Abstract
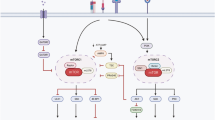
Aging is associated with remodeling of the immune system to enable the maintenance of life-long immunity. In the CD8+ T cell compartment, aging results in the expansion of highly differentiated cells that exhibit characteristics of cellular senescence. Here we found that CD27−CD28−CD8+ T cells lost the signaling activity of the T cell antigen receptor (TCR) and expressed a protein complex containing the agonistic natural killer (NK) receptor NKG2D and the NK adaptor molecule DAP12, which promoted cytotoxicity against cells that expressed NKG2D ligands. Immunoprecipitation and imaging cytometry indicated that the NKG2D–DAP12 complex was associated with sestrin 2. The genetic inhibition of sestrin 2 resulted in decreased expression of NKG2D and DAP12 and restored TCR signaling in senescent-like CD27−CD28−CD8+ T cells. Therefore, during aging, sestrins induce the reprogramming of non-proliferative senescent-like CD27−CD28−CD8+ T cells to acquire a broad-spectrum, innate-like killing activity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon request. The complete microarray data set is available online from the NCBI Gene Expression Omnibus public repository (GEO accession number GSE98640). The scRNA-seq data are available on EGA (accession number EGAS00001004255).
References
Akbar, A. N., Beverley, P. C. & Salmon, M. Will telomere erosion lead to a loss of T-cell memory? Nat. Rev. Immunol. 4, 737–743 (2004).
Gray, D. A role for antigen in the maintenance of immunological memory. Nat. Rev. Immunol. 2, 60–65 (2002).
Mitri, D. et al. Reversible senescence in human CD4+CD45RA+CD27− memory T cells. J. Immunol. 187, 2093–2100 (2011).
Henson, S. M. et al. p38 signaling inhibits mTORC1-independent autophagy in senescent human CD8+ T cells. J. Clin. Invest. 124, 4004–4016 (2014).
Lanna, A., Henson, S. M., Escors, D. & Akbar, A. N. The kinase p38 activated by the metabolic regulator AMPK and scaffold TAB1 drives the senescence of human T cells. Nat. Immunol. 15, 965–972 (2014).
Tarazona, R. et al. Increased expression of NK cell markers on T lymphocytes in aging and chronic activation of the immune system reflects the accumulation of effector/senescent T cells. Mech. Ageing Dev. 121, 77–88 (2001).
Lanna, A. et al. A sestrin-dependent Erk-Jnk-p38 MAPK activation complex inhibits immunity during aging. Nat. Immunol. 18, 354–363 (2017).
Dunne, P. J. et al. Quiescence and functional reprogramming of Epstein–Barr virus (EBV)-specific CD8+ T cells during persistent infection. Blood 106, 558–565 (2005).
Pereira, B. I. et al. Senescent cells evade immune clearance via HLA-E-mediated NK and CD8+ T cell inhibition. Nat. Commun. 10, 2387 (2019).
Krizhanovsky, V. et al. Senescence of activated stellate cells limits liver fibrosis. Cell 134, 657–667 (2008).
Dominguez, C. X. et al. The transcription factors ZEB2 and T-bet cooperate to program cytotoxic T cell terminal differentiation in response to LCMV viral infection. J. Exp. Medicine 212, 2041–2056 (2015).
Kovalovsky, D. et al. PLZF induces the spontaneous acquisition of memory/effector functions in T cells independently of NKT cell-related signals. J. Immunol. 184, 6746–6755 (2010).
Raberger, J. et al. The transcriptional regulator PLZF induces the development of CD44 high memory phenotype T cells. Proc. Natl Acad. Sci. USA 105, 17919–17924 (2008).
Liu, D. et al. Integrin-dependent organization and bidirectional vesicular traffic at cytotoxic immune synapses. Immunity 31, 99–109 (2009).
Bernardini, G., Sciumè, G. & Santoni, A. Differential chemotactic receptor requirements for NK cell subset trafficking into bone marrow. Front. Immunol. 4, 12 (2013).
van Lier, R. A., ten Berge, I. J. & Gamadia, L. E. Human CD8+ T-cell differentiation in response to viruses. Nat. Rev. Immunol. 3, 931–939 (2003).
Rufer, N. et al. Ex vivo characterization of human CD8+ T subsets with distinct replicative history and partial effector functions. Blood 102, 1779–1787 (2003).
Henson, S. M. et al. KLRG1 signaling induces defective Akt (ser473) phosphorylation and proliferative dysfunction of highly differentiated CD8+ T cells. Blood 113, 6619–6628 (2009).
Plunkett, F. J. et al. The loss of telomerase activity in highly differentiated CD8+CD28−CD27− T cells is associated with decreased Akt (Ser473) phosphorylation. J. Immunol. 178, 7710–7719 (2007).
Henson, S. M., Riddell, N. E. & Akbar, A. N. Properties of end-stage human T cells defined by CD45RA re-expression. Curr. Opin. Immunol. 24, 476–481 (2012).
