs-association score (m) and phagocytosis scores for the neutrophil clusters (n) as in f. o, Heatmap of hierarchically clustered regulon activity (t-value versus other clusters) obtained with SCENIC for the neutrophil clusters (as in g) is shown. Only regulons with an absolute value of relative activity >1.2 are visualized. p, Heatmap showing the expression of selected transcription factor-related genes for neutrophil clusters (as in o). In f, h, j, l, m and n, Bonferroni-corrected Wilcoxon rank sum test, (*P ≤ 0.05, **P ≤ 0.01, ****P ≤ 0.0001); all statistical comparisons for violin plots are reported in Supplementary Table 3; boxes in violin plots extend from the 25th to 75th percentiles, midline denotes the median.
We next focused on the main clusters scNeu1, scNeu2 and scNeu3 and interrogated the scRNA-seq dataset using transcriptional signatures-based scores. Because scNeu1 displayed the highest bone marrow (BM) proximity score18, probably reflecting recent egress from the BM (Fig. 3f) we compared our neutrophil clustering structure with the signature of a published scRNA-seq dataset17 that had mapped nine maturing neutrophil stages: hematopoietic stem and progenitor cells (HSPC), common myeloid progenitor (CMP)/granulocyte monocyte progenitor (GMP) (G0), proNeu (G1), preNeu (G2), immature neutrophils (G3), BM maturing neutrophils (G4) and peripheral mature blood neutrophils (G5a, G5b, G5c) (Fig. 3g). ScNeu1 correlated with G4, while scNeu2 and scNeu3 displayed some similarity to G5a, G5c and G5b (Fig. 3g). Neutrophils mobilized from the BM after stroke may undergo activation17, maturation17 or ‘aging’ in the circulation21. scNeu1 displayed among the main clusters a higher activation score (Extended Data Fig. 3h), a lower maturation score (Fig. 3h), a higher ‘aging’ score (Fig. 3i,j and Extended Data Fig. 3i) and a lower chemotaxis score (GO:0030593; Extended Data Fig. 3j). By applying a two-component mixture model, we found that 36.6% of scNeu1 corresponded to ‘aging’ neutrophils, more than that present in scNeu2 (15.9%) or scNeu3 (12.1%) (Fig. 3k).
Interrogation of the transcriptional programs of neutrophil effector functions, including granule biosynthesis22, oxidative burst, phagocytosis and NETosis indicated that scNeu1, scNeu2 and scNeu3 had a relatively low score for primary (azurophil) and secretory vesicles, but an higher score for specific and tertiary (gelatinase) granules (Extended Data Fig. 3k,l), with scNeu1 displaying the highest compared with the other neutrophil subsets (Extended Data Fig. 3l). Further scNeu1 had a significantly higher ROS production, NADPH oxidase and NETs-association score compared with scNeu2 and scNeu3 (Fig. 3l,m), while the phagocytosis score was higher for scNeu2 (Fig. 3n). The correlation index between functional scores for each cell within scNeu1 indicated significant correlations between NAPDH oxidase and ROS scores, and between NAPDH oxidase and NETosis scores (Extended Data Fig. 3m,n and Supplementary Table 3). There was no overt difference within scNeu1 between neutrophils from aged and young samples in terms of correlation of functional scores (Supplementary Table 3).
Single-cell regulatory network inference and clustering (SCENIC) analysis indicated that the scNeu1, scNeu2, scNeu3 and scNeu4 clusters were characterized by specific gene regulatory networks (Fig. 3o,p and Supplementary Table 3). scNeu1 showed negative activity for the Fosl2 regulon, a transcription factor upregulated during maturation23 and a positive regulatory activity for Cebpe—a master regulator of early neutrophil development24—as well as for Foxo3, which is associated to survival of neutrophils in inflammation25 (Fig. 3o,p). scNeu2 regulatory networks controlled mainly proinflammatory functions (Ets2, Klf2, Xbp1) and effector properties, including degranulation (Psmd12)26. scNeu3 expressed gene regulatory networks associated with canonical neutrophil maturation (Junb27, Fos, Elf1 and Spi1 (ref. 28)) and effector functions (Jund, Irf1, Stat6 (ref. 17,23,29)), (Fig. 3o,p). ScNeu4 displayed enrichment for IRF regulons (Irf9, Irf7, Irf2), defined as late-maturation and activated neutrophil17 transcription factors (Fig. 3o,p). scNeu3 and scNeu4 downregulated the Cebpa regulon (Fig. 3o), which is associated with neutrophil commitment in early myelopoiesis30. Of note, expression of transcription-factor-related genes across clusters was consistent with the relative regulatory network activity enrichments (Fig. 3o). After stroke, blood neutrophil subsets in aged mice exhibited a transcriptional program linked to effector functions, including oxidative stress and NETosis, as well as a BM proximity signature, indicating premature release from the BM.
Aged emergency granulopoiesis is dysregulated poststroke
We used a neutrophil-specific MCFC panel that targeted signature molecules emerging from the scRNA-seq neutrophil analysis to characterize granulopoiesis during stroke in aging. The PHATE algorithm, a dimensionality reduction method for visualizing trajectory structures in high-dimensional data, charted a continuous process of neutrophil differentiation and maturation (Fig. 4a) from proNeu2 in the BM to mature CD101+CD62Lhi neutrophils (designated as cluster Neu3a) in the blood. This process was characterized by the downregulation of CD11b, Ly6C, CXCR4 and CD177 and the upregulation of Ly6G, followed by the upregulation of CXCR2, CD101 and CD62L (Fig. 4b,c). In the blood, Neu3a were present at the steady state (0 h) in both age groups, and evolved to CD101+CD62Llo neutrophils (Neu3b) poststroke (Fig. 4c). Neu3b accumulated rapidly in the blood 6 h poststroke and were increased significantly in aged compared with young mice (Fig. 4c,d and Extended Data Fig. 4a). At 24 h poststroke, two clusters of CD101lo neutrophils, CD177hiCD101loCD62Llo (Neu1) and CD177loCD101loCD62Lhi (Neu2), emerged in the blood of aged mice (Fig. 4c,d and Extended Data Fig. 4a), partially resembling, based on the expression of CD101, CD62L, CD177, Ly6C, CXCR2 and CD11b, the expression profile of scNeu1 and scNeu2 clusters, respectively (Extended Data Fig. 4b). Accordingly, neutrophils retrieved from the brain of aged mice 24 h poststroke showed higher CD177 and lower CD101 median fluorescence intensity by MCFC compared with young mice, resembling blood Neu1 (Extended Data Fig. 4c). Furthermore, immunofluorescence staining in aged mice confirmed the presence of CD177+Ly6G+, CD101-Ly6G+ and CD14+7/4+ neutrophils within the peri-ischemic area (Extended Data Fig. 4d).
a, Dimensionality reduction method for visualizing trajectory using the PHATE algorithm of blood live CD45+CD11b+Ly6G+ neutrophils at baseline (0 h), 6 h and 24 h poststroke and BM live CD45+CD11b+ neutrophils and direct precursors at 0 h and 24 h poststroke combining young and aged mice (n = 4 or 5 mice per group per timepoint). b, Heatmap displaying the MAE of markers used to identify neutrophil subsets as in a. c, PHATE dimensionality reduction method plotting blood neutrophils of young and aged mice at 0 h, 6 h and 24 h poststroke (n = 6 young (Y), n = 4 aged (A) mice at 0 h and 24 h, n = 5 per group at 6 h). d, Graphs displaying dynamic changes in the subset composition of the circulating neutrophil population over time poststroke and barplots of the relative frequencies of blood neutrophil subsets (groups as in c, shown is the mean, centerline ± s.d., colored area). e, PHATE dimensionality reduction method plotting BM neutrophils of young and aged mice at 0 and 24 h poststroke (n = 6 young, n = 4 aged per timepoint). f, Barplots showing the relative frequencies of neutrophils among total BM neutrophils and among mature BM neutrophils (as in e). In d and f, two-way-ANOVA, Bonferroni post hoc test; one of two independent experiments is shown; *P ≤ 0.05, **P ≤ 0.01, ***P ≤ 0.001, ****P ≤ 0.0001.
MCFC of CD11b+Ly6G–CXCR2– neutrophil progenitors and CD11b+Ly6GloCXCR2– immature neutrophils in the BM (Fig. 4a,b,e) indicated a lower frequency of CD11b+Ly6G–CD177hiCD49d+ proNeu2 and significantly higher frequencies of CD11b+Ly6G–CD177+ preNeu and Ly6Glo neutrophils in aged compared with young mice at 24 h poststroke (Fig. 4e,f and Extended Data Fig. 4e), suggestive of an accelerated neutropoiesis in aged BM. Alongside, mature CXCR2+ neutrophils, namely Neu2, Neu3a and Neu3b, were reduced among the BM neutrophils in the aged compared with young mice (Fig. 4e,f and Extended Data Fig. 4e), suggesting a concurrent, inverse kinetic to that observed in the blood. Consistently, the mature neutrophil compartment of the BM showed a precocious (6 h compared with 0 h poststroke) contraction in the frequency of Neu3a, which was more pronounced in the aged mice (Extended Data Fig. 4f). Higher concentrations of neutrophil growth and mobilization factors (G-CSF, CXCL1) and proinflammatory mediators (IL-6, IP10/CXCL10) were detected in the plasma of aged mice compared with young mice (Extended Data Fig. 4g). These findings indicated that the emergency granulopoiesis response to stroke was dysregulated in aged mice and lead to a rapid accumulation of proinflammatory neutrophils in the blood and ischemic brain, along with the gradual depletion of the neutrophil BM reservoir.
Age-related HSC changes drive early neutrophil differentiation
To investigate whether age affected the hematopoietic progenitor response to stroke, we interrogated the Lin–c-kit+BM progenitor compartment in aged and young mice at steady state and 24 h poststroke using MCFC and algorithm-guided analysis. We found higher frequencies of CMP and GMP and significantly lower frequencies of lymphoid-biased multipotent progenitors (MPP)4, and CLP at steady state (Extended Data Fig. 5a–c) indicating myeloid-biased hematopoiesis in aged mice. At 24 h poststroke, aged mice displayed a significant increase of long-term (LT)-HSC, short term (ST)-HSC and the myeloid-biased MPP3/MPP2, as well as further contraction of CLP and a significant increase in GMP and direct progenies (common monocyte progenitors, cMOP) compared with young mice (Extended Data Fig. 5c), indicating the diffuse activation of the Lin–SCA-1+c-Kit+ compartment and a stronger myeloid shift.
Next, we performed scRNA-seq analysis on sorted CD45+CD11b+ cells from the blood of young and aged mice at steady state and Gr1+ and c-kit+-enriched cells from the BM of young and aged mice at steady state and 24 h poststroke (Supplementary Figs. 2a–c and 3a,b). Rigorous quality controls were performed on individual samples (Supplementary Table 4) and, for further quality assurance, the steady-state young and aged BM dataset were integrated with a published dataset from the Tabula Muris Senis31. Unsupervised clustering revealed a similar clustering structure and a strong correlation among the average gene expression per cluster between the two datasets (Supplementary Fig. 4a–g). Next we integrated20 our blood and BM scRNA-seq datasets of steady state and poststroke young and aged mice. We obtained 33,664 high-quality cells with a total of 25,017 mouse genes detected in at least five cells (Supplementary Figs. 2a–h and 3b–h and Supplementary Table 4). Unbiased, graph-based clustering identified 27 main cell populations. To specifically interrogate neutropoiesis, we restricted our analysis to 18,154 cells annotated as HSPC compartment, myeloid progenitors and neutrophils (Supplementary Figs. 2c and 3b). Unsupervised clustering partitioned differentiating and mature neutrophils into 13 clusters representing the cell state continuum from hematopoietic stem cells to the mature neutrophil (Extended Data Fig. 6a–c). Annotation according to their similarity to previously mapped neutrophil subsets (HSPC and G0-G5)17, based on a GSVA-based score, indicated that the dataset covered HSPC (cluster 14), granulocyte monocyte progenitors (G0_CMP and G0_CMP, cluster 6 and 3), proneutrophils (G1, cluster 9 and 12), preneutrophils (G2, cluster 7 and 10), immature neutrophils (G3, G4, cluster 8 and 2) and mature neutrophils (G5a, G5b, G5c, cluster 0, 13, 1, 5)17,24 (Extended Data Fig. 6d). Aged mice had an increased frequency of HSPC clusters at steady state and poststroke and an increased proportion of G4 and G5a neutrophils in the blood at steady state compared with young mice, with a further increase poststroke (Extended Data Fig. 6e).
Using Monocle32, we inferred a linear differentiation trajectory for the integrated blood and BM dataset starting from the HSPC cluster, through G0 (CMP) to G0 (GMP) continuing, after branching towards monocyte-committed (cMOP), to neutrophil-committed G1(proNeu), G2 (preNeu), G3 and G4 (immature neutrophils in the BM), to -G4- G5a (immature neutrophils in blood) and G5a-c (mature neutrophils of the blood) (Extended Data Fig. 6f). Several transcripts showed a different pattern of expression along the pseudotime between aged and young mice at steady state or poststroke (Extended Data Fig. 6g). Ifitm1, a marker of HSPC quiescence and G-CSF signaling in myeloid progenitors33, and the tetraspanin Cd63, a marker of early myeloid differentiation24 and involved in granule targeting34, increased early in pseudotime in aged mice, both in steady state and poststroke (Extended Data Fig. 6g); Cebpe, which is associated with granulocyte commitment, Cd14 and the chemokine Cxcl2, involved in thrombo-inflammation7, were increased in aged mice poststroke (Extended Data Fig. 6g), while the effector genes Elane and Prtn3 were increased in the late differentiation states (G2–G3) in aged mice at steady state (Extended Data Fig. 6g).
Differential gene expression (DGE) analysis indicated that aged HSPC and G0 displayed higher expression of genes associated with myeloid bias and HSPC-aging, such as Nupr1, Sult1a1, Selp, Mt1, Aldh1a1 and Vwf35 at steady state—a characteristic retained poststroke (Extended Data Fig. 7a–c). The stem score, based on a Hlf-expressing LT-HSC signature33, was increased, while the proliferation score33 was lower in aged compared with young HSPCs at steady state (Extended Data Fig. 7d). G-CSF signature score was increased in aged HSPCs at steady state and poststroke compared with young HSPCs whereas, in the young, G-CSF signature and proliferation score33 were highly increased in G1 and G2 poststroke (Extended Data Fig. 7d). Poststroke, HSPC and G0–G1 progenitors upregulated genes expressed only in more differentiated stages (G2–G4; for example Ngp, Camp, CD117) or in mature neutrophils (G5, for example Cxcr2, S100a8 and S100a9) (Extended Data Fig. 7b). Mature neutrophil signature genes (Elane, Retnlg), which are induced at G1 and G4, respectively, in the young, were found increased in cluster G0–G1 in the BM of the aged (Extended Data Fig. 7b,c). Mature neutrophil subsets of the blood upregulated genes associated with thrombo-inflammation, such as Mmp8, Cxcl2, Cxcl3, Prok2 and Cd14, in aged mice poststroke (Extended Data Fig. 7e and Supplementary Table 4). Our results demonstrated a transcriptional bias towards myelopoiesis in early progenitor cells of aged mice, which was further enhanced poststroke.
