Abstract
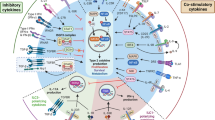
Elucidating the mechanisms that sustain asthmatic inflammation is critical for precision therapies. We found that interleukin-6- and STAT3 transcription factor-dependent upregulation of Notch4 receptor on lung tissue regulatory T (Treg) cells is necessary for allergens and particulate matter pollutants to promote airway inflammation. Notch4 subverted Treg cells into the type 2 and type 17 helper (TH2 and TH17) effector T cells by Wnt and Hippo pathway-dependent mechanisms. Wnt activation induced growth and differentiation factor 15 expression in Treg cells, which activated group 2 innate lymphoid cells to provide a feed-forward mechanism for aggravated inflammation. Notch4, Wnt and Hippo were upregulated in circulating Treg cells of individuals with asthma as a function of disease severity, in association with reduced Treg cell-mediated suppression. Our studies thus identify Notch4-mediated immune tolerance subversion as a fundamental mechanism that licenses tissue inflammation in asthma.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The data presented in the manuscript, including de-identified patient results, will be made available to investigators after a request to the corresponding author. Any data and materials to be shared will be released via a material transfer agreement. RNA-seq datasets have been deposited in the Gene Expression Omnibus with the accession no. GSE151763. Source data are provided with this paper.
Change history
19 November 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41590-020-00841-w
26 April 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41590-021-00929-x
References
Lambrecht, B. N. & Hammad, H. The immunology of asthma. Nat. Immunol. 16, 45–56 (2015).
Martinez, F. D. & Vercelli, D. Asthma. Lancet 382, 1360–1372 (2013).
Noval Rivas, M. & Chatila, T. A. Regulatory T cells in allergic diseases. J. Allergy Clin. Immunol. 138, 639–652 (2016).
Krishnamoorthy, N. et al. Early infection with respiratory syncytial virus impairs regulatory T cell function and increases susceptibility to allergic asthma. Nat. Med. 18, 1525–1530 (2012).
Noval Rivas, M. et al. Regulatory T cell reprogramming toward a Th2-cell-like lineage impairs oral tolerance and promotes food allergy. Immunity 42, 512–523 (2015).
Massoud, A. H. et al. An asthma-associated IL4R variant exacerbates airway inflammation by promoting conversion of regulatory T cells to TH17-like cells. Nat. Med. 22, 1013–1022 (2016).
Xia, M. et al. Vehicular exhaust particles promote allergic airway inflammation through an aryl hydrocarbon receptor–notch signaling cascade. J. Allergy Clin. Immunol. 136, 441–453 (2015).
Xia, M., Harb, H., Saffari, A., Sioutas, C. & Chatila, T. A. A Jagged 1–Notch 4 molecular switch mediates airway inflammation induced by ultrafine particles. J. Allergy Clin. Immunol. 142, 1243–1256.e17 (2018).
Tsai, V. W. W., Husaini, Y., Sainsbury, A., Brown, D. A. & Breit, S. N. The MIC-1/GDF15-GFRAL pathway in energy homeostasis: implications for obesity, cachexia, and other associated diseases. Cell Metab. 28, 353–368 (2018).
Luan, H. H. et al. GDF15 is an inflammation-induced central mediator of tissue tolerance. Cell 178, 1231–1244.e11 (2019).
Chen, W. et al. Conversion of peripheral CD4+CD25- naive T cells to CD4+CD25+ regulatory T cells by TGF-β induction of transcription factor Foxp3. J. Exp. Med. 198, 1875–1886 (2003).
Shi, S. & Stanley, P. Protein O-fucosyltransferase 1 is an essential component of Notch signaling pathways. Proc. Natl Acad. Sci. USA 100, 5234–5239 (2003).
Charbonnier, L. M., Wang, S., Georgiev, P., Sefik, E. & Chatila, T. A. Control of peripheral tolerance by regulatory T cell–intrinsic Notch signaling. Nat. Immunol. 16, 1162–1173 (2015).
Han, H. et al. Inducible gene knockout of transcription factor recombination signal binding protein-J reveals its essential role in T versus B lineage decision. Int. Immunol. 14, 637–645 (2002).
Tachdjian, R. et al. Pathogenicity of a disease-associated human IL-4 receptor allele in experimental asthma. J. Exp. Med. 206, 2191–2204 (2009).