Aktas, E., Kucuksezer, U., Bilgic, S., Erten, G. & Deniz, G. Relationship between CD107a expression and cytotoxic activity. Cell. Immunol. 254, 149–154 (2009).
Lanier, L. L. NKG2D receptor and its ligands in host defense. Cancer Immunol. Res. 3, 575–582 (2015).
Wu, J. et al. An activating immunoreceptor complex formed by NKG2D and DAP10. Science 285, 730–732 (1999).
Upshaw, J. L. et al. NKG2D-mediated signaling requires a DAP10-bound Grb2-Vav1 intermediate and phosphatidylinositol-3-kinase in human natural killer cells. Nat. Immunol. 7, 524–532 (2006).
Diefenbach, A. et al. Selective associations with signaling proteins determine stimulatory versus costimulatory activity of NKG2D. Nat. Immunol. 3, 1142–1149 (2002).
Gilfillan, S., Ho, E. L., Cella, M., Yokoyama, W. M. & Colonna, M. NKG2D recruits two distinct adapters to trigger NK cell activation and costimulation. Nat. Immunol. 3, 1150–1155 (2002).
Wu, J., Cherwinski, H., Spies, T., Phillips, J. H. & Lanier, L. L. Dap10 and Dap12 form distinct, but functionally cooperative, receptor complexes in natural killer cells. J. Exp. Med. 192, 1059–1068 (2000).
Akondy, R. S. et al. Origin and differentiation of human memory CD8 T cells after vaccination. Nature 552, 362–367 (2017).
Akondy, R. S. et al. The yellow fever virus vaccine induces a broad and polyfunctional human memory CD8+ T cell response. J. Immunol. 183, 7919–7930 (2009).
Wang, X. et al. Human invariant natural killer T cells acquire transient innate responsiveness via histone H4 acetylation induced by weak TCR stimulation. J. Exp. Med. 209, 987–1000 (2012).
Mingueneau, M. et al. Loss of the LAT adaptor converts antigen-responsive T cells into pathogenic effectors that function independently of the T cell receptor. Immunity 31, 197–208 (2009).
Wencker, M. et al. Innate-like T cells straddle innate and adaptive immunity by altering antigen-receptor responsiveness. Nat. Immunol. 15, 80–87 (2014).
Pawelec, G. Immunosenenescence: role of cytomegalovirus. Exp. Gerontol. 54, 1–5 (2014).
Khan, N. et al. Herpesvirus-specific CD8 T cell immunity in old age: cytomegalovirus impairs the response to a coresident EBV infection. J. Immunol. 173, 7481–7489 (2004).
Jackson, S. E. et al. CMV immune evasion and manipulation of the immune system with aging. Geroscience 39, 273–291 (2017).
Vallejo, A. N. et al. Expansions of NK-like αβT cells with chronologic aging: novel lymphocyte effectors that compensate for functional deficits of conventional NK cells and T cells. Ageing Res. Rev. 10, 354–361 (2011).
Coppé, J.-P. et al. Senescence-associated secretory phenotypes reveal cell-nonautonomous functions of oncogenic RAS and the p53 tumor suppressor. PLoS Biol. 6, 2853–2868 (2008).
Campisi, J. & di Fagagna, F. Cellular senescence: when bad things happen to good cells. Nat. Rev. Mol. Cell Biol. 8, 729–740 (2007).
Baker, D. J. et al. Naturally occurring p16Ink4a-positive cells shorten healthy lifespan. Nature 530, 184–189 (2016).
Baker, D. J. et al. Clearance of p16Ink4a-positive senescent cells delays ageing-associated disorders. Nature 479, 232–236 (2011).
Sagiv, A. et al. NKG2D ligands mediate immunosurveillance of senescent cells. Aging 8, 328–344 (2016).
Ucar, D et al. The chromatin accessibility signature of human immune aging stems from CD8+ T cells. J. Exp. Med. 214, 3123–3144 (2017).
Callender, L. A. et al. Human CD8+ EMRA T cells display a senescence-associated secretory phenotype regulated by p38 MAPK. Aging Cell 17, e12675 (2018).
Gérart, S. et al. Human iNKT and MAIT cells exhibit a PLZF-dependent proapoptotic propensity that is counterbalanced by XIAP. Blood 121, 614–623 (2013).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, 281–291.e9 (2019).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
McInnes, L., Healy, J. & Melville, J. UMAP: uniform manifold approximation and projection for dimension reduction. Preprint at aRxiv https://arxiv.org/abs/1802.03426v2 (2018).
Polański, K et al. Fast batch alignment of single cell transcriptomes unifies multiple mouse cell atlases into an integrated landscape. Bioinformatics 36, 964–965 (2020).
Allez, M. et al. CD4+NKG2D+ T cells in Crohn’s disease mediate inflammatory and cytotoxic responses through MICA interactions. Gastroenterology 132, 2346–2358 (2007).