BM rejuvenation ameliorates stroke
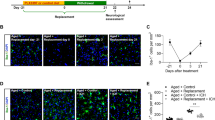
To assess the contribution of aged hematopoietic progenitors to altered neutrophil generation, we transplanted total BM cells from young (YtoA chimeras) or aged (AtoA chimeras) mice into lethally irradiated aged mice. At week 6 postengraftment (0 h), YtoA mice had fewer Neu3a neutrophils but more lymphoid cells in the blood (Fig. 5a,b) and more preNeu and Ly6Glo Neu neutrophil precursors, but fewer Neu1 and Neu3a mature neutrophils in the BM compared with AtoA mice (Fig. 5c,d). Further, there was an increase in MPP4 and CLPs and a significant reduction of GMPs in YtoA compared with AtoA (Fig. 5c,d). At 24 h poststroke, compared with AtoA mice, YtoA had a significantly lower frequency of Neu1 neutrophils in the blood (Fig. 5a,b) and of GMPs in the BM (Fig. 5e), higher reperfusion values (Fig. 5f), lower mNSS disability scores and a nonsignificant trend towards lower mortality (Fig. 5g,h). Immunoflourescence stainings indicated significantly lower Ly6G+ neutrophil infiltration in the ischemic hemisphere (Fig. 5i), lower number of 3NT+ cells in the ischemic brain (Fig. 5j) and significantly less neuronal loss (Fig. 5k) in YtoA mice compared with AtoA mice. As such, rejuvenation of the hematopoietic compartment rebalanced the subsets of circulating neutrophils and ameliorated stroke outcome.
a,b, Barplots showing the frequencies of blood neutrophil subsets among total neutrophils (a) and graphs showing the frequencies of blood neutrophil subsets, lymphoid cells and monocytes among CD45+ cells at 0 h and 24 h poststroke (b) in irradiated aged chimeric mice that received young (YtoA) or aged (AtoA) total BM (YtoA, n = 3; AtoA, n = 4 at 0 h and n = 5 per group at 24 h, two-way ANOVA with Sidak post hoc test, Neu1, **P = 0.0021, Neu3a **P = 0.009; #P = 0.0738). c–e, Barplots of relative frequencies of neutrophil subsets among BM neutrophils (c) and graphs showing the frequencies of hematopoietic stem and progenitor cells at 0 h (§P = 0.09, #P = 0.0794, *P = 0.0367) (d) and 24 h (e) poststroke (#P = 0.06) in AtoA and YtoA mice as in b. f, LDF recordings during stroke induction and upon reperfusion in YtoA and AtoA mice (YtoA, n = 11; AtoA, n = 10, two-way ANOVA, Benjamini–Hochberg posttest, one experiment, mean ± s.e.m., *P ≤ 0.05). g,h, Disability mNSS score (g) and Kaplan–Meier survival curve (h) up to 3 days poststroke in YtoA and AtoA mice (two-way ANOVA with Sidak’s post hoc test, **P ≤ 0.01 and Mantel–Cox test for survival, P = 0.1474; n = 11 mice per group). i, Representative confocal microscopy images of neutrophils (Ly6G+, green) and blood vessels (tomato lectin+, red) in the perilesional area at 72 h poststroke and quantification of neutrophils, (AtoA, n = 7; YtoA, n = 8; P = 0.044). Scale bar, 50 µm. j,k, 3NT+ cells (AtoA, n = 7; YtoA, n = 7, **P = 0.0055) (j) and quantification of neurons (NeuN+ cells) (AtoA, n = 8; YtoA, n = 9, **P = 0.0047) (k) in the perilesional area of mice at 72 h poststroke. In b, boxes extend from the 25th to 75th percentiles, the midline denotes the median and whiskers plot minimum and maximum. Dots in barplots correspond to a single mouse, mean ± s.e.m. is shown. Two-tailed t-test, if not otherwise indicated. *P ≤ 0.05; **P ≤ 0.01.
Aging-associated neutrophil transfer worsens stroke
We next analyzed the production of ROS, NETosis, platelet conjugate formation and phagocytosis in blood neutrophil subsets from both aged and young mice using MCFC. Neu1 from aged mice showed increased ROS production and phagocytosis compared with young mice (Extended Data Fig. 8a,b). Neutrophil-platelets aggregates (NPA, Ly6G+CD41+) and NETing neutrophils (CitH3+MPO+) were increased in aged mice compared with young mice 24 h poststroke (Extended Data Fig. 8c–f), with Neu3a and Neu2 displaying the highest NPA and NETing activity, respectively (Extended Data Fig. 8d,f). Neu1 and Neu3b neutrophils, sorted from the blood of young and aged mice, exhibited a mainly multilobulated and hypersegmented nuclear morphology36, independent of donor origin (Extended Data Fig. 8g). On poststroke blood smears, aged mice had a significantly higher frequency of hypersegmented neutrophils with a prominent multilobulated nucleus compared with young mice (Extended Data Fig. 8h). To test the role of aging-associated neutrophils in stroke in vivo, we injected young recipients intravenously at stroke onset with Ly6G+ neutrophils from either the BM of young (YtoY) or aged (AtoY) mice. Neutrophils from aged donors were enriched in CD101loCD62LloNeu1 and CD101+CD62LloNeu3b subsets (Fig. 6a,b). AtoY mice displayed significantly worse reperfusion (on LDF) and disability (mNSS) (Fig. 6c,d), increased lesion volume (Fig. 6e), perilesional neuronal loss (Fig. 6f), brain neutrophil infiltration (Fig. 6g) and frequencies of both total and hypersegmented neutrophils in the blood 24 h poststroke compared with YtoY (Fig. 6h). Transferred neutrophils were able to reach the ipsilesional hemisphere (Extended Data Fig. 8i,j). Similarly, the transfer of blood leukocytes, from aged mice 24 h poststroke to young mice at stroke onset, resulted in worse reperfusion (LDF) and mNSS score, increased perilesional neuronal loss and edema compared with the transfer of neutrophil-depleted blood leukocytes from aged mice (Extended Data Fig. 8k–m). PEdKO mice, which lack P- and E-selectins, have augmented circulating CD62Llo neutrophils37. Transfer of neutrophils from steady-state PEdKO mice to young mice at stroke onset (NKOtoY) worsened reperfusion, mNSS score and perilesional neurons loss, but not the ischemic volume compared with mice transferred with neutrophils from young WT mice (NtoY) (Extended Data Fig. 8n,o). Thus, the transfer of aging-associated CD62Llo neutrophils was sufficient to worsen no-reflow, brain tissue damage and neurological outcomes of experimental ischemic stroke.
a, Representative flow cytometry plots of BM neutrophils (after negative selection using magnetic beads) from young (Y donor) or aged (A donor) mice used for adoptive intravenous transfer into young recipients with stroke (YtoY or AtoY, respectively, in b–h). b, Barplots showing the relative frequencies of neutrophil subsets among BM neutrophils isolated from Y donors (n=6) and A donors (n=3). c,d, LDF recordings of YtoY and AtoY adoptively transferred mice after stroke induction and reperfusion (two-way ANOVA, Benjamini–Hochberg post hoc test, *P ≤ 0.05) (c), and mNSS disability score up to 3 days poststroke (mixed-effect model, Sidak’s post hoc test, **P = 0.0094, ***P = 0.0003, **P = 0.0010) (d) in mice as in a (YtoY, n = 8; AtoY, n = 6). e,f, Representative serial 3D reconstructions of mouse brains at 3 days poststroke (same colors as in Fig. 1i) and ischemic volume quantification by Cresyl Violet staining (e) and IF images and quantification of neurons (NeuN+, red) in the peri-ischemic area (**P = 0.0036, ***P = 0.0008, YtoY, n = 8; AtoY, n = 5) of mice (f) as in a. g, Representative IF images and quantification of infiltrating neutrophils (Ly6G+, green) in the ischemic brain at day 3 poststroke (blood vessels stained by tomato lectin+ are in red, YtoY, n = 8; AtoY, n = 5, ***P = 0.0004). h, Frequencies of lymphocytes, neutrophils and hypersegmented neutrophils 24 h poststroke quantified by peripheral blood smear analysis (YtoY, n = 8; AtoY, n = 6, *P ≤ 0.05). In the scatterplots each dot corresponds to a single mouse; mean ± s.e.m. is shown. Two-tailed t-test if not otherwise indicated, *P ≤ 0.05; **P ≤ 0.01; ***P ≤ 0.001. Scale bars, 50 µm f,g).
Aging-associated neutrophils are prothrombotic in stroke
To further dissect the pathogenic transcriptional program of neutrophils of aged mice, we performed deep bulk RNA-seq in CD62Llo neutrophils sorted from the blood of young and aged mice at steady state (0 h) and 24 h poststroke. Unsupervised hierarchical clustering separated neutrophils from aged and young mice under both conditions (Fig. 7a). DGE analysis, at 0 h, identified 97 upregulated genes and 75 downregulated genes (false discovery rate (FDR) ≤0.05; Fig. 7a); at 24 h poststroke, there were 329 upregulated genes and 252 downregulated genes in CD62Llo neutrophils from aged mice compared with young mice (FDR ≤ 0.05; Fig. 7a). At 24 h poststroke, CD62Llo neutrophils from aged mice had higher expression of genes associated with neutrophil migration and chemotaxis (for example, Cxcl3, Olfm4, Lbp), metallopeptidase genes (for example, Mmp8, Mmp19, Mmp23 and Adam22), genes involved in coagulation and thrombosis (for example, Factor X (F10), Factor V (F5), Thbs1, Annexin A5 (Anxa5), Tlr4, Hemolytic complement (Hc)) (Fig. 7b,c) and genes involved in the superoxide metabolic pathway, including Nos2, compared with their younger counterparts (Supplementary Table 6). Consensus binding sequence analysis revealed that transcription factors involved in inflammatory responses, including AP-1, Sp1, Nrf2, OCT1 (refs. 29,38,39) or transcription factors expressed by immature neutrophils (CEBPa, CEBP_Q2 and Nfe2)17 were enriched in the aged at 24 h poststroke (Fig. 7d). The projection of the bulk RNA aging-associated neutrophil signature score on the blood poststroke scRNA-seq dataset indicated that scNeu1 cluster, prevalent in aged mice, had the highest bulk signature score (Fig. 7e and Supplementary Table 3).
a, Unsupervised hierarchical clustering of bulk-RNA-seq data from sorted CD62Llo blood neutrophils of young and aged mice at 0 h and 24 h poststroke showing the relative expression level of the significantly DEGs between aged and young mice (FDR ≤ 0.05, adjusted P value (Padj) ≤ 0.05, n = 4 mice per group). b, Dot plot showing GO analysis (Biological Process, Canonical Pathways and Molecular Function terms) of DEGs (as in a). Selected GO terms (in red) at 24 h poststroke are reported in c. c, GSEA of selected GO terms (as in b). Genes were ranked by log2 fold change (LogFC). Color-coded ratio of aged over young mice (Deseq2 model). d, Transcription factor enrichment analysis of DEGs (as in a), color-coded ratio of aged over young mice. e, UMAP and barplot showing the neutrophil gene expression signature generated from the bulk RNA-seq data (selected DEGs at 24 h poststroke), projected onto the main neutrophil clusters (scNeu1, scNeu2 and scNeu3) defined by scRNA-seq (as in Fig. 3b) (Bonferroni-corrected Wilcoxon rank sum test, ****P ≤ 0.0001). f, Volcano plot of DEGs from bulk-RNA-seq data (as in a)(DEGs above or below 1.5 log2FC are color labeled, red upregulated, blue downregulated). g,h, Cd101, Cd177, Cd14 (*P ≤ 0.05; **P ≤ 0.01; ***P ≤ 0.001) (g) and Cxcl3 expression (h) in blood CD62Llo neutrophils (as in a, *P = 0.0172, **P = 0.0035). i, Confocal microscopy images of blood neutrophils (Ly6G+, white) at 24 h poststroke showing expression of CXCL3+ (green) and ROS (DHE+, in red). j, Representative IF image of CXCL3+ (in red) neutrophils (Ly6G+, in green) infiltrating the ischemic lesion of an aged mouse at 24 h poststroke. k, CXCL3 levels in plasma of young and aged mice 48 h poststroke (n = 8 mice/group, *P = 0.045, one experiment). In the scatterplots each dot corresponds to a single mouse; mean ± s.e.m. is shown. In k two-tailed t-test; in g, h, two-way ANOVA with Sidak’s post hoc test. Scale bar, 25 µm (i, j). In i and j, cell nuclei stained with DAPI (blue).
Aged poststroke CD62Llo neutrophils had lower expression of Cd101 and higher expression of Cd177 and Cd14 (Fig. 7f,g), consistent with their relative immaturity and higher expression of genes involved in the inflammatory response, including Cxcl3, Prok2 (ref. 40) (encoding prokineticin 2), Thbs1 (encoding thrombospondin 1) and Selp (P-selectin) (Fig. 7f). The gene set enrichment analysis (GSEA) for the signaling pathway genes of CXCR2, the CXCL3 cognate receptor, trended toward enrichment in poststroke aged CD62Llo neutrophils compared with young mice (Extended Data Fig. 9a) and the CXCR2 signature score was highest on blood stroke scRNA-seq neutrophils in scNeu2 (Extended Data Fig. 9a). Cxcl3 mRNA expression had the highest LogFC (3.73) between aged and young mice in poststroke CD62Llo neutrophils (Fig. 7f,h), and this correlated with protein expression (Fig. 7i–k).
To test the role of CXCL3 on neutrophils, we added recombinant CXCL3 to ex vivo blood neutrophils. CD62L and CXCR2 median expression measured by MCFC was significantly reduced in CXCL3-stimulated compared with unstimulated neutrophils (Extended Data Fig. 9b,c). At 2 h postintravenous injection of CXCL3 in young mice, we observed an increase of blood neutrophils and in particular of Neu2 and Neu3b compared with vehicle-injected mice (Extended Data Fig. 9d–g). Young mice injected intravenously with CXCL3 had significantly worse reperfusion (LDF) and mNSS score, increased neuronal death, Ly6G+ neutrophil infiltration and 3-NT+cells in the brain (Extended Data Fig. 9h–k) compared with vehicle-injected controls. Conversely, the administration of an CXCL3 neutralizing antibody before and after stroke onset in aged mice (Extended Data Fig. 9l–p) significantly ameliorated no-reflow and disability (Extended Data Fig. 9l,m), neuronal loss (Extended Data Fig. 9n), brain edema (Extended Data Fig. 9o) and neutrophil infiltration in the brain (Extended Data Fig. 9p). Furthermore, antibody neutralization of CXCL3 in young mice receiving intravenous Ly6G+ neutrophils from aged BM at stroke onset improved mNSS score, LDF, decreased neuronal loss and neutrophil infiltration, with no effect on the overall ischemic volume (Extended Data Fig. 9q–u). Taken together, these results suggested that CD62Llo neutrophils derived from aged mice express genes that promote proinflammatory and prothrombotic functions, such as Cxcl3.