Shi, H. et al. Hippo kinases Mst1 and Mst2 sense and amplify IL-2R-STAT5 signaling in regulatory T cells to establish stable regulatory activity. Immunity 49, 899–914.e6 (2018).
Geng, J. et al. The transcriptional coactivator TAZ regulates reciprocal differentiation of TH17 cells and Treg cells. Nat. Immunol. 18, 800–812 (2017).
van Loosdregt, J. et al. Canonical Wnt signaling negatively modulates regulatory T cell function. Immunity 39, 298–310 (2013).
van Loosdregt, J. & Coffer, P. J. The role of WNT signaling in mature T cells: T cell factor is coming home. J. Immunol. 201, 2193–2200 (2018).
Misra, J. R. & Irvine, K. D. The Hippo signaling network and its biological functions. Annu Rev. Genet 52, 65–87 (2018).
Clevers, H. & Nusse, R. Wnt/β-catenin signaling and disease. Cell 149, 1192–1205 (2012).
Feng, Y. et al. Control of the inheritance of regulatory T cell identity by a cis element in the Foxp3 locus. Cell 158, 749–763 (2014).
Li, X., Liang, Y., LeBlanc, M., Benner, C. & Zheng, Y. Function of a Foxp3 cis-element in protecting regulatory T cell identity. Cell 158, 734–748 (2014).
Vivier, E. et al. Innate lymphoid cells: 10 years on. Cell 174, 1054–1066 (2018).
Esty, B. et al. Treatment of severe persistent asthma with IL-6 receptor blockade. J. Allergy Clin. Immunol. Pract. 7, 1639–1642.e4 (2019).
Hirota, T. et al. Genome-wide association study identifies three new susceptibility loci for adult asthma in the Japanese population. Nat. Genet. 43, 893–896 (2011).
Ferreira, M. A. et al. Identification of IL6R and chromosome 11q13.5 as risk loci for asthma. Lancet 378, 1006–1014 (2011).
Gruzieva, O. et al. Prenatal particulate air pollution and DNA methylation in newborns: an epigenome-wide meta-analysis. Environ. Health Perspect. 127, 57012 (2019).
Li, X. et al. Genome-wide association study identifies TH1 pathway genes associated with lung function in asthmatic patients. J. Allergy Clin. Immunol. 132, 313–320.e15 (2013).
Savenije, O. E. et al. Association of IL33-IL-1 receptor-like 1 (IL1RL1) pathway polymorphisms with wheezing phenotypes and asthma in childhood. J. Allergy Clin. Immunol. 134, 170–177 (2014).
Soroosh, P. et al. Lung-resident tissue macrophages generate Foxp3+ regulatory T cells and promote airway tolerance. J. Exp. Med. 210, 775–788 (2013).
Magee, C. N. et al. Notch-1 inhibition promotes immune regulation in transplantation via regulatory T cell-dependent mechanisms. Circulation 140, 846–863 (2019).
Lee, P. P. et al. A critical role for Dnmt1 and DNA methylation in T cell development, function, and survival. Immunity 15, 763–774 (2001).
Messerschmidt, D. et al. β-Catenin-mediated adhesion is required for successful preimplantation mouse embryo development. Development 143, 1993–1999 (2016).
Rubtsov, Y. P. et al. Regulatory T cell-derived interleukin-10 limits inflammation at environmental interfaces. Immunity 28, 546–558 (2008).
McFarland-Mancini, M. M. et al. Differences in wound healing in mice with deficiency of IL-6 versus IL-6 receptor. J. Immunol. 184, 7219–7228 (2010).
Yang, X. et al. Notch activation induces apoptosis in neural progenitor cells through a p53-dependent pathway. Dev. Biol. 269, 81–94 (2004).
McCright, B., Lozier, J. & Gridley, T. Generation of new Notch2 mutant alleles. Genesis 44, 29–33 (2006).
Krebs, L. T. et al. Characterization of Notch3-deficient mice: normal embryonic development and absence of genetic interactions with a Notch1 mutation. Genesis 37, 139–143 (2003).
Barnden, M. J., Allison, J., Heath, W. R. & Carbone, F. R. Defective TCR expression in transgenic mice constructed using cDNA-based α- and β-chain genes under the control of heterologous regulatory elements. Immunol. Cell Biol. 76, 34–40 (1998).