Neutrophil heterogeneity is a biomarker in human stroke
We next investigated the association of neutrophil subtypes with stroke outcome in aged humans. We applied single-cell 37-parameter mass cytometry by time-of-flight (CyTOF) to blood samples collected during the first 24 h (D1), at day 3 or 7 (D3, D7) poststroke from young (mean 60.6 ± 12.7 years, n = 9) and old (86.2 ± 3.7 years, n = 23) stroke patients(SP) (young SP and old SP), and in young (64.7 ± 5.8 years, n = 12) and old (82.4 ± 2.3 years, n = 5) age-matched controls (MC) (young MC and old MC) (Supplementary Table 7). Using the uniform manifold approximation and projection (UMAP) dimensionality reduction41 and FlowSOM-guided clustering42,43, we identified nine blood leukocyte populations (Extended Data Fig. 10a–d). To focus on the heterogeneity of neutrophils, we repeated FlowSOM-guided clustering and manual annotations and assigned human blood neutrophils to six neutrophil subsets (Fig. 8a,b). Among these, CD24–CXCR4– and CD49d+ neutrophils were rare subsets of immature neutrophils22 corresponding to putative promyelocytes and metamyelocytes, respectively. The other four main populations of neutrophils, defined by differential expression of CD101, CD62L and CD177 (CD66b+CD177hiCD101loCD62LloCXCR4+ hNeu1, CD66b+CD177loCD101loCD62LhiCXCR2lo hNeu2, CD66b+CD177loCD101+CD62LhiCXCR4+ hNeu3a, CD66b+CD177hiCD101+CD62LloCXCR4+ hNeu3b) resembled Neu1, Neu2, Neu3a and Neu3b in mice (Fig. 8a–c). hNeu3a neutrophils increased significantly after stroke in old SP at D1 and D3 (Fig. 8d,e). hNeu3b were increased significantly at D1 in old SP compared with old MC (Fig. 8d,e). hNeu3b were increased in old SP compared with young SP at D1 and D3, and returned similar to young SP and MC at D7 (Fig. 8d,e). Compared with young SP, old SP patients showed an increase of hNeu2 at D7, while the frequency was reduced at D1 (Fig. 8d,e).
a, Representative UMAP showing the supervised FlowSOM clustering of CD66b+ neutrophils and antigen markers based on mass cytometry analysis of blood CD66b+neutrophils from young and old SPs (young SP, n = 9; old SP, n = 23) at day (D) 1, D3 and D7 poststroke and in young and old MC (young MC, n = 12; old MC, n = 5). b, Heatmap displaying the MAE intensity of markers used to generate the UMAP as in a. c, Heatmaps showing the MAE of shared neutrophil antigens targeted by mouse flow cytometry (from Fig. 4) and human CyTOF panels respectively in mouse blood and human blood neutrophil subsets as in a. d, Donut graphs showing relative frequencies of neutrophil subsets (among total neutrophils) (groups as in a). e, Violin plots displaying absolute cells counts of neutrophil subsets (groups as in a). f, Absolute cell counts and frequencies among total neutrophils of CD62LloCXCR4+ neutrophils (CD45+CD16+CD15+CD14-CD62LloCXCR4+ cells) of blood samples from young SP (n = 34), old SP (n = 48) at D1, D3 and D7 poststroke, and from young MC (n = 25) and old MC (n = 19). g, Violin plot of CD62LloCXCR4+ neutrophil counts in blood from SPs with good (mRS 0–3) and poor (mRS 4–6) functional outcome at 3 months poststroke, and ROC of CD62LloCXCR4+ neutrophils at D3 poststroke for predicting poor functional outcome at 3 months. h, Violin plot of CD62LloCXCR4+ neutrophil counts in SPs undergoing endovascular thrombectomy with complete (mTICI 3) or partial (mTICI 0-2b) reperfusion. i, Average thrombograms recorded for FV- and FVIII-deficient plasma in presence of 3 × 106 ml–1 fMLP-stimulated neutrophils, from healthy controls with reduced frequency of CD62Llo neutrophils (HC[lo]CD62LloNeu, n = 5), SPs with either reduced frequency of CD62Llo neutrophils (SP[lo]CD62LloNeu, n = 5) or high frequency (SP[hi] Cd62Llo Neu, n = 6) or with high frequency of CD62Llo neutrophils and thrombomodulin (SP[hi]CD62LloNeu-TM, n = 2). j, Histograms displaying Lagtime, the ETP, the peak and the time to peak (ttPeak) (three groups as in j). Dots represent individual samples. e,f, Multiple two-tailed Mann–Whitney; *refers to comparisons between SP-young and SP-old. For comparison of young-SP or old-SP with respectively young-MC or old-MC Kruskall–Wallis with Dunnʼs post hoc test: ^refers to comparisons with MC-young; #refers to comparisons with MC-old; *,^,#P ≤ 0.05, **,^^,## P ≤ 0.01, ***,^^^,### P ≤ 0.001, ****,#### P ≤ 0.0001. g,h, Two-tailed Mann–Whitney. j, Kruskall–Wallis test with Dunnʼs multiple comparison test, *P ≤ 0.05.
Next we used a clinically applicable eight-fluorochrome flow cytometry panel on blood samples of a larger cohort of young and old SPs (young SP, mean 56.1 ± 14.9 years, n = 34; old SP, mean 87.2 ± 5.2 years, n = 48) and MCs (young MC, mean 57.0 ± 7.0 years, n = 25; old MC, mean 85.4 ± 4.2 years, n = 19) (Supplementary Table 8). Old SP had significantly higher numbers of total circulating neutrophils at D1 and D3 poststroke compared with old MC and young SP, together with significantly lower lymphocyte counts, resulting in a higher neutrophil-to-lymphocyte ratio6 (Extended Data Fig. 10e). Further CD62LloCXCR4+ neutrophils were significantly more abundant in the blood of SP compared with MC at all timepoints, and in old SP compared with young SP at D1 and D3 (Fig. 8f). Multivariate analysis showed that the frequency of CD62LloCXCR4+ neutrophils at D1 and D3 after stroke remained independently associated with patient age after adjustments for sex, hypertension, atrial fibrillation, stroke severity and thrombolysis administration (respectively, adjusted odds ratio(aOR), aOR 2.60 per quartile, 95% confidence interval (CI), 1.10–6.14; aOR 2.10 per quartile, 95% CI 1.22–3.61).
Considering clinical characteristics, patients with poor functional outcomes at 3 months poststroke had significantly more abundant CD62LloCXCR4+ neutrophils at D3 (Fig. 8g). The receiving operating curve (ROC) of CD62LloCXCR4+ neutrophils at D3 for predicting poor functional outcome showed an area under the curve (AUC) of 0.695 (Fig. 8g). Patients who achieved only partial reperfusion after revascularization treatment for endovascular thrombectomy had significantly higher CD62LloCXCR4+ neutrophil counts at D3 compared with those with complete reperfusion (Fig. 8h).
We next sought to validate in human neutrophils the coagulation pathway emerged in the transcriptional analysis of aged mouse CD62Llo neutrophil subsets. Thrombin generation was monitored in factor V (FV)- and factor VIII (FVIII)-deficient plasma by adding purified platelet-free-neutrophils from healthy controls (HC), with low frequencies of CD62Llo neutrophils (HC[lo]CD62LloNeu), or neutrophils from SPs with either high or low frequency of CD62Llo neutrophils (SP[hi]CD62LloNeu or SP[lo]CD62LloNeu) (Fig. 8i,j and Extended Data Fig. 10f,g). Thrombin was generated in all neutrophil preparations; however, peak velocity of thrombin generation and endogenous thrombin potential (ETP) were significantly higher and lagtime significantly faster in SP[hi]CD62LloNeu compared with the other groups (Fig. 8i,j). Activated protein C, generated by addition of thrombomodulin (TM) to the deficient plasma-neutrophil reaction mixtures, abrogated thrombin generation, by targeting factor V/Va, in SP[hi]CD62LloNeu (Fig. 8i). Thus, increased CD62Llo neutrophil levels in aged SPs were linked to poorer outcomes and reperfusion and these neutrophils promoted thrombin formation through factor V provision, accelerating the coagulation pathway.
Discussion
Here, we found that stroke in aged mice was characterized by worse no-reflow phenomenon, followed by increased neuronal loss, disability and mortality. This impaired outcome was due to stroke-evoked emergency granulopoiesis that, in the aged, leads to the release of atypical neutrophils endowed with increased ROS production, and procoagulant features contributing to the occlusion of brain capillaries.
Neutrophils have been identified as key contributors to the development of no-reflow phenomenon in vascular disorders9,15,44 as well as in chronic neurological conditions such as Alzheimer’s disease45. Although the neutrophil-to-lymphocyte ratio correlates with clinical prognosis in stroke6, trials of neutrophil-targeted therapeutics in stroke have demonstrated little benefit, probably due the incomplete understanding and targeting of the functional heterogeneity of neutrophils46, and the rapid dynamic changes of the granulopoietic response together with the heterogeneity of SPs enrolled. The functional role of neutrophil subsets is emerging46,47. For instance, CD62Llo ‘senescent’ neutrophils, which persist longer in the circulation compared with CD62Lhi neutrophils7,21,48 while endowed with tissue homeostasis functions, are highly detrimental in pathological conditions such as sickle-cell disease48, acute myocardial infarction7,18 and other neurodegenerative diseases49. In experimental stroke, aged mice displayed increased counts and heterogeneity of blood neutrophils, characterized by a rapid accumulation of CD101+CD62Llo(Neu3b) mature neutrophils, followed by the emergence and predominance of CD177hiCD101loCD62Llo(Neu1) and CD177loCD101loCD62Lhi(Neu2) immature neutrophils with, at the same time, an erosion of BM neutrophil subsets compared with younger mice. Of note, we found a corresponding age-related heterogeneity and a similar kinetic of blood neutrophils in patients with stroke. These observations support a model in which stroke in the elderly initiates massive and persistent mobilization of immature proinflammatory granulocytes from the BM. In analogy, a recent study on emergency granulopoiesis triggered by acute bacterial infection reported that infection can shortcircuit the release and maturation of BM immature neutrophils directly in the blood, changing the neutrophil maturation dynamics17. Severe infections, such as COVID-19, lead to the emergence of hypersegmented neutrophils in the blood50 and patients with severe COVID-19 accumulate immature neutrophils in blood51 and lungs52, which might facilitate immunothrombosis53.
Functionally, in experimental stroke, aging-associated neutrophils displayed a peculiar transcriptional signature encompassing gene sets related to immaturity and senescence as well as to enhanced effector functions, such as oxidative burst, NETosis and phagocytosis as well as coagulation processes. Indeed, the transfer of aging-associated neutrophils to young mice was sufficient to phenocopy the worse stroke outcome observed in the elderly. We showed that, in stroke, CD62Llo neutrophils in the aged produce CXCL3 and that CXCL3 leads to the emergence of the CD62Llo neutrophils functionally contributing to worse outcomes in stroke. CXCL3 binds to CXCR2 (ref. 54) and triggers neutrophil mobilization from the BM55. CXCR2 signaling is part of an autocrine loop initiating the transition to a ‘senescent neutrophil’ stage in the context of the circadian rhythm7. A subset of tissue-infiltrating neutrophils with high expression of Cxcl3 has been described in lung-tumor-bearing mice19 and in mouse myocardial infarction18.
Using single-cell transcriptomics, we observed an anticipation of transcriptional programs of granulopoiesis in the aged that culminated in the development and early mobilization of inflammatory and prothrombotic neutrophil subsets. Sustained high levels of G-CSF, CXCL1 and CXCL3 that circulate after stroke in the elderly might, in fact, contribute to boosting granulopoiesis. These evidence mechanistic link the age-related myeloid skewing of hematopoiesis to the aging-associated atypical neutrophils responsible for the worsening of tissue damage in stroke in the elderly.
Consequently, normalization of the neutrophil response could become a prominent therapeutic strategy for cerebrovascular diseases. A broad intervention like BM rejuvenation was able to revert age-related granulopoietic adaptations and to improve the outcome of experimental stroke; harvesting from our and future phenotypic, molecular and functional kinetic studies of neutrophil differentiation and mobilization will pave the way to the development of effective and more selective approaches to rebalance the deranged granulopoiesis of the aged or to interfere in a timely manner with the emergence of pathogenic neutrophil subsets.
Methods
Animal model
All experiments were approved by the Institutional Animal Care and Use Committee (IACUC nr. 581 and 798 at the IRCCS, San Raffaele Hospital Milano (Italy)). Young (2–4 months) and aged (18–24 months) adult male and female C57BL/6 mice were purchased from Charles River Laboratories and Jackson Laboratories. Young adult (2–4 months) male C57BL/6-Ly5.1 were purchased from Charles River Laboratories. Female mice were analyzed in Extended Data Fig. 1c–e; all other ischemia experiments were performed on male mice. PEdKO LySM-eGFP mice were kindly provided by P. Frenette37, bred and maintained at the IRCCS, San Raffaele Hospital animal facility. Mice were housed socially (three to five mice per cage) on a 12 h light/dark cycle in individually ventilated cages and with ad libitum access to food and water. The experimental holding room had temperature (21.5 °C set point) and humidity control (40% set point) and was supplied with HEPA-filtered air. Only animals without any signs of scratches, and with no overt skin lesions were included in the study. Experimental procedures were all performed during the middle of the day in a blinded fashion for treatment.
Human samples
This study was approved by the Institutional Review Board at IRCCS, San Raffaele Hospital (San Raffaele Ethics Committee) and has been performed in accordance with the ethical standards laid down in the 1964 Declaration of Helsinki and its later amendments. Written informed consent was obtained from all patients and controls. The study prospectively included patients with acute ischemic stroke admitted to the Emergency Department and the Stroke Unit of IRCCS San Raffaele Hospital, Milan, Italy, between January 2018 and December 2019. All patients were treated according to international and national guidelines. Patients aged ≥18 years, with a diagnosis of ischemic stroke and presented within 24 h from symptoms onset, were included.
Patients with signs or symptoms of infection on admission, or a history of immunological or hematological disease, were excluded. Participants were categorized according to age as ‘young’ (≤70 years) and ‘old’ (≥80 years) SPs. Information collected prospectively for each patient included: demographic data, clinical history and vascular risk factors, acute stroke treatments, classification of stroke etiology, stroke severity on admission and at discharge (assessed by the National Institute of Health Stroke Scale (NIHSS)), occurrence of hemorrhagic transformation, functional outcome assessed at discharge and at 3 months from stroke onset (assessed by the modified Rankin scale (mRS)). ‘Young’ (≤70 years) and ‘old’ (≥80 years) MC subjects, without history of ischemic stroke, acute infectious diseases, immunological or hematological diseases, were included (Supplementary Table 8). Laboratory measurements were made without knowledge of the participants’ status or age group.
Proximal MCA occlusion
All experimental stroke mice underwent 45-min left MCA occlusion (MCAO)56. Briefly, animals were anesthetized with 1–1.5% isofluorane (Merial) in 30% O2; their temperature was maintained between 36.0 and 36.5 °C and LDF (PeriFlux System 5000, Perimed) was used for monitoring the procedure. Focal cerebral ischemia of the left MCA was induced with a 7-0 silicon-coated filament (Doccol). Animals that successfully reached a stable cerebral blood flow below 30% of their initial baseline were included in the study. Animals that experienced bleeding during surgery or other complications were excluded from the study. Laser Doppler recordings during surgery that had saturated, negative values or other technical registration problems (for example, detachment of probe during ischemia) were excluded from LDF analysis.
Behavioral analyses
The mNSS grades neurologic function on a scale of 0 to 12 (normal score 0; maximal deficit score 12). mNSS is a composite of motor and balance tests56. Behavioral tests were performed by a researcher blinded to the treatment group during the light phase of the circadian cycle beginning 4 h after lights were turned on.
Mouse brain tissue preparation for flow cytometry
Mice were anesthetized, perfused with saline and brain tissue removed. The forebrain (separated from the bulbi, cerebellum and brainstem) was divided into a left ipsilateral (ischemic) hemisphere and a right (non-ischemic) contralateral hemisphere. Each hemisphere was processed separately. Brain tissues were cut with scissors into small pieces and incubated with 0.4 mg ml–1 of collagenase type IV (Sigma-Aldrich) for 30 min at 37 °C. After incubation, they were passed repeatedly through a 19 G needle with a syringe to obtain a homogeneous cell suspension, filtered through a 70 μm mesh, suspended in PBS and washed. Tissue homogenate was loaded onto an isotonic 90% Percoll (GE Heathcare) gradient for enrichment of CNS infiltrates57. Brain cell suspensions enriched on the Percoll gradient were then labeled and analyzed by flow cytometry58 using a FACS Symphony (BD Biosciences), with data analysis by FlowJo software (Tree Star). The whole sample was processed. For brain analysis, cells were always gated on single and live cells by excluding dead cells with the live/dead Zombie NIR Fixable Viability Kit (Biolegend).
Mouse blood sample preparation for flow cytometry
Blood samples were collected in EDTA. Whole blood was immediately labeled for 30 min at room temperature, red blood cells (RBCs) were lysed in ACK Lysing Buffer (Gibco), and remaining cells were analyzed using a FACS Canto II or FACS Symphony (BD Biosciences). Data analysis was carried out using FlowJo software (Tree Star). Leukocytes were identified on manual FlowJo based analysis (Fig. 6a, Extended Data Figs. 3a, 8a–g,i,n and 9b–g and Supplementary Table 2) as neutrophils (CD45+CD11b+Ly6G+ or CD45+CD11b+CXCR2+), further gated as mature (CD45+CD11b+Ly6G+CD101+) and immature neutrophils (CD45+CD11b+Ly6G+CD101–), then gated as Neu1 (CD177hiCD101loCD62Llo), Neu2 (CD177loCD101loCD62Lhi), Neu3a (CD101+CD62Lhi) and Neu3b (CD101+CD62Llo). Ly6Clo monocytes were gated as (CD45+CD11b+Ly6G–Ly6Clo), Ly6Chi monocytes (CD45+CD11b+Ly6G–Ly6Chi), eosinophils (CD45+CD11b+Ly6G–Ly6CintSSC-Ahi), lymphocytes (CD45+CD11b–) and T cells (CD45+CD11b–CD3+). For the 18-marker panel the cell clusters were categorized using self-organizing maps (FlowSOM)42,43.
Absolute counts of each white blood cell (WBC) population were obtained by multiplying the percentages of CD45+ cells calculated by flow cytometry analyses by the WBC counts obtained using a certified hemocytometer (ProCyte Dx, Idexx Laboratories).
Mouse BM preparation for flow cytometry
BM was flushed from tibiae and femurs through a 27G syringe with PBS/1% BSA and filtered through a 40 µm cell strainer. Cells were harvested, RBCs were lysed in ACK Lysing buffer (Gibco) and remaining cells were labeled for 30 min at 4 °C. Cell data were acquired on a FACS Canto II or FACS Symphony (BD Biosciences) and analyzed with FlowJo software (Tree Star). HSPCs were identified by FlowSom-guided clustering. Viability was assessed with Zombie Aqua Fixable Viability Kit (Biolegend).
BM transplantation
Five million BM cells from 2- to 4-month-old or 17-month-old wild type (WT) mice were injected intravenously into sublethally irradiated (11 Gy ref. 59)) 17-month-old WT mice. Blood chimerism was tested after 5 weeks, and stroke was induced 6 weeks after BM reconstitution.
In vivo neutrophil depletion
Young and aged WT mice were injected intravenously with 400 µg of anti-Ly6G (in vivo plus anti-mouse Ly6G, 1A8, Bioxcell) or isotype control antibody (Rat IGg2b, Bioxcell) 30 min before stroke onset.
Neutrophil adoptive transfer experiments
Neutrophils were obtained from the BM of young and aged mice. BM was flushed from tibiae and femurs through a 27G syringe with cold PBS/0.5% BSA and filtered through 40 µm cell strainer. Cells were harvested, RBCs lysed in ACK Lysing buffer (Gibco) and neutrophils were isolated by negative selecti with the ‘Neutrophil Isolation Kit’ (Miltenyi) according to the manufacturer’s instructions. Qualitative analysis of sorted neutrophils was performed by flow cytometry before transfer. Five million neutrophils were injected intravenously into young mice a few minutes before and 24 h poststroke.
For transfer of blood leukocytes enriched with the diverse neutrophils’ subsets, young or aged WT donor female C57BL/6N mice at 24 h after stroke were injected intraperitoneally with 5 mg kg–1 of AMD-3100 (Sigma-Aldrich) 2 h before bleeding as described7.
For transfer of leukocytes enriched with CD62Llo neutrophils, untreated PEdKO LySM-eGFP mice were bled. For transfer of leukocytes depleted of CD62Llo neutrophils, female PEdKO LySM-eGFP mice were injected intravenously with 300 µg of anti-Gr1 (Bioxcell, clone RB6-8C5,) 3 h before bleeding. For all transfers, whole blood was collected, lysed in ACK lysing buffer (Gibco) and cells washed in PBS. WBCs were quantified with a ProCyte Dx hemocytometer (Idexx Laboratories) and 2.5 × 106 neutrophils were injected intravenously into young WT male recipient mice 30 min before and 24 h poststroke. Because anti-Gr1-treated PEdKO LySM-eGFP had almost no circulating neutrophils, the same volume of blood as untreated PEdKO LySM-eGFP was collected, treated as above and injected.
For transfer of leukocytes depleted of neutrophils, donor blood was treated with Anti-Ly-6G MicroBeads UltraPure, mouse (Miltenyi), and the neutrophil-deprived flow-through was injected as described above.
Transferred neutrophil tracing
Young WT CD45.1 recipient mice were injected intravenously twice with 5 million BM neutrophils from either young or aged WT CD45.2 donors 30 min before and 24 h poststroke. At 2 days poststroke, the ischemic forebrain and contralateral hemisphere were analyzed separately and donor neutrophils quantified by flow cytometry. Alternatively, BM neutrophils were labeled with CellTracker Green CMFDA Dye (Thermo Fisher) 10 µM, according to the manufacturer’s instructions. The recipient’s brains were analyzed by immunofluorescence 48 h poststroke.
CXCL3 administration to healthy mice
Recombinant mouse CXCL3 (150 µg kg–1; R&D Systems) or vehicle (PBS with 0.1% BSA) was injected intravenously into young WT male mice, which were subsequently bled at 2 h, 4 h and 6 h after administration.
CXCL3 administration in MCAO
Recombinant mouse CXCL3 (150 µg kg–1; R&D Systems) or vehicle (PBS + 0.1% BSA) were injected intravenously into young WT male mice 2 h before, 24 h and 48 h poststroke.
In vivo CXCL3 blockade
For in vivo CXCL3 blockade, aged mice were injected intraperitoneally twice with 100 µg anti-CXCL3 (Thermo Fisher) or isotype the night before (–17 h) and the same day (–5 h) of stroke induction7. Young mice receiving BM neutrophils from aged mice, were injected intraperitoneally twice with 100 µg of anti-CXCL3 30 min before and at 24 h poststroke.
Ex vivo CXCL3 treatment of mouse neutrophils
Blood was collected into EDTA, RBCs were lysed in ACK lysing buffer (Gibco) and remaining cells resuspended in Roswell Park Memorial Institute (RPMI) 1640 medium (Gibco). WBCs were then incubated with CXCL3 (150 ng ml–1, R&D Systems) or left untreated for 45 min at 37 °C. Cells were then washed twice in PBS 1× and labeled for 20 min at 4 °C for flow cytometry analyses.
ROS measurement
Whole blood EDTA from young and aged mice 24 h poststroke underwent RBCs lysis in ACK (Gibco) and was resuspended in RPMI 1640 medium (Gibco) containing 5 µM of CellROX (Thermo Fisher) at 37 °C for 30 min. Cells were then washed twice in PBS and labeled for 20 min at 4 °C for flow cytometry analyses.
NETs measurement by flow cytometry
Whole blood in EDTA from young or aged mice 24 h poststroke, underwent flow cytometry surface staining and RBCs lysis as described above. Blood was then fixed in 2% paraformaldehyde (PFA), blocked with 2% BSA (Sigma-Aldrich), stained with anti-mouse citrullinated histone H3 (Abcam) and FITC anti-MPO (Abcam) and subsequently analyzed by flow cytometry as previously described60,61.
Neutrophil-platelets aggregates measurement by flow cytometry
Whole blood EDTA from young or aged mice at 24 h poststroke was processed as described62. Briefly, blood samples were diluted 1:2 in Hepes-Tyrode Buffer and simultaneously stained with FACS antibodies for 30 min at room temperature. Samples were then fixed in FACS Lysing solution (BD), washed and analyzed by flow cytometry.
In vitro phagocytosis assay
Phagocytosis assay was performed as described22. Briefly, DH5a Escherichia coli expressing green fluorescent protein (GFP)63 were grown overnight in LB medium at 37 °C and then diluted and grown for 1–2 h to an optical density at 600 nm of ~0.5. Sorted BM neutrophils subsets (1 × 105 for each subset) were incubated with E. coli-GFP at a ratio of 1:200. Cells were then fixed in 2% PFA and analyzed by flow cytometry.
Preparation and analysis of human blood for flow cytometry
A group of 82 patients with acute ischemic stroke was analyzed. Patients were stratified by age into ‘Young’ (n = 34, median age 56.1 ± 14.9 years) and ‘Old’ (n = 48, median age 87.2 ± 5.2 years). An MC group of ‘Young’ (n = 25, median age 57.0 ± 7.0 years) and ‘Old’ (n = 19, median age 85.4 ± 4.2 years) subjects without history of ischemic stroke, acute infectious, immunological or hematological diseases was included. Venous blood samples were taken from the antecubital vein of stroke patients within 24 h (D1), at day 3 (D3) and at day 7 (D7) from stroke onset with a 19–20G needle. Control patients had a single blood collection. Blood was collected between 12:00 and 16:00, to avoid biases due to circadian oscillations of neutrophil subtypes7, into vacuum EDTA Tubes (BD Vacutainer, 6 ml) and processed immediately. Complete blood cell count was assessed with an automated hemocytometer. For flow cytometry, whole blood samples were labeled for 30 min at room temperature, subsequently fixed and RBCs were lysed in BD FACS lysing solution (BD Biosciences) overnight. The samples were then washed and resuspended in PBS. Cell data were then acquired on a FACS Canto II (BD Biosciences) and analyzed with FlowJo software (Tree Star). Forward and side scatter were used to identify the granulocyte population and to gate out debris. A tota–––l of 200,000 CD45+ events per sample were analyzed. CD62Llo neutrophils were identified as CD45+CD16+CD15+CD62LloCXCR4+ cells.
Preparation of human blood for mass cytometry
Leukocytes from a group of 32 patients with acute ischemic stroke were isolated, labeled and analyzed. Patients were stratified according to the age into ‘Young’ (n = 9, median age 60.6 ± 12.7) and ‘Old’ (n = 23, median age 86.2 ± 3.7). An MC group of ‘Young’ (n = 12, median age 64.7 ± 5.8) and ‘Old’ (n = 5, median age 82.4 ± 2.3) subjects without history of ischemic stroke, acute infectious, immunological or hematological diseases were included. Immediately after sampling, venous blood was RBCs-lysed and leukocytes were fixed in BD Phosflow Lyse/Fix Buffer diluted 1:5 in distilled water (BD Biosciences). WBCs were thereafter collected by centrifugation and washed twice in HBSS without Ca2+ and Mg2+ (Euroclone). Leukocytes were then resuspended in PBS/BSA 0.5% containing 10% DMSO and stored at –80 °C until labeling for CyTOF.
Mass-tag cellular barcoding
Mass-Tag cellular barcoding was performed as described64. Briefly, the first barcoding scheme was composed of unique combinations of three out of five palladium metals (104Pd, 105Pd, 106Pd, 108Pd, 110Pd) (Trace Sciences International)65. The second included a unique CD45 antibody preconjugated to 89Y, 194Pt, 198Pt (Fluidigm), 113In, 115In (Trace Sciences International)66. Merging of these two strategies allowed us to pool 50 samples. Palladium isotopes were conjugated to Bromoacetamidobenzyl-EDTA (Dojindo Laboratories). The concentrations were adjusted to 100 nM for all metals. After thawing samples, cells were washed once with Cell Staining Medium (CSM: PBS, 0.5% BSA (Sigma-Aldrich) and 0.02%NaN3), once with PBS and once with 0.03% Saponin (Sigma-Aldrich) in PBS and centrifuged at 600g for 5 min at 4 °C. Next, 1:1,000 diluted barcoding reagent in PBS with 0.03% saponin was mixed with the sample and incubated at room temperature for 15 min. Cells were then washed three times in CSM, and ten samples were pooled for CD45 antibody labeling. 194Pt, 198Pt, 113In, 115In metals were conjugated with purified anti-human CD45 antibody (Biolegend) in house using the Maxpar ×8 chelating polymer kit (Fluidigm) according to the manufacturer’s instructions. Ten pooled samples were incubated on ice in PBS and a unique combination of metal-tagged CD45 antibodies. Samples were then washed twice in PBS, and 50 samples were pooled for antibody surface labeling.
Metal-isotope-tagged antibodies
All anti-human antibodies, corresponding clones and tagged metal isotopes for mass cytometry analysis are listed in the Reporting Summary. Metal isotope preconjugated antibodies were purchased from Fluidigm or commercial suppliers in purified form and conjugated in house using the Maxpar ×8 chelating polymer kit (Fluidigm) according to the manufacturer’s instructions.
Cell surface/intracellular labeling for mass cytometry
To avoid nonspecific binding of antibodies, the samples were incubated at 4 °C for 30 min in Human TruStain FcX (Fc Receptor Blocking Solution; Biolegend) and Heparin 100 U ml−1 (ref. 67). Without washing, the cells were spun down, resuspended in the antibody mixture in PBS, and incubated at 4 °C for 30 min. Cells were permeabilized using Foxp3/Transcription Factor Staining Buffer Set (eBioscience) according to the manufacturer’s instructions for 45 min at 4 °C. Subsequently, the sample was washed once in Perm/Wash buffer (eBioscience) and incubated in the antibody mixture in Perm/Wash buffer (including intracellular antigens, Ki67 and Basophils) for 30 min at 4 °C. The sample was washed once in Perm/Wash buffer (eBioscience) and centrifuged to pellet the cells. Thereafter, the cells were incubated in 4% PFA overnight. Before acquisition the cells were pelleted and resuspended in 1:3,000 Cell-ID Intercalator-Ir (Fluidigm) + Maxpar Fix and Perm Buffer (Fluidigm) for 1.5–3 h. The sample was washed twice in PBS and twice in ddH2O, diluted to 1.5–106 cells ml–1 in ddH2O containing 10% EQ Four Element Calibration Beads (Fluidigm). Samples were analyzed with a Helios CyTOF2 (Fluidigm).
Preprocessing of cytometry data
Raw mass cytometry data were normalized using the MATLAB version of the Normalizer tool68 Cells were assigned by manually gating on Event length and DNA (191Ir and 193Ir) channels using FlowJo Software (Tree Star). Doublets were excluded using Gaussian discrimination channels (Center, Offset, Width, Residual). Next, samples were distinguished based on unique CD45 expression and debarcoded using Boolean gating in FlowJo software (Tree Star). The normalized data containing living cells were exported manually from FlowJo Software (Tree Star) and imported into R studio of R using the R packages ‘flowCore’ and ‘flowWorkspaceData’ (R Foundation for Statistical Computing)69,70. Before automated high-dimensional data analysis, the mass cytometry data were transformed with a cofactor in the range of 5 and 60 using an inverse hyperbolic sine (arcsinh) function71. All cytometry data were normalized between 0 and 1 to the 99–999th percentile of the merged sample in each batch. To control for the batch effect and verify the marker expression distributions, we used the same clinical sample.