Moh, A. et al. Role of STAT3 in liver regeneration: survival, DNA synthesis, inflammatory reaction and liver mass recovery. Lab. Invest. 87, 1018–1028 (2007).
Zhang, N. et al. The Merlin/NF2 tumor suppressor functions through the YAP oncoprotein to regulate tissue homeostasis in mammals. Dev. Cell 19, 27–38 (2010).
Reginensi, A. et al. Yap- and Cdc42-dependent nephrogenesis and morphogenesis during mouse kidney development. PLoS Genet. 9, e1003380 (2013).
Binder, A. K. et al. Expression of human NSAID activated gene 1 in mice leads to altered mammary gland differentiation and impaired lactation. PLoS ONE 11, e0146518 (2016).
Charbonnier, L. M. et al. Functional reprogramming of regulatory T cells in the absence of Foxp3. Nat. Immunol. 20, 1208–1219 (2019).
Zheng, Y. et al. Role of conserved non-coding DNA elements in the Foxp3 gene in regulatory T-cell fate. Nature 463, 808–812 (2010).
Povoleri, G. A. M. et al. Human retinoic acid–regulated CD161+ regulatory T cells support wound repair in intestinal mucosa. Nat. Immunol. 19, 1403–1414 (2018).
Acknowledgements
This work was supported by National Institutes of Health grants (nos. R01 AI115699 and R01 AI065617 to T.A.C., U01AI110397 and R01 HL137192 to W.P.) a National Health and Medical Research Council grant (no. APP1163249 to B.G.) and a German Research Society grant (no. HA 8465/1-1 to H.H.).
Author information
Authors and Affiliations
Contributions
H.H. and T.A.C. designed experiments. H.H., E.S-V, M.B., A.M., Y.C, L-M.C. and S.A. performed experiments and developed experimental models. E.C., S.B., A.C. and W.P. recruited patients and analyzed their demographics. K.S.A. and B.G. analyzed the RNA-seq data. J.M.L.C. and R.S.G. provided RoraCre and RoraCreIl4∆/∆Il13∆/∆ mice. C.S. and A.J.M provided UFPs. H.H. and T.A.C. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
T.A.C., H.H. and A.M. are inventors on published US patent application no. WO2019178488A1 submitted by the Children’s Medical Center Corporation, titled ‘Method for treating asthma or allergic disease’. B.G. is a director of Pacific Analytics and SMRTR, Australia; a founding member of the International Cerebral Palsy Genetics Consortium; and a member of the Australian Genomics Health Alliance. W.P. is a Consultant for Genentech, Novartis, Regeneron, Sanofi Genzyme and Glaxo Smith Kline, and receives clinical trial support from Genentech, Novartis, Regeneron, Circassia, Thermo Fisher, Monaghan, Lincoln Diagnostics, Alk Abello and Glaxo Smith Kline.
Additional information
Peer review information Zoltan Fehervari was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Notch4 expression on lung Treg cells in allergic airway inflammation.
a-c, Flow cytometric analysis, cell frequencies and (MFI) of Notch1, 2 and 3 expression on lung Treg and Teff cells in Foxp3YFPCre (n = 5). d, Cell frequencies of Notch4 expression on OT-II+CD4+Foxp3+ T cells generated in co-cultures with sham or OVA323-339 + UFP-pulsed alveolar macrophages without or with IL-1β, IL-25, IL-33, TSLP or TNF (n = 5). e, ChIP assays for the binding of STAT3 and control (IgG) antibodies to the Notch1, 2 and 3 promoters in lung Treg cells of OVA + UFP-treated Foxp3YFPCre, and Foxp3YFPCreStat3∆/∆ mice (n = 5). Each symbol represents one mouse. Numbers in flow plots indicate percentages. Error bars indicate SEM. Statistical tests: One−way ANOVA with Dunnett’s post hoc analysis (a-c); two-way ANOVA with Sidak’s post hoc analysis (d,e). **P < 0.01, ***P < 0.001, ****P < 0.0001. Data representative of two or three independent experiments.
Extended Data Fig. 2 Notch4 expression on lung Treg cells licenses allergic airway inflammation.
a, RT-PCR analysis of Notch4 expression in CD4Cre mice in B-cells and T-cells (n = 5). b, RT-PCR analysis of Notch4 expression in Foxp3YFPCre mice in both Treg and Teff cells (n = 5). c,d, IL-4 and IFN-γ expression in lung Foxp3+CD4+ Treg. (c) and Foxp3–CD4+Teff cells. (d) derived from the respectively treated Foxp3YFPCre, CD4CreNotch4∆/∆ and Foxp3YFPCreNotch4∆/∆ mice (n = 5). e, Airway hyperresponsiveness in Foxp3YFPCre sensitized either with PBS or OVA, then challenged with OVA + UFP following transfer of OTII+Foxp3YFPCre or OTII+Foxp3YFPCreNotch4∆/∆ iTreg cells (n = 5). f, Eosinophil numbers for the respective mouse groups (n = 5). g, IL-4, IL-13, IL-17 and IFNγ expression in lung Foxp3–CD4– Teff cells. Each symbol represents one mouse (n = 5). Error bars indicate SEM. Statistical tests: two-way ANOVA with Sidak’s post hoc analysis (a,c,d); One-way ANOVA with Dunnett’s post hoc analysis (e,f). *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. Data representative of two or three independent experiments.
Extended Data Fig. 3 Allergic airway inflammatory responses in mice with Treg cell-specific Pofut1 or Rbpj1 deletion.
a, Representative PAS-stained sections of lung tissues isolated from Foxp3YFPCre, Foxp3YFPCrePofut1∆/∆ or Foxp3YFPCreRbpj1∆/∆ mice segregated into PBS, OVA or OVA + UFP-treated groups (200X magnification). b, Inflammation scores in the respective lung tissues. c, AHR in the respective mouse groups in response to methacholine. d,e, serum total and OVA-specific IgE concentrations. f,g, absolute numbers of lung CD4+ T cells and eosinophils. h,i, IL-4, IL-13, IL-17 and IFNγ expression in lung Foxp3+CD4+ Treg (h) and Foxp3–CD4+Teff cells (i). Each symbol represents an independent sample. Error bars indicate SEM. Statistical tests: two-way ANOVA with Sidak’s post hoc analysis (b-i). *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. Data representative of two or three independent experiments. n = 5 mice per group.
Extended Data Fig. 4 Allergic airway inflammatory responses in mice with Treg cell-specific Notch1 or Notch2 deletion or global Notch3 deletion.
a-c, AHR in Foxp3YFPCre, Foxp3YFPCreNotch1∆/∆, Foxp3YFPCreNotch2∆/∆, or Foxp3YFPCreNotch3–/– mice segregated into PBS, OVA or OVA + UFP-treated groups (200X magnification). d, serum OVA-specific IgE concentrations. e,f, absolute numbers of lung CD4+ T cells and eosinophils. g,h, IL-4, IL-13, and IL-17 expression in lung Foxp3–CD4+Teff (g) and Foxp3+CD4+Treg cells (h). Each symbol represents an independent sample. Error bars indicate SEM. Statistical tests: two-way ANOVA with Sidak’s post hoc analysis a-h. Data representative of two or three independent experiments. n = 5 mice per group.
Extended Data Fig. 5 Treg cell-specific Il6r and stat3 deletions attenuate allergic airway inflammation.
a, Representative PAS-stained sections of lung tissues isolated from Foxp3YFPCre, Foxp3YFPCreIl6r∆/∆ or Foxp3YFPCreStat3∆/∆ mice segregated into PBS, OVA or OVA + UFP-treated groups (200X magnification). b, Inflammation scores in the respective lung tissues. c, AHR in the respective mouse groups in response to methacholine. d,e, serum total and OVA-specific IgE concentrations. f,g, absolute numbers of lung CD4+ T cells and eosinophils. h,i, IL-13 and IL-17 expression in lung Foxp3+CD4+ Treg (h) and Foxp3–CD4+ Teff cells (i). Each symbol represents an independent sample. Error bars indicate SEM. Statistical tests: two-way ANOVA with Sidak’s post hoc analysis (b-i). *P < 0.05, *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. Data representative of two or three independent experiments. n = 5 mice per group.