Human neutrophil-sustained thrombin generation
Citrated normal pooled plasma (NPP, 700 ml) obtained from healthy donors was processed, frozen and stored as previously described72. FV- and FVIII-deficient plasma was obtained by collecting blood from 20 apparently healthy volunteers in saline containing 1% EDTA (9:1). Platelet-free plasma were obtained after two centrifugations at 2,000g for 10 min and 8,500g for 5 min at room temperature, pooled and processed as described73. Neutrophils, starting from the peripheral blood of SP and healthy donors, were isolated with the MACSxpress Whole Blood Neutrophil Isolation Kit (Miltenyi) according to the manufacturer’s instructions, diluted to 30 × 106 neutrophils ml–1 in Hepes-Tyrode buffer added with 1 mM CaCl2 and MgCl2, stained and examined by cytofluorimetry (Symphony, BD).
Calibrated automated thrombin generation74 was measured in FV- and FVIII-deficient plasma samples (40 µl) containing the neutrophil preparations (40 µl) stimulated with 0.5 µM of N-formyl-l-methionyl-l-leucyl-l-phenylalanine (fMLP, Merck), dispended in U-bottom 96 wells microtiter plates (Greiner bio-one). Plates were warmed at 37 °C in a Fluoroskan Ascent plate reader (Thermo Fisher Scientific) for 10 min, followed by the addition of 20 µl of prewarmed fluorogenic substrate (Z–Gly–Gly–Arg–AMC) per well. Fluorescence development with excitation and emission spectra of 390 and 460 nm respectively, was triggered by addition of either (1) PPP reagent (20 µl, 5pM approximate tissue factor final concentration); (2) Dade Actin FSL, containing purified soy phosphatides and rabbit brain phosphatides in 10–4 M ellagic acid (20 µl, Siemens Healthcare) or (3) MP reagent (20 µl, containing synthetic phospholipid vesicles with no tissue factor) and further tested in presence or absence of rabbit thrombomodulin (Sekisui Diagnostics) diluted in PBS (3 nM final concentration). Hemker Thrombinoscope Software (v.3.0.0.29, Thrombinoscope BV) was used to obtain the curve of thrombin concentrations as a function of time and to calculate the following parameters: Lagtime (time (min) between the addition of the triggers and the generation of 10 nM thrombin), Peak (nM, maximum velocity of thrombin generation); time to peak (ttPeak (min) needed to reach the peak); ETP (nM × time, the area under the curve, defined as endogenous thrombin potential). Each sample was tested in duplicate, and each thrombin generation measurement was calibrated against the fluorescence curve of a fixed amount of thrombin-α2-macroglobulin complex (thrombin calibrator, 20 µl) to correct for inner filter effects and substrate consumption. In parallel experiments thrombin generation was tested in NPP and in FV- and FVIII-deficient plasma with the addition of NPP diluted in PBS (final FV and FVIII concentration 1% and 0.5% of NPP).
Single-cell collection, library construction and sequencing
Myeloid cells (CD11b+ Ly6G–) and neutrophils (CD11b+ Ly6G+) from blood (named collectively CD11b+CD45+ blood cells), or Gr1+ and c-kit+-enriched cell fractions from BM, were isolated using a FACS Aria Fusion (BD Bioscience), as detailed in the gating strategy flow-chart (Fig. 3 and Supplementary Fig. 1) and collected into 0.04% BSA/PBS. The MCAO was performed the same day in young and aged mice. Sacrifice, FACS sorting and single-cell collection were performed at the same time of day, to avoid any possible effect deriving from circadian oscillation of leukocytes populations. Pools of cells from four mice were created for each group. scRNA-seq libraries were generated using a microfluidics-based approach on a Chromium Single-Cell Controller (10x Genomics) using the Chromium Single Cell 3′ Reagent Kit v.2 (PN-120235) and v.3.1 (PN-1000269), according to the manufacturer’s instructions. The cell preparation time before loading onto the Chromium 10x controller was ≤2 h. Briefly, sorted cells were counted and the viability was assessed to be ≥85% with trypan blue by microscopy. For blood, single cells were combined in ratio of 1:3 (myeloid cells: neutrophils). Sorted BM cells were combined in ratio 2:3 (Gr1+: c-kit+).
The cells were then partitioned into gel beads in emulsion (GEMs) and lysed, followed by RNA barcoding, reverse transcription and PCR amplification (14 cycles). The concentration of the scRNA-seq libraries was determined using Qubit v.3.0 and size distribution was assessed with 4200 TapeStation system (Agilent). Libraries were sequenced on an Illumina NextSeq500 and an Illumina NovaSeq6000 instruments.
scRNA-seq data processing
Cell Ranger v.2.1.1 (for stroke blood samples, analyzed in Fig. 3 and Supplementary Fig. 1) and unique molecular identifier (UMI) (UMItool v.1.0.0 (for blood and BM samples, at steady state and after stroke, analyzed in Extended Data Figs. 6 and 7) were used to align the raw sequencing reads to the mm10 mouse genome and to quantify the expression level of genes in each cell (basic annotations from Gencode version M22). Gene counts were imported into the R environment (R v.3.6.0, R v.4.0.3) and analyzed with the Seurat package (http://satijalab.org/seurat/). For the experiment in Fig. 3, cells from the two stroke blood samples were pooled together using the Merge function in Seurat and analyzed with the standard Seurat analysis workflow (Seurat v.2.3.1). Then, neutrophils were identified by considering the expression of Ly6G, Csf3r and Ngp genes and a new complete Seurat analysis was performed on this selected population (Seurat v.3.2.1). Cells expressing <200 or >3,500 unique genes and cells with a ratio of mitochondrial versus endogenous genes expression >0.1, were discarded. A total of 5,793 cells was retained in the analysis, associated with 11,691 comprehensively detected genes. Counts were log-normalized using the NormalizeData function, with a scale factor of 10,000, to remove the differences in sequencing depth across cells. Data were then scaled using the ScaleData function, regressing on number of UMI and percentage of expressed mitochondrial genes. On the scaled data, cell cycle scores were estimated through the CellCycleScoring function and differences between S and G2M scores were regressed out as well. The most variably expressed genes within each dataset were identified through the FindVariableFeatures function (y.cutoff = 0.5 for the analysis on total cells and top 1,000 most variable genes according to Variance Stabilize Transformation for neutrophils) and used for the principal component analysis (PCA), implemented with the RunPCA function. Next, we selected principal components 1–15 (for total cells) or 1–20 (for neutrophil population) as input for the subsequent UMAP algorithm, performed with the RunUMAP function. Unsupervised clustering was obtained using the original Louvain algorithm (resolution = 0.2 for total cells and 0.4 for neutrophils) based on the same principal components as for the RunUMAP function. For the integrated analysis, shown in Supplementary Fig. 1, the same filtered young and aged stroke blood samples were analyzed together using the Seurat (v.4.1.0) anchoring-integration standard workflow20, and then subsetted to consider the neutrophil population only.
For the experiment described in Extended Data Figs. 6 and 7, cells from four datasets (at steady state and 24 h poststroke for blood and BM) including young and aged condition for each dataset, were pooled and analyzed together. Cells expressing <200 or >3,500 unique genes, and cells with a ratio of mitochondrial versus endogenous genes expression >0.1, were excluded from the blood stroke samples. Cells expressing <200 or >4,000 unique genes, and with a ratio of mitochondrial versus endogenous genes expression >0.05, were excluded from blood steady-state samples. Cells expressing < 200 or >7,000 unique genes, or with a ratio of mitochondrial versus endogenous genes >0.1, were excluded from the stroke BM samples. Cells expressing <200 or >6,000 unique genes, or with a ratio of mitochondrial versus endogenous genes >0.05, were excluded from the steady state BM samples. A total of 33,664 cells was retained in the analysis, associated with 25,017 detected genes and was log-normalized within each cell (the LogNormalize method normalizes the gene expression for each cell by the total expression, multiplies this by a scale factor of 10,000, and log-transforms the results). The top 2,000 variable genes calculated by Seurat (selection method vst, 2,000 features per dataset) were used to harmonize and integrate sample datasets by sample of origin (FindIntegrationAnchors and IntegrateData functions). The dataset was then scaled to remove unwanted sources of variation such as batch effects. On scaled data the PCA, was performed. Next, we selected principal components 1–40 (for total cells) or 1–30 (for neutrophils) as input to perform the RunUMAP function to obtain bidimensional coordinates for each cell.
Unsupervised clustering was performed using the FindClusters function (resolution: 0.9 for both total cells and neutrophils) to cluster cells using the Louvain algorithm based on the same principal components as for the RunUMAP function.
For the experiment described in Supplementary Fig. 4, the filtered and high-quality cells from the steady-state BM dataset were pooled and analyzed together with cells of steady-state BM from the Tabula Muris Senis publicly available dataset31. Only data generated with 10x Genomics-based scRNA-seq technology and from 1-month and 18-months-old male mice were considered. A total of 21,340 cells was retained, associated with 24,950 comprehensively detected genes and was log-normalized within each cell. The top 2,000 variable genes calculated by Seurat (selection method vst, 2,000 features per dataset) were used to harmonize and integrate sample datasets (FindIntegrationAnchors and IntegrateData functions). The dataset was then scaled to remove unwanted sources of variation such as batch effects. On scaled data, linear dimension reduction, the PCA, was performed. Next, we selected principal components 1–60 (for both total cells and neutrophil lineage cells) as input to perform the RunUMAP function to obtain bidimensional coordinates for each cell. Unsupervised clustering was performed using the FindClusters function (resolution: 1.1 for total cells and 0.65 for neutrophils) to cluster cells using the Louvain algorithm, as described above.
The assignment of cell identity to clusters was based on the comparison of top50 DEG calculated for each cluster against the others (Identification of DEGs) and by signature-based comparison (Scoring of biological processes) with a previously published and reported mouse neutrophil scRNA-seq studies17.
Trajectory analysis was performed on each dataset using the Monocle3 (v.3.0.2.2) Bioconductor package to infer the cells-associated pseudotime, taking advantage of the SeuratWrappers R package to load the original Seurat object in the Monocle3 environment.
Identification of DEGs
The FindAllMarkers function (logfc.threshold = 0.25, min.pct = 0.25, test.use = ‘Bimod’ for total cells and ‘Wilcox’ for neutrophils) to the normalized data to identify significantly more highly expressed genes within clusters was applied. For the integrated datasets, the data used in the test were retrieved from the ‘RNA’ slot of the Seurat objects. P value adjustment was performed using Bonferroni correction based on the total number of tested genes. Only genes with logFC≥0.25 and expressed by at least 25% of the cells analyzed were tested. An analogous procedure was performed to compare cells from aged and from young mice, within each selected cluster.
Scoring of biological processes
Individual neutrophils were scored for their expression of gene signatures representing certain biological functions as previously described17. For all signatures except neutrophil aging, functional scores were defined using the AddModuleScore function. Aging score was defined as the weighted average of normalized expression of age-related genes. Gene weights were set to either 1 or –1 to reflect positive or negative relationships with the aging process. To correlate different cluster signature across different dataset we used the GSVA algorithm75 as implemented in the GSVA Bioconductor R package. The neutrophil maturation, stem and cell cycle signatures were derived by using genes identified by Xie et al.17. G-csf signature was derived from Giladi et al.33 and included only genes with logFC ≥1 or ≤–1. Granule signatures were derived from Xie et al.17 and Cowland and Borregaard76. NETs-associated gene signature was derived from Wang et al.77. Other functional signatures were derived from the Gene Ontology (GO) database78, with the full gene list provided in Supplementary Table 3 and Supplementary Table 5. Score differences between clusters were evaluated using Wilcoxon rank sum test. For comparing the functional state of cells in the cluster scNeu1, Pearson’s correlation was calculated between each pair of functional signatures, considering the scores obtained by the cells in cluster scNeu1.
To compare the similarity of the clusters of the Tabula Muris Senis to the clusters of the steady-state BM, we calculated a correlation score for each considered cluster, considering the top 20 marker genes defined according to the top logFC. The average expression of each gene identified was calculated within the four groups, Tabula Muris Senis young (TM-Y), BM steady-state young (BM-Y), Tabula Muris Senis aged (TM-A), BM steady-state aged (BM-A), in the considered cluster. Pearson’s correlation between TM-Y and BM-Y and between TM-A and BM-A was evaluated by comparing the average expression levels of these 20 genes.
SCENIC analysis
Transcription factor analysis was conducted following the standard workflow of the SCENIC79 package (https://doi.org/10.1038/nmeth.4463) in its R implementation (v.1.1.2), starting from the normalized counts of blood stroke samples and using GENIE3 to retrieve coexpression networks.
Bulk RNA isolation, sequencing and analysis
The CD62LloCXCR2lo neutrophil enriched fraction was isolated with FACS Aria Fusion (BD Bioscience) and collected into 0.5% BSA/PBS, from blood of young and aged mice in steady state and 24 h poststroke. Pools of neutrophils from three mice were created for each group. RNA was isolated using ReliaPrep RNA Cell Miniprep System (Promega), according to the manufacturer’s instructions. Purified RNAs were quantified using the Agilent Bioanalyzer, RNA used had an RNA integrity number ≥ 7.
Libraries were prepared using SMART-Seq v.4 Ultra Low Input RNA Kit for Sequencing (Takara Bio) performing 17 cycles of amplification, according to the manufacturer’s instructions. Sequencing was performed on a NextSeq500 machine (Illumina) obtaining 30 million single-end reads per sample on average. Reads were trimmed using Trimmomatic, v.0.32 to remove adapters and to exclude low-quality reads from the analysis. The remaining reads were then aligned to the reference genome mm10, Gencode v. M16, using STAR aligner, v.2.5.3a. FeatureCounts function from Rsubread package (v.1.16)80 was used to assign reads to the corresponding genes. Only genes with a count per million (CPM) value >1 in at least four samples were retained. Gene expression read counts were exported and analyzed in the R environment (v.3.1.1) to identify differentially expressed genes (DEGs), using the DESeq2 Bioconductor library81,82. DEGs for each comparison were defined setting a cutoff of FDR ≤ 0.05.
Preranked GSEA83 was performed for each comparison, on all the expressed genes. The gene sets included in the GSEA analyses were obtained from C2: Curated Gene Sets, Canonical Pathways subset, C5: Ontology Gene Sets, Biological Process and Molecular Function subsets collections and C3: Regulatory Target Gene Set, as are reported in the MSigDB database.
Tissue histological preparation and analysis
On the day of sacrifice, animals were weighed, deeply anesthetized and perfused transcardially with 25 ml of PBS (pH 7.4 with EDTA), followed by 25 ml of PFA4% in PBS. Brains were carefully dissected, postfixed overnight, cryoprotected in sucrose and embedded in tissue-freezing medium for cryostat sectioning. Frozen brains were cut into 20 µm-thick coronal cryostat sections.
Analysis of infarct size
For the measurement of the ischemic lesion volume, 20-μm-thick coronal cryostat sections were sliced from 2 mm rostral to 4 mm caudal to the Bregma. One systematic random series of sections per mouse was stained with cresyl violet (Sigma), digitalized and analyzed with Image J (National Institutes of Health). The lesion area was measured for each reference level by the ‘indirect method,’ as previously described84.
For the representative three-dimensional (3D) volume rendering image, a representative brain per group was traced using the assistance of the Stereo Investigator v.3.0 software (MicroBrightField, Inc.)84,85. Images were first acquired with a CCD-IRIS color video camera and the cerebral hemispheres, lesioned area and ventricles were interactively delineated at low magnification on a video image of the section.
DHE staining
For DHE (Thermo Fisher, catalog no. D1168) staining, mice were intraperitoneally injected with 30 mg kg–1 of DHE 3 h before sacrifice and then processed as described above.
Immunofluorescence
Immunofluorescence labeling was performed as previously described85. Briefly, coronal brain sections (20 µm thickness) were incubated with blocking solution (10% secondary antibody species serum and Triton 0.05-0.1% in PBS), for 1 h. Primary antibodies were diluted in blocking solution (1% secondary antibody species serum and Triton 0.05–0.1% in PBS, Sigma) and sections were incubated overnight at 4 °C with proper primary antibodies (detailed information is provided in the Reporting Summary). Adequate fluorophore-conjugated secondary antibodies (Thermo Fisher Scientific) were used. Omission of the primary antibodies did not reveal any nonspecific labeling. Confocal (Leica SP5 and SP8 equipped with ×40 and ×63 objectives; Leica, Germany) images were obtained for tissue analysis. Blood vessel were counterstained with Lycopersicon esculentum (tomato) lectin Dylight594 and 649 in PBS (Vector Laboratories).
NeuN+ neuronal cell quantification
Immunostained NeuN+ cells were counted in the perilesional areas of the cortex and striatum. Ischemic lesions were visible on microscopy because of signs of overt brain tissue damage and cell loss; the borderline area was considered as perilesional area.
Detection of entrapped RBCs
At 6 h after reperfusion, mice were sacrificed and processed as described above. Extracted brains were postfixed in 4% PFA for 24 h and then embedded in paraffin; 6-µm-thick microtome sections were cut. Paraffin-embedded slices were deparaffinized in xylene and rehydrated in decreasing ethanol concentration solutions. Slides were incubated 30 min in sodium borohydride (NaBH4, Merck, catalog no. 13462) 0.2% in PBS at room temperature, washed twice in PBS. Vessels were counterstained with tomato lectin Dylight649 as described above15.
Detection of microvascular patency
At 6 h after reperfusion, mice were injected intravenously with 200 mg kg–1 of HRP IV (Sigma, catalog no. P8375) in PBS 20 min before sacrifice and brains collected after sacrifice without any previous perfusion, as described15. Brains were processed as described above; 60-µm-thick cryostat sections were cut in free-floating. Image analysis was performed with a brightfield Leica microscope connected to Neurolucida v.3.0 software (MicroBrightField Inc.). The number of microvascular constrictions was evaluated.
Peripheral blood smear preparation and analysis
Mouse peripheral blood smears were prepared using the coverslip method, as described86. Briefly, a drop of blood was placed at base of the slide, a second ‘spreader’ slide was placed at the base of the first to touch the drop of blood that was then smeared with the spreader87. Blood smears were then stained with May-Grunwald/Giemsa as described86 and analyzed in brightfield microscopy with ×40 magnification; 100 WBCs per slide were counted, about 1 mm away from the tail88.
Luminex assay
Mouse Luminex Screening Assay (R&D) was performed to measure G-CSF, CXCL1, M-CSF, GM-CSF, IL-6, CXCL10 and IL-17A in platelet-free plasma collected in basal conditions (0 h) and 48 h poststroke, according to the manufacturer’s instructions.
ELISA for CXCL3
ELISA assay was performed to measure CXCL3 in human (Abcam) and mouse (Abcam) platelet-free plasma collected at day 3 after stroke (D3), or in steady state (0 h) and 48 h poststroke, according to the manufacturer’s instructions.
Quantification and statistical analysis
Investigator blinding to experimental conditions was applied and all statistical analyses were conducted using GraphPad Prism software (v.8).
A two-tailed Student’s t-test, Mann–Whitney U-test, one-way analysis of variance (ANOVA), two-way ANOVA, repeated-measures two-way ANOVA or mixed-effect model (restricted maximum likelihood (REML)) when values were missing, and Chi-square test for categorical variables, as appropriate, were used for statistical analyses. Wilcoxon rank sum test was applied to calculate statistical differences among scRNA-seq scores. For human datasets, Mann–Whitney U-test was applied. ROC curves were used to assess the predictive value of leukocyte subtype counts for long-term functional outcome. Stepwise forward conditional logistic regression was employed for multivariate analysis.
For ANOVAs, Sidak or Original FDR method of Benjamini and Hochberg post hoc comparison were used. There are no missing data, animals were excluded as stated above if not exhibiting correct middle cerebral artery occlusion. Values are represented as mean ± s.e.m., with differences considered significant when P ≤ 0.05.
Automated population identification
First, we carried out FlowSOM clustering to generate a starting point of 100 nodes using pr-processed and combined mass and flow cytometry datasets42,43. This was then followed by expert-guided manual metaclustering, using parameters listed in each figure legend. The respective k-value was chosen manually; identified clusters were annotated and merged based on a similarity of antigen expression to uphold the biological relevance of the dataset. Manually annotated clusters were used to calculate the relative frequencies of immune populations. Heatmaps display median expression levels of all markers per merged population and plotted using the R package ‘pheatmap’89. For human samples, we preselected CD66b expressing cells and performed additional FlowSOM analysis to identify neutrophil subsets; to define CD62L low/high clusters we set median expression level at 0.03. To define CD101 low/high clusters we set median expression level at 0.4.
For a complex overview of the immune/neutrophil compartment, we used UMAP90. To create a UMAP, we pooled an equally proportioned number of cells per group. To visualize a developmental trajectory of neutrophils we applied PHATE algorithm91.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data that support the findings of this study are available from the corresponding author upon request. All sequencing data generated in this study have been deposited at NCBI’s Gene Expression Omnibus (GEO) repository and are accessible through GEO Series accession number GSE174440. Source data are provided with this paper. Further information and requests for resources and reagents should be directed to and will be fulfilled by the Lead Contact, M. Bacigaluppi (bacigaluppi.marco@hsr.it). This study did not generate new or unique reagents.
Code availability
Code that supports the findings of this study are available from the corresponding author upon request. All sequencing code generated in this study have been deposited at NCBI’s GEO repository and are accessible through GEO Series accession number GSE174440.
References
Benjamin, E. J. et al. Heart disease and stroke statistics—2019 update: a report from the American Heart Association. Circulation 139, e56–e528 (2019).
Feigin, V. L., Lawes, C. M., Bennett, D. A. & Anderson, C. S. Stroke epidemiology: a review of population-based studies of incidence, prevalence, and case-fatality in the late 20th century. Lancet Neurol. 2, 43–53 (2003).
Iadecola, C., Buckwalter, M. S. & Anrather, J. Immune responses to stroke: mechanisms, modulation, and therapeutic potential. J. Clin. Investig. 130, 2777–2788 (2020).
Gelderblom, M. et al. Temporal and spatial dynamics of cerebral immune cell accumulation in stroke. Stroke 40, 1849–1857 (2009).
Courties, G. et al. Ischemic stroke activates hematopoietic bone marrow stem cells. Circ. Res. 116, 407–417 (2015).
Semerano, A. et al. Leukocyte counts and ratios are predictive of stroke outcome and hemorrhagic complications independently of infections. Front. Neurol. 11, 201 (2020).
Adrover, J. M. et al. A neutrophil timer coordinates immune defense and vascular protection. Immunity 50, 390–402 e310 (2019).
Jordan, J. E., Zhao, Z. Q. & Vinten-Johansen, J. The role of neutrophils in myocardial ischemia-reperfusion injury. Cardiovasc. Res. 43, 860–878 (1999).
El Amki, M. et al. Neutrophils Obstructing brain capillaries are a major cause of no-reflow in ischemic stroke. Cell Rep. 33, 108260 (2020).
Rossi, D. J. et al. Cell intrinsic alterations underlie hematopoietic stem cell aging. Proc. Natl Acad. Sci. USA 102, 9194–9199 (2005).
Maryanovich, M. et al. Adrenergic nerve degeneration in bone marrow drives aging of the hematopoietic stem cell niche. Nat. Med. 24, 782–791 (2018).
Akunuru, S. & Geiger, H. Aging, clonality, and rejuvenation of hematopoietic stem cells. Trends Mol. Med. 22, 701–712 (2016).
Jaiswal, S. et al. Age-related clonal hematopoiesis associated with adverse outcomes. N. Engl. J. Med. 371, 2488–2498 (2014).
Jaiswal, S. et al. Clonal hematopoiesis and risk of atherosclerotic cardiovascular disease. N. Engl. J. Med. 377, 111–121 (2017).
Yemisci, M. et al. Pericyte contraction induced by oxidative-nitrative stress impairs capillary reflow despite successful opening of an occluded cerebral artery. Nat. Med. 15, 1031–1037 (2009).
Nortley, R. et al. Amyloid beta oligomers constrict human capillaries in Alzheimer’s disease via signaling to pericytes. Science 365, eaav9518 (2019).
Xie, X. et al. Single-cell transcriptome profiling reveals neutrophil heterogeneity in homeostasis and infection. Nat. Immunol. 21, 1119–1133 (2020).
Vafadarnejad, E. et al. Dynamics of cardiac neutrophil diversity in murine myocardial infarction. Circ. Res. 127, e232–e249 (2020).
Zilionis, R. et al. Single-cell transcriptomics of human and mouse lung cancers reveals conserved myeloid populations across individuals and species. Immunity 50, 1317–1334 e1310 (2019).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 e1821 (2019).
Casanova-Acebes, M. et al. Rhythmic modulation of the hematopoietic niche through neutrophil clearance. Cell 153, 1025–1035 (2013).
Evrard, M. et al. Developmental analysis of bone marrow neutrophils reveals populations specialized in expansion, trafficking, and effector functions. Immunity 48, 364–379 e368 (2018).
Grassi, L. et al. Dynamics of transcription regulation in human bone marrow myeloid differentiation to mature blood neutrophils. Cell Rep. 24, 2784–2794 (2018).
Kwok, I. et al. Combinatorial single-cell analyses of granulocyte-monocyte progenitor heterogeneity reveals an early uni-potent neutrophil progenitor. Immunity 53, 303–318 e305 (2020).
Jonsson, H., Allen, P. & Peng, S. L. Inflammatory arthritis requires Foxo3a to prevent Fas ligand-induced neutrophil apoptosis. Nat. Med. 11, 666–671 (2005).
Yan, K. et al. Haploinsufficiency of PSMD12 causes proteasome dysfunction and subclinical autoinflammation. Arthritis Rheumatol. 74, 1083–1090 (2022).
Fischer, J. et al. Safeguard function of PU.1 shapes the inflammatory epigenome of neutrophils. Nat. Immunol. 20, 546–558 (2019).
Bjerregaard, M. D., Jurlander, J., Klausen, P., Borregaard, N. & Cowland, J. B. The in vivo profile of transcription factors during neutrophil differentiation in human bone marrow. Blood 101, 4322–4332 (2003).
Khoyratty, T. E. et al. Distinct transcription factor networks control neutrophil-driven inflammation. Nat. Immunol. 22, 1093–1106 (2021).
Zhang, D. E. et al. Absence of granulocyte colony-stimulating factor signaling and neutrophil development in CCAAT enhancer binding protein alpha-deficient mice. Proc. Natl Acad. Sci. USA 94, 569–574 (1997).
Tabula Muris, C. A single-cell transcriptomic atlas characterizes ageing tissues in the mouse. Nature 583, 590–595 (2020).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Giladi, A. et al. Single-cell characterization of haematopoietic progenitors and their trajectories in homeostasis and perturbed haematopoiesis. Nat. Cell Biol. 20, 836–846 (2018).
Kallquist, L. et al. The tetraspanin CD63 is involved in granule targeting of neutrophil elastase. Blood 112, 3444–3454 (2008).
Flohr Svendsen, A. et al. A comprehensive transcriptome signature of murine hematopoietic stem cell aging. Blood 138, 439–451 (2021).
Pillay, J., Tak, T., Kamp, V. M. & Koenderman, L. Immune suppression by neutrophils and granulocytic myeloid-derived suppressor cells: similarities and differences. Cell Mol. Life Sci. 70, 3813–3827 (2013).
Frenette, P. S., Mayadas, T. N., Rayburn, H., Hynes, R. O. & Wagner, D. D. Susceptibility to infection and altered hematopoiesis in mice deficient in both P- and E-selectins. Cell 84, 563–574 (1996).
Tamassia, N. et al. Induction of OCT2 contributes to regulate the gene expression program in human neutrophils activated via TLR8. Cell Rep. 35, 109143 (2021).
Khanna-Gupta, A. et al. Growth factor independence-1 (Gfi-1) plays a role in mediating specific granule deficiency (SGD) in a patient lacking a gene-inactivating mutation in the C/EBPepsilon gene. Blood 109, 4181–4190 (2007).
Shojaei, F. et al. Bv8 regulates myeloid-cell-dependent tumour angiogenesis. Nature 450, 825–831 (2007).
Becht, E. et al. Dimensionality reduction for visualizing single-cell data using UMAP. Nat. Biotechnol. 37, 38–44 (2018).
Van Gassen, S. et al. FlowSOM: using self-organizing maps for visualization and interpretation of cytometry data. Cytometry A 87, 636–645 (2015).
Hartmann, F. J. et al. High-dimensional single-cell analysis reveals the immune signature of narcolepsy. J. Exp. Med. 213, 2621–2633 (2016).
Niccoli, G., Burzotta, F., Galiuto, L. & Crea, F. Myocardial no-reflow in humans. J. Am. Coll. Cardiol. 54, 281–292 (2009).
Cruz Hernandez, J. C. et al. Neutrophil adhesion in brain capillaries reduces cortical blood flow and impairs memory function in Alzheimer’s disease mouse models. Nat. Neurosci. 22, 413–420 (2019).
Ng, L. G., Ostuni, R. & Hidalgo, A. Heterogeneity of neutrophils. Nat. Rev. Immunol. 19, 255–265 (2019).
Montaldo, E. et al. Cellular and transcriptional dynamics of human neutrophils at steady state and upon stress. Nat. Immunol. 23, 1470–1483 (2022).
Zhang, D. et al. Neutrophil ageing is regulated by the microbiome. Nature 525, 528–532 (2015).
Dong, Y. et al. Neutrophil hyperactivation correlates with Alzheimer’s disease progression. Ann. Neurol. 83, 387–405 (2018).
Salib, C. & Teruya-Feldstein, J. Hypersegmented granulocytes and COVID-19 infection. Blood 135, 2196 (2020).
Schulte-Schrepping, J. et al. Severe COVID-19 Is marked by a dysregulated myeloid cell compartment. Cell 182, 1419–1440 e1423 (2020).
Silvin, A. et al. Elevated calprotectin and abnormal myeloid cell subsets discriminate severe from mild COVID-19. Cell 182, 1401–1418 e1418 (2020).
Middleton, E. A. et al. Neutrophil extracellular traps contribute to immunothrombosis in COVID-19 acute respiratory distress syndrome. Blood 136, 1169–1179 (2020).
Sokulsky, L. A. et al. A critical role for the CXCL3/CXCL5/CXCR2 neutrophilic chemotactic axis in the regulation of type 2 responses in a model of rhinoviral-induced asthma exacerbation. J. Immunol. 205, 2468–2478 (2020).
Eash, K. J., Greenbaum, A. M., Gopalan, P. K. & Link, D. C. CXCR2 and CXCR4 antagonistically regulate neutrophil trafficking from murine bone marrow. J. Clin. Investig. 120, 2423–2431 (2010).
Bacigaluppi, M. et al. Delayed post-ischaemic neuroprotection following systemic neural stem cell transplantation involves multiple mechanisms. Brain 132, 2239–2251 (2009).
De Feo, D. et al. Neural precursor cell-secreted TGF-beta2 redirects inflammatory monocyte-derived cells in CNS autoimmunity. J. Clin. Investig. 127, 3937–3953 (2017).