Extended Data Fig. 6 Treg cell-specific Notch4 deletion rescues HDM induced allergic airway inflammation.
a, scheme of the house dust mite (HDM) airway inflammation protocol. b, Representative PAS-stained sections of lung tissues isolated from Foxp3YFPCre or Foxp3YFPCreNotch4∆/∆ mice segregated into PBS, OVA or OVA + UFP-treated groups (200X magnification). c, Inflammation scores in the respective lung tissues. d, AHR in the respective mouse groups in response to methacholine. e, serum total IgE concentrations. (f-h), absolute numbers of lung CD4+ T cells, neutrophils and eosinophils. i,k, IL-4, IL-13, IL-17 and IFNγ expression in lung Foxp3+CD4+ Treg (i) and Foxp3–CD4+ Teff cells (k). Each symbol represents an independent sample. Numbers in flow plots indicate percentages. Error bars indicate SEM. Statistical tests: two-way ANOVA with Sidak’s post hoc analysis (c-k). **P < 0.01, ***P < 0.001, ****P < 0.0001. Data representative of two or three independent experiments. n = 5 mice per group.
Extended Data Fig. 7 Treg cell-specific Notch4 deletion rescues chronic allergic airway inflammation.
a, Scheme for the chronic airway inflammation mouse protocol b, Representative Sirius-Red-stained sections of lung tissues isolated from Foxp3YFPCre or Foxp3YFPCreNotch4∆/∆ mice segregated into PBS, OVA or OVA + UFP-treated groups (200X magnification). c, Collagen disposition measurement in the respective lung tissues. d, AHR in the respective mouse groups in response to methacholine. e,f, absolute numbers of lung CD4+ T cells and eosinophils. g,h, IL-4, IL-13, and IL-17 expression in lung Foxp3+CD4+ Treg (g) and Foxp3–CD4+Teff cells (h). i, Serum OVA-specific IgE titers in the respective groups. Each symbol represents an independent sample. Error bars indicate SEM. Statistical tests: two-way ANOVA with Sidak’s post hoc analysis (c-h). *P < 0.05, ****P < 0.0001. Data representative of two or three independent experiments. n = 5 mice per group.
Extended Data Fig. 8 Notch receptor expression in human Treg and Teff cells.
a,b, Flow cytometric analysis, cell frequencies and mean fluorescence intensity (MFI) of Notch1, 2 and 3 expression in peripheral blood Treg cells (a) and Teff cells (b) of control and asthmatic subjects, the latter segregated for asthma severity (control n = 22, M.P n = 15, Mod n = 16. S.P n = 11). c, Flow cytometric analysis and cell frequencies of Notch4 peripheral blood Treg cells of healthy control, food allergy (FA), eczema and FA + eczema (Control n = 37, FA n = 28, Eczema n = 10 and FA + Eczema n = 20) d, Serum GDF15 concentrations in asthmatic subjects plotted as a function of Notch4 expression on circulating Treg cells (n = 73) e, Cell frequencies of Notch4 expression in peripheral blood Treg cells in healthy subjects, allergic and non-allergic asthmatics (control = 56, non-allergic n = 21, allergic n = 85). Error bars indicate SEM. Statistical tests: One-way ANOVA with Dunnett’s post hoc analysis. (a-c,e); simple regression analysis (d). ***P < 0.001, ****P < 0.0001. Data representative of two or three independent experiments.
Supplementary information
Supplementary information
Supplementary Table 1.
Supplementary Dataset 1
RNA-seq analysis of lung Treg cells from OVA + UFP-treated Foxp3YFPCre and FoxpeYFPCreNotch4∆/∆.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Rights and permissions
About this article
Cite this article
Harb, H., Stephen-Victor, E., Crestani, E. et al. A regulatory T cell Notch4–GDF15 axis licenses tissue inflammation in asthma. Nat Immunol 21, 1359–1370 (2020). https://doi.org/10.1038/s41590-020-0777-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-020-0777-3
This article is cited by
-
Protective effect of low‐intensity pulsed ultrasound on immune checkpoint inhibitor-related myocarditis via fine-tuning CD4+ T-cell differentiation
Cancer Immunology, Immunotherapy (2024)
-
GDF15 as a key disease target and biomarker: linking chronic lung diseases and ageing
Molecular and Cellular Biochemistry (2024)
-
Pathogenesis of allergic diseases and implications for therapeutic interventions
Signal Transduction and Targeted Therapy (2023)
-
Exploring redox imbalance and inflammation for asthma therapy
Molecular Biology Reports (2023)
-
Neutrophil activation and NETosis are the predominant drivers of airway inflammation in an OVA/CFA/LPS induced murine model
Respiratory Research (2022)