Lelios, I. & Greter, M. Isolation of leukocytes from mouse central nervous system. Methods Mol. Biol. 1193, 15–19 (2014).
Capotondo, A. et al. Brain conditioning is instrumental for successful microglia reconstitution following hematopoietic stem cell transplantation. Proc. Natl Acad. Sci. USA 109, 15018–15023 (2012).
Gavillet, M. et al. Flow cytometric assay for direct quantification of neutrophil extracellular traps in blood samples. Am. J. Hematol. 90, 1155–1158 (2015).
Perdomo, J. et al. Neutrophil activation and NETosis are the major drivers of thrombosis in heparin-induced thrombocytopenia. Nat. Commun. 10, 1322 (2019).
Poli, V. et al. Inhibition of transcription factor NFAT activity in activated platelets enhances their aggregation and exacerbates gram-negative bacterial septicemia. Immunity 55, 224–236 e225 (2022).
Chua, R. Y. & Wong, S. H. SNX3 recruits to phagosomes and negatively regulates phagocytosis in dendritic cells. Immunology 139, 30–47 (2013).
Friebel, E. et al. Single-Cell mapping of human brain cancer reveals tumor-specific instruction of tissue-invading leukocytes. Cell 18, 1626–1642.e20 (2020).
Zunder, E. R. et al. Palladium-based mass tag cell barcoding with a doublet-filtering scheme and single-cell deconvolution algorithm. Nat. Protoc. 10, 316–333 (2015).
Mei, H. E., Leipold, M. D. & Maecker, H. T. Platinum-conjugated antibodies for application in mass cytometry. Cytometry A 89, 292–300 (2016).
Rahman, A. H., Tordesillas, L. & Berin, M. C. Heparin reduces nonspecific eosinophil staining artifacts in mass cytometry experiments. Cytometry A 89, 601–607 (2016).
Finck, R. et al. Normalization of mass cytom- etry data with bead standards. Cytometry A 83, 483–494 (2013).
Ellis, B. et al. flowCore: Basic structures for flow cytometry data. R package version 2.3.0. https://bioconductor.org/packages/flowCore/ (2019).
Finak, G. flowWorkspaceData: A data package containing two flowJo, one diva xml workspace and the associated fcs files as well as three GatingSets for testing the flowWorkspace, openCyto and CytoML packages. https://bioconductor.org/packages/release/data/experiment/html/flowWorkspaceData.html (2018).
Bendall, S. C. et al. Single-cell mass cytometry of differential immune and drug responses across a human hematopoietic continuum. Science 332, 687–696 (2011).
D’Angelo, A. et al. Effect of clot-detection methods and reagents on activated partial thromboplastin time (APTT). Implications in heparin monitoring by APTT. Am. J. Clin. Pathol. 94, 297–306 (1990).
D’Angelo, S. V., Gilardoni, F. & D’Angelo, A. Evaluation of coagulometric assays in the assessment of protein C anticoagulant activity; variable sensitivity of commercial APTT reagents to the cofactor effect of protein S. Thromb. Haemost. 62, 861–867 (1989).
Hemker, H. C. & Kremers, R. Data management in thrombin generation. Thromb. Res. 131, 3–11 (2013).
Hanzelmann, S., Castelo, R. & Guinney, J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14, 7 (2013).
Cowland, J. B. & Borregaard, N. Granulopoiesis and granules of human neutrophils. Immunol. Rev. 273, 11–28 (2016).
Wang, J. et al. Excessive neutrophils and neutrophil extracellular traps in COVID-19. Front. Immunol. 11, 2063 (2020).
The Gene Ontology, C. The Gene Ontology Resource: 20 years and still GOing strong. Nucleic Acids Res. 47, D330–D338 (2019).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Escobar, G. et al. Interferon gene therapy reprograms the leukemia microenvironment inducing protective immunity to multiple tumor antigens. Nat. Commun. 9, 2896 (2018).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Butti, E. et al. Subventricular zone neural progenitors protect striatal neurons from glutamatergic excitotoxicity. Brain 135, 3320–3335 (2012).
Bacigaluppi, M. et al. Neural stem cell transplantation induces stroke recovery by upregulating glutamate transporter GLT-1 in astrocytes. J. Neurosci. 36, 10529–10544 (2016).
Houwen, B. Blood film preparation and staining procedures. Clin. Lab. Med. 22, 1–14 (2002).
Adewoyin, A. S. & Nwogoh, B. Peripheral blood film—a review. Ann. Ib. Postgrad. Med. 12, 71–79 (2014).
Bain, B. J. Diagnosis from the blood smear. N. Engl. J. Med. 353, 498–507 (2005).
Kolde, R. pheatmap: pretty heatmaps. https://rdrr.io/cran/pheatmap/ (2019).
McInnes, L., Healy, J. & Melville, J. Umap: uniform manifold approximation and projection for dimension reduction. Preprint at https://arxiv.org/abs/1802.03426 (2018).
Moon, K. R. et al. Visualizing structure and transitions in high-dimensional biological data. Nat. Biotechnol. 37, 1482–1492 (2019).
Acknowledgements
We thank A. Hidalgo for scientific advice, counseling and critical reading of the manuscript; P. Frenette for PEdKO mice; R. Ostuni and M. Genua for advice and help on blood scRNA-seq; C. Doglioni and F. Sanvito for advice; Advanced Light and Electron Microscopy BioImaging Center (ALEMBIC); F. Simeoni, D. Lazarevic and M. Morelli, Center for Translational Genomics and BioInformatics, IRCCS San Raffaele Hospital, for support in bioinformatics analysis; P.Panina-Bordignon and C. Bordignon for critical reading of the manuscript; M. Raso, hematological and chemistry laboratory service, Mouse Clinic, IRCCS, San Raffaele Hospital. This work has been supported in part by: TargetBrain (EU Framework 7 project HEALTH-F2-2012-279017) to G.M. and to B.B.; Cariplo, (‘Ricerca Medica Giovani ricercatori’ 2015–1057) to M.B. and A.U.; Ministero della Salute Italiana (Progetto Giovani Ricercatori 58/GR-2011−02348160) to M.B.; European Union (EU) funding within the NextGenerationEU-MUR PNRR Extended Partnership initiative on Consequences and challenges of ageing, Ageing Well in an Ageing Society (Project no. PE00000015, Age-It) to G.M.; European Research Council (ERC) under the EU’s Horizon 2020 research and innovation program grant agreement no. 882424 to B.B.; Swiss National Science Foundation (733 310030_170320, 310030_188450 and CRSII5_183478 to B.B.; Swiss Multiple Sclerosis Society to B.B. and to D.D.F.); University of Zurich postdoctoral fellowship to D.D.F.
Author information
Authors and Affiliations
Contributions
M.B. and D.D.F. conceptualized the study. G.S.G., D.D.F., E.F., A.S., N.M., and M.B. developed the methodology. D.D.F., E.F. and G.M.S. wrote the software. G.S.G., D.D.F., E.F., A.S. and M.B. performed formal analyses. G.S.G., D.D.F., E.F., A.S., N.M., E.Butti., E.Brambilla, A.B., T.V., M.G., A.dʼA., P.D.V., P.R., S.C., M.P., A.C., M.G.S. and M.B. performed the investigation. G.S.G., A.S., A.G., G.M., B.B., A.U., L.R., G.C. and M.B. provided resources. G.S.G., D.D.F., E.F., G.M.S. and M.B. curated the data. D.D.F., G.S.G. and M.B. wrote the original draft of the manuscript. D.D.F., G.S.G., M.B., B.B. and G.M. contributed to writing, review and editing of the manuscript. G.S.G., D.D.F. and M.B. worked on visualization. B.B., G.M. and M.B. were responsible for funding acquisition and supervision of the work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks Steffen Massberg, Andres Hidalgo and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Ioana Visan was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Worse stroke outcome in aged male and female mice.
a, mNSS scores of young and aged male mice up to 7 days poststroke (n = 10 young, n = 6 aged, Mixed- effect Model, with Bonferroni post-hoc, one experiment, *p ≤ 0.05, **p ≤ 0.01). b, Kaplan-Meier survival curve showing mortality up to 8 days poststroke (n = 10 young and n = 8 aged male mice, Mantel-Cox test, *p = 0.0341). c- e, Stroke in young (2–4 months) and aged (18–24 –months) female mice. c, mNSS of young (n = 7) and aged (n = 4) female mice measured with the mNSS at 24h poststroke (2-tailed, Mann-Whitney test, **p = 0.003) and Kaplan-Meier survival curve up to 3days poststroke (n = 7 young, n = 5 aged female mice; Mantel-Cox test, **p = 0.0053). d, Staining for neurons (NeuN+ in brown) at 3 days poststroke and quantification of neurons (NeuN+) in the peri-lesional area (n = 7 young, n = 5 aged female mice, ***p = 0.001). e, Quantification of ischemic volume by Cresyl Violet staining at 3 days poststroke (n = 7 young n = 8 aged female mice). f-i, Representative images, and density of DHE+ (in red) astrocytes (GFAP+, green in f), neurons (NeuN+, green in g), blood vessels (CD31+, green in h) and pericytes (PDGFRb+, green in i) in the peri-lesional area at 24h poststroke in male young and aged mice (n = 3 mice per group *p = 0.0358). Cell nuclei are stained in blue (DAPI). j, Representative confocal microscopy images of 3NT+ cells (in green) in the ischemic cortex, and density of 3NT+ cells in the ischemic area of young and aged mice at 3 days poststroke (n = 8 young, n = 4 aged mice, **p = 0.0094). In the scatterplots each dot corresponds to a single mouse; mean ± SEM is shown. Scale bar in d, f, g, h, i, j, 50µm. 2-tailed t-test if not otherwise indicated.
Extended Data Fig. 2 Aged mice with stroke have increased brain and blood neutrophils.
UMAP displaying 17 resident and infiltrating immune cell subsets in the ischemic brain from young and aged mice at 24h poststroke analyzed by 19-color flow cytometry. b, Frequency of immune cell subsets identified in a (as percentage among total CD45+ cells, n = 5 mice per group, *p ≤ 0.05, **p ≤ 0.01, ****p ≤ 0.0001). c, Graph showing the number of eosinophils (SiglecF+) in the lesional and perilesional areas at 24h poststroke (n = 5 mice per group). d, UMAP of 18-color flow cytometry analysis of blood leukocytes in young and aged mice at steady state conditions (0h), at 6h and 24h poststroke (n = 5 mice per group). e, Heatmap showing the MAE values for the clustering markers for each cell cluster. f, Frequencies of blood leukocyte sub-populations among total CD45+ cells in young and aged mice (groups as in d, Mixed- effect Model, Benjamini-Hochberg post-hoc test; *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, one of two independent experiments is shown). In the scatterplots in b each dot corresponds to a single mouse; in f the boxes extend from the 25th to 75th percentiles, the midline denotes the median while whiskers plot min and max. Two tailed t-test if not otherwise indicated.
Extended Data Fig. 3 Neutrophil depletion and scRNAseq on blood leukocytes in stroke of the aged.
a, Neutrophil depletion by intravenous injection of anti-Ly6G antibody(1A8) or isotype-control (iso) in aged mice (a-f); MCFC plots for blood CD11b+CXCR2+neutrophils (gated on CD45+7/4+cells) and quantification of neutrophils 1.5h after treatment (n = 4 mice/group, **p = 0.0048). b, LDF-recordings (n = 15 Aged+1A8, n = 13 Aged+iso, 2-ANOVA, Benjamini-Hochberg post-test, **p ≤ 0.01). c, Immunofluorescence images and quantification of capillaries (t.lectin+vessels, white) with entrapped erythrocytes (NaBH4+, red) in the peri-ischemic cortex at 6h poststroke (n = 4, Aged+1A8; n = 5 Aged+iso, **p = 0.029). d, Representative images and density of 7/4+neutrophils (red). Stained are capillaries (t.lectin+, white) and nuclei (DAPI, blue) in the ischemic hemisphere at 3days poststroke (groups described in b, 2-way ANOVA, Sidak’s post-hoc test; n = 8 mice/group; pool of 2-independent experiments; ***p = 0.0001, *p = 0.048). e, mNSS score (n = 12 Aged+iso, n = 11 Aged+1A8; data are pooled from 3-independent experiments, Mixed-Effect-Model, Bonferroni post-hoc test, *p = 0.0319 and *p = 0.038). f, Quantification of peri-ischemic NeuN+neurons at 3days poststroke (n = 7. Aged+iso; n = 9, Aged+1A8, *p = 0.0117, data pooled from 2-independent experiments), ischemic volume and edema (n = 11, Aged+iso; n = 12 Aged+1A8). Dots in scatterplots correspond to a single mouse; mean±SEM is shown. 2-tailed t-test if not otherwise indicated. Scale bar c, d, 25µm. g-n, Analysis of blood neutrophils at 24h poststroke by scRNAseq. g, Scatterplots of differentially expressed genes between neutrophil clusters (as in Fig. 3). h-j, Violin plots showing, h, neutrophil activation score, i, neutrophil aging-score (from genes reported in Xie et al.17). j, chemotaxis score for neutrophil cluster as in Fig. 3b. k, Heatmap showing row-scaled expression of granule-related genes and related l, violin plots of azurophil, specific, gelatinase and secretory scores for neutrophil clusters (as in Fig. 3b). In h, i, j, l Bonferroni-corrected Wilcoxon rank sum test, *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001; ****p ≤ 0.0001. All comparisons in Supplementary Table 3; boxes in violin plots extend from the 25th-75th percentiles, midline at median. m, n, UMAPs showing the weighted average expression for the signature of NADPH Oxidase, Neutrophil activation, ROS Production and NETs associated genes (as in Fig. 3b) corresponding to violin plots shown in Extended Data Fig. 3h and Fig. 3l, m. In n the correlations between the functional scores within the cluster scNeu1 (as in Fig. 3b) are shown.
Extended Data Fig. 4 Neutrophil subsets in the blood, brain and BM of young and aged stroke mice.
a, Quantification of circulating neutrophil subsets by 18-color MCFC at 0h, 6h and 24h poststroke, (n = 4 to 6 mice/group/ timepoint, 2-ANOVA, Bonferroni-post-hoc test, **p ≤ 0.01; ****p≤0.0001). b, Heatmap displaying the median antigen intensity in blood neutrophil clusters analyzed by MCFC at 24h poststroke and dotplot showing the scaled expression of selected genes, corresponding to markers of MCFC, of the scRNAseq stroke blood neutrophil subsets at 24h poststroke (neutrophil subsets from Fig. 3). The table shows the nomenclature and marker antigens and genes for major neutrophil subsets and GSVA based comparison (shown in Fig. 3g) with the nomenclature used by Xie et al.17. c, Histograms, and barplots showing the median fluorescence intensity (MFI) of CD177, CD11b and CD101 on neutrophils infiltrating the ischemic lesion analyzed by MCFC at 24h poststroke in young and aged mice (n = 5 mice/group, 2-tailed, Wilcoxon Mann-Whitney, *p = 0.031, **p = 0.0079, each dot corresponds to a single mouse). d, Representative IF images of CD177, CD101 or CD14 (in red) on neutrophils (Ly6G+in green, or white and 7/4+ in white) infiltrating the ischemic lesion of aged mice at 24h poststroke (blood vessels, stained by t-lectin, white) Scale bar, 25µm. e, Quantification of neutrophils and progenitors in the BM by MCFC at 0h and 24h poststroke of young and aged mice (n = 4- 6 mice per group per timepoint, 2-ANOVA Bonferroni post-hoc test; *p ≤ 0.05; **p ≤ 0.01; ****p≤0.0001). f, Barplots showing frequencies among mature BM neutrophils or BM neutrophil at 6h poststroke (n = 3 young and 4 aged mice, 2-tailed t-test, *p ≤ 0.05). g, ELISA for selected blood cytokines and chemokines at 0h and 48h poststroke; (n = 5 to 7 mice per group per timepoint, 2-ANOVA Bonferroni post-hoc test *p ≤ 0.05; **p≤0.01, ***p ≤ 0.001; §p≤0.055). In a, c, e and g, the box extends from the 25th- 75th percentiles, midline at median, whiskers plot min and max.
Extended Data Fig. 5 MCFC analysis of BM hematopoietic stem cells and progenitors after stroke.
a, 18-color MCFC analysis of live CD45+Lin-Ly6G-CD11b-ckit+ BM progenitor cells from young and aged mice at steady-state (0h) and 24h poststroke (n = 6young, n = 4 aged at 0h; n = 6 mice per group at 24h). Shown is the PHATE-algorithm based map showing BM progenitor clustering. b, Individual PHATE plots are overlaid with the clustering markers from the panel and c, corresponding quantification (groups as in a). The box plots extend from the 25th to 75th percentiles, the midline denotes the median while whiskers plot min and max, each dot corresponds to a single mouse (n = 4 to 6 mice per group per timepoint). 2-ANOVA, Benjamini-Hochberg post-hoc test *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001.
Extended Data Fig. 6 scRNAseq of BM and blood granulopoiesis of young and aged mice at steady-state and after stroke.
Integrated analysis of scRNAseq datasets of blood and BM cells from young (2-4months) and aged (18–24 months) male mice at steady state (0h) and 24h poststroke. a, Combined UMAP of the extracted neutrophil and neutrophil precursor subsets (total of 18’154 cells, see Supplementary Fig. 2, Supplementary Table 4) from blood and BM (see inset) of young and aged male mice at steady state and 24h poststroke colored by Seurat guided cluster identity. In brackets the inferred neutrophil population name following the nomenclature of Xie et al.17. b. The Blood and BM neutrophil precursors and neutrophil subsets shown in panel a, are here displayed divided for condition (steady state or 24h poststroke) and for age (young and aged mice). c, Dot plot for scaled expression of the gene signature for each neutrophil and neutrophil progenitor cluster (as in a). d, GSVA based comparison of the indicated neutrophil seurat clusters (as in a) with scRNA-seq-defined neutrophil populations reported by Xie et al.17 (HSPC, G0 to G5); GSVA score calculated on the list of top genes expressed per cluster75. The score of indicated defined clusters (HSPC, G0-G5) for each group of neutrophil subsets is shown (cluster 4, cMOP, is not computed). e, Graphs showing the proportion of cells for each cluster in young or aged mice, for the blood or BM at steady state and 24h poststroke (cluster 4, cMOP, not computed). f, Projection of pseudo-time generated with Monocle version 3 to infer the potential lineage differentiation trajectory onto the combined UMAP; the selected starting point is cluster 14 (HSPC). g, Expression of selected genes along pseudotime for granulocyte differentiation (Cluster 14, 6, 3, 9, 12, 10, 7, 8, 2, 0, 13, 1, 5) either at steady state or at 24h poststroke in young and aged mice.
Extended Data Fig. 7 Differentially expressed genes of granulopoietic clusters in the BM and blood among young and aged mice.
a, b, Heatmap showing the log2 [fold change] of selected differential expressed genes of aged over young mice in steady state (a) and 24h poststroke (b) for each main cluster of the BM (similar clusters have been computed together, 9 and 12 (G1) and 7 and 10 (G2)). Genes are marked with an asterisk if their expression changed significantly, as identified by 2-tailed t-test, Bonferroni-corrected p value, *p ≤ 0.05). c, Dotplot showing the scaled expression of signature genes for each cluster in the BM 24h poststroke in young (Y) and aged (A) mice, colored by the average expression of each gene in each cluster scaled across the indicated clusters (as in a). Dot size represents the percentage of cells in each cluster with more than one read of the corresponding gene. d, Projection of the stem-score, proliferation-score and G-CSF score signatures33 onto the combined UMAP (as in Extended Data Fig. 6a) comprehending blood and BM neutrophils and neutrophil progenitors of young and aged mice at steady state and 24h poststroke. The dotplots displayed below each score report for HSPC (cluster 14), G0_CMP (cluster 6), G0_GMP (cluster 3), G1 (cluster 9 and 12) and G2 (cluster 7 and 10) and G3 (cluster 8) the mean signature score. e, Scatterplots showing the differentially expressed genes in the blood of aged stroke mice compared to young stroke mice within the four mostly represented neutrophil clusters of the blood (cluster 2, 0, 1, 5 as in Extended Data Fig. 6a). Colored dots display an adjusted p- value p≤0.05, as identified by 2-tailed t-test, Bonferroni-corrected p value *p ≤ 0.05.
Extended Data Fig. 8 Neutrophils’ functional assays and heterochronic neutrophil transfer in stroke.
a, ROS production by blood neutrophil subsets (as defined in Fig. 4) in young and aged mice 24h poststroke assessed by MCFC. b, Phagocytosis of GFP+ E. coli by mature BM neutrophil subsets in young and aged mice. c, Neutrophil-platelet aggregates (NPAs) (Ly6G+ CD41+ cells) in the blood at 24h poststroke. d, Representative MCFC-plots for NPA in Neu3a and quantification of NPAs among neutrophils. e, NETting neutrophils (citH3+MPO+ cells) in the blood of young and aged mice at 24h poststroke. f, Representative MCFC plots of the NETting Neu2 subset and frequencies of citH3+MPO+NETting neutrophils subsets. In a-f n = 4 to 5 mice/group). g, MCFC plots of CD62LhiCXCR4lo and CD62LloCXCR4hi gated either on Ly6G+CD101+ or on Ly6G+CD101lo blood neutrophils of young and aged mice 24h poststroke. Microscopy images of sorted neutrophils stained by May-Grunwald/Giemsa. h, Quantification of neutrophils, band cells and hypersegmented neutrophils, on blood smears of young and aged mice at 0h, 1h, 24h poststroke (insets are May-Grünwald/Giemsa stained cells, n = 3 to 15 mice per group per timepoint). In h, the box extends from the 25th - 75th percentiles, midline at median, whiskers plot min and max. In a, b, d, f, h 2-ANOVA, Sidak’s post-hoc test, in c, e two-tailed t-test. *p ≤ 0.05; **p ≤ 0.01. i-j, MCFC at 48h post-stroke on the inflammatory brain infiltrate of CD45.1+ young recipients intravenously transferred with BM neutrophils from young or aged CD45.2+ donors. i, MCFC plots of inflammatory cells within the ipsilateral (ipsi) and contralateral (contra) hemispheres and quantification (n = 4 mice per group, contra-hemispheres were pooled in groups of two). j, Representative immunofluorescence image of labeled (CellTracker+ cells, green) aged-donor neutrophils (Ly6G+, red) in the ischemic lesion of young stroke recipients. Blood vessels (t.lectin+, white). k-m, Adoptive intravenous transfer of blood leukocytes from young(YtoY) or aged (AtoY) or aged donors with neutrophil depleted leukocytes(AnoNEUtoY) at 24h poststroke to young stroke recipients. k, LDF-recordings, l, mNSS-score, m, quantification of peri-ischemic NeuN+neurons, ischemic volume and edema at 72h poststroke. In k, l, 2-ANOVA, Tukey post-hoc test; m, 1-ANOVA, Tukey post-hoc test; n = 6 YtoY, n = 6 AtoA, n = 5 AnoNEUtoY, except in k n = 4 mice; *p ≤ 0.05; **p ≤ 0.01. n-o, Adoptive intravenous-transfer of neutrophils from wild-type donors (NtoY), or PEdKO donors (NKOtoY), or neutrophil-depleted PEdKO donors (NKO+𝜶Gr-1toY) into young stroke mice. n, MCFC plots of transferred cells; LDF-recordings and mNSS-score and o, immunofluorescence-images and quantification of peri-ischemic NeuN+neurons (green), and ischemic volume evaluation at 3 days poststroke. (n = 5 to 6 NtoY, n = 5 to 6 NKOtoY and n = 4 NKO+𝜶Gr-1toY). In n 2-ANOVA, Tukey post-hoc test; in o, 1-ANOVA, Tukey post hoc test; *p ≤ 0.05; ***p ≤ 0.001. In the scatterplots each dot corresponds to a single mouse; mean±SEM is shown. Scale bar, h, 5µm, j, o, 50µm. Nuclei (DAPI, blue).
Extended Data Fig. 9 CXCL3 worsens stroke outcome.
a, GSEA-enrichment curve for IL8 and CXCR2 signaling in aged over young mice at 24h poststroke calculated on bulk RNA-seq data (Fig. 7) and violin plot showing CXCR2 signature score in scRNAseq neutrophil subsets (as in Fig. 3b). b, Bar graphs of CD62L, CXCR4, CXCR2 and CD11b MCFC-based MFI on blood neutrophils after in vitro CXCL3 (or vehicle) stimulation (2-tailed t-test; n = 4 mice/group, *p ≤ 0.05) and c, histograms. d-g, In vivo intravenous injection of CXCL3 (or vehicle) in young mice. d, Analysis of total blood neutrophils 2h post-injection, e, representative MCFC plots, f, quantification of neutrophil subsets (as in Fig. 4d). In d-g n = 5 to 6 mice per group; one of three independent experiments is shown; two-tailed t-test; ***p ≤ 0.001, **p ≤ 0.01,. h-k, In vivo intravenous injection of CXCL3 (or vehicle) to young mice and stroke induction. h, LDF-recordings (n = 9 vehicle vs n = 8 CXCL3, 2-ANOVA, Benjamin-Hochberg post hoc test), i, mNSS-score, and j, quantification of peri-ischemic NeuN+neurons and ischemic volume 3 days poststroke. k, Immunofluorescence images and quantification of perilesional Ly6G+neutrophils (green) at 3 days poststroke. Blood vessels (t.lectin+, red), nuclei (DAPI, blue). In i-k, n = 5 to 8 mice per group, two-tailed t-test except for mNSS, 2-ANOVA, Sidak’s post hoc test. l-p, In vivo intraperitoneal injection of anti-CXCL3 antibody (A+αCXCL3) or isotype (A+isotype) in aged mice undergoing stroke. l, LDF (n = 9 A+isotype; n = 8, A+αCXCL3, 2-ANOVA, Benjamin-Hochberg post hoc test), m, mNSS-score and n, quantification of peri-ischemic NeuN+neurons, o, ischemic volume and edema at 3 days poststroke. p, Quantification of perilesional Ly6G+neutrophils 3days poststroke. In m-p, n = 5 mice per group, 2-tailed t-test, except for mNSS, Mixed-effects model, Holm-Sidak’s post hoc test. q-u, Adoptive transfer of BM neutrophils from aged donors into young mice with stroke, pre-treated with anti-CXCL3 (AtoY+αCXCL3). The control groups YtoY and AtoY are reported from Fig. 6c. q, LDF, r, mNSS-score and s, quantification of peri-ischemic NeuN+neurons, t, ischemic volume, and edema. u, Quantification of perilesional Ly6G+ neutrophils at 3days poststroke. n = 5 to 8 mice per group; 1-ANOVA, Tukey post-hoc test; for mNSS, Mixed-effects model, Sidak’s post hoc test; for LDF, 2-ANOVA Tukey post-hoc test. Dots in scatterplots corresponds to a single mouse; mean±SEM is shown. *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001.
Extended Data Fig. 10 Leukocytes subsets in young and old stroke patients.
a-d. Mass cytometry analysis of blood leukocytes (groups as in Fig. 8a). a, Representative UMAP map showing the FlowSOM-guided meta-clustering of blood leukocytes. b, Heatmap displaying the MAE of markers used to generate the UMAP. c, Bar graphs showing frequencies of leukocyte subsets, and donut graphs of non-neutrophil leukocytes in MC and in SP at the indicated timepoints. d, Violin plots displaying quantifications of circulating leukocytes in MC and in SP at D1, D3 and D7 after stroke. e, Flow cytometry analyis of total white blood cells (WBCs), neutrophils, lymphocytes, and neutrophil-to-lymphocytes ratio (NLR) (groups as in Fig. 8f). f, Average thrombogram of control conditions for tissue factor-triggered thrombin generation in FV and FVIII-deficient plasma (depleted), in normal pool plasma (NPP), in mixtures of factor V and VIII-deficient plasma and NPP with final factor V and VIII concentrations of 1% and 0.5% (NPP0.5% and NPP 1%), and ellagic acid-triggered thrombin generation in factor V and VIII-deficient plasma (depleted_FSL). No ellagic acid-triggered thrombin generation was observed in factor V and VIII-deficient plasma-neutrophils mixtures (any neutrophil preparation). Thrombin generation did not occur with a reagent containing phospholipid vesicles in the absence of tissue factor. g, Representative flow cytometry plots of purified human neutrophils. In d, e 2-tailed multiple Mann-Whitney Wilcoxon test; *refers to comparisons between young-SP and old-SP. For comparison of young-SP or old-SP with respectively young-MC or old-MC Kruskall-Wallis with Dunn’s post-hoc test: ^refers to comparisons with MC-young; # refers to comparisons with MC-old; *,^,#p ≤ 0.05, **,^^,## p ≤ 0.01, ***,^^^,### p ≤ 0.001, ****,#### p ≤ 0.0001.
Supplementary information
Supplementary Information
Supplementary Tables 1,7 and 8 and Figs 1–5.
Supplementary Table 2
Flow cytometry leukocytes counts of blood from young and aged mice.
Supplementary Table 3
Stroke blood scRNA-seq quality control (QC) parameters, cell cluster signature genes and gene lists used for enrichment scores with related P values.
Supplementary Table 4
Blood steady state (0 h), blood stroke (24 h), BM steady state (0 h) and BM stroke (24 h) scRNA-seq.
Supplementary Table 5
Tabula Muris Senis.
Supplementary Table 6
Bulk RNA-seq.
Source data
Source Data Fig. 1
Statistical Source Data.
Source Data Fig. 2
Statistical Source Data.
Source Data Fig. 4
Statistical Source Data.
Source Data Fig. 5
Statistical Source Data.
Source Data Fig. 6
Statistical Source Data.
Source Data Fig. 7
Statistical Source Data.
Source Data Fig. 8
Statistical Source Data.
Source Data Extended Data Fig. 1
Statistical Source Data.
Source Data Extended Data Fig. 2
Statistical Source Data.
Source Data Extended Data Fig. 3
Statistical Source Data.
Source Data Extended Data Fig. 4
Statistical Source Data.
Source Data Extended Data Fig. 5
Statistical Source Data.
Source Data Extended Data Fig. 6
Statistical Source Data.
Source Data Extended Data Fig. 8
Statistical Source Data.
Source Data Extended Data Fig. 9
Statistical Source Data.
Source Data Extended Data Fig. 10
Statistical Source Data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gullotta, G.S., De Feo, D., Friebel, E. et al. Age-induced alterations of granulopoiesis generate atypical neutrophils that aggravate stroke pathology. Nat Immunol 24, 925–940 (2023). https://doi.org/10.1038/s41590-023-01505-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-023-01505-1
This article is cited by
-
An aging-related immune landscape in the hematopoietic immune system
Immunity & Ageing (2024)
-
Type I Interferon Signalling and Ischemic Stroke: Mechanisms and Therapeutic Potentials
Translational Stroke Research (2024)
-
Inflammaging aggravates stroke pathology
Nature Immunology (2023)
-
Immunology of human fibrosis
Nature Immunology (2023)