Trivedi, S. G. et al. Essential role for hematopoietic prostaglandin D2 synthase in the control of delayed type hypersensitivity. Proc. Natl Acad. Sci. USA 103, 5179–5184 (2006).
Acknowledgements
We thank A. Toubert from INSERM U.1160 and Laboratoire d´Immunologie et d´Histocompatibilité, Hôpital Saint-Louis, Université Paris Diderot, Sorbonne Paris Cité for the kind gift of the C1R−MICA cell line. B.I.P. was supported by the Portuguese Foundation for Science and Technology and the Gulbenkian Institute for Science sponsoring the Advanced Medical Program for Physicians (PFMA). This work was supported by the Medical Research Council (grant MR/P00184X/1 to A.N.A.), the Ministry of Education of Brazil (grant BEX9414/14-2 to L.P.C.), the Wellcome Trust (grant AZR00630 to A. Lanna), University College London Business (to S.M.H. and A.N.A. for the microarray work), the National Institutes of Health (R01DK102850 and R01DK111465 to J.H.L.), the NIH/NIAID (R01 AG052608 and R01 AI142086 to J.B.) and the Biotechnology and Biological Science Research Council (grant BB/L005336/1 to N.E.R.). R.P.H.D was supported, in part, by the NIHR UCL Hospital Biomedical Research Centre, and S.M.H. is funded by the Springboard award from the Academy of Medical Science and the Wellcome Trust. A.L. is a Sir Henry Wellcome Trust Fellow sponsored by M. L. Dustin (University of Oxford). S.M.J. is a Wellcome Trust Senior Fellow in Clinical Science and is supported by the Rosetrees Trust, the Welton Trust, the Garfield Weston Trust and the UCLH Charitable Foundation. S.M.J. and V.H.T. have been funded by the Roy Castle Lung Cancer Foundation. D.U. is supported by the National Institute of General Medical Sciences (NIGMS) under award number GM124922. G.A.K. is supported by the Travelers Chair in Geriatrics and Gerontology, as well as the National Institute on Aging (AG061456, AG048023, AG063528, AG060746, AG021600, AG052608 and AG051647).
Author information
Authors and Affiliations
Contributions
B.I.P., L.P.C and R.P.H.D. designed and performed the experiments, analyzed the data and wrote the manuscript. D.N.-B. designed and analyzed the scRNA-seq data under the supervision of J.B. and D.U. R.M. performed all the experiments with the healthy older adult subjects. G.A.K. recruited all the healthy older adult donor subjects in Farmington, CT. A. Lanna, E.S.C. and N.E.R. designed and performed experiments. S.W. and J.S. designed and performed in vivo cytotoxicity studies. S.M.H. and A.N.A. designed and performed the microarray studies. A. Larbi provided support for studies of NKR on T cells. N.A.M., V.H.T. and S.M.J. analyzed the microarray and RNA-seq data. D.C.O.G., D.W.G., J.H.L. and M.K.M. facilitated mouse experiments. A.N.A. designed the experiments, reviewed and edited the manuscript, and organized the collaborative infrastructure.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Ioana Visan was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 CD8+ T cell gating and NKR expression.
a, Representative flow cytometry plots showing T cell gating and NKR expression on peripheral blood lymphocytes, specifically focusing on CD8+ T cell subsets stratified by the expression of CD27/CD45RA in healthy donors. Defined subsets are CD27+CD45RA+ TN, CD27+CD45RA− TCM, CD27−CD45RA− TEM, and CD27−CD45RA+ TEMRA cells. b, Confirmation of expression of CD57, KLRG1, CD244, NKG2D, NKG2C, and KIR2DL on TN (naive), TCM, TEM, and TEMRA CD8+ T cell subsets. Numbers in quadrants represent percentages of cells in each subset. Numbers above the histograms indicate the MFI. c, Flow cytometry gating of CD8+ T cells to confirm CD27 and CD28 expression in subpopulations based on CD27/CD45RA gating.
Extended Data Fig. 2 NKR expression in CD8+ and CD4+ T cells defined by CD27/CD28.
Expression of NK cell receptors (NKR) on a, CD8+ and b, CD4+ T cells assessed by flow cytometry on PBMCs from 22 healthy donors (median age = 52, range 25-83). Total CD8+ and CD4+ T cells were stratified into three subsets according to CD27/CD28 expression as shown in Extended Data Fig. 1a.
Extended Data Fig. 3 scRNA-seq method and quality control.
a, Overview of the scRNA-seq processing pipeline. Raw data (n=82,061 sorted CD8+ T cells) from six healthy older adult donors (six IL7R+ and six IL7R− CD8+ T cell samples) were first cleaned from the multiplets, using Scrublet36, then merged, resulting in a data set containing 62,343 cells. After batch correction using BBKNN37, the Scanpy66 -based pipeline was ran (see Methods section). b, Number of cells per individual (n=12). IL7R+ (n=6, in green) and IL7R- (n=6; in purple). c, Number of genes per distribution across the IL7R+ (in green) and IL7R- (in purple) cells. d, Number of cells before (light orange) and after (light blue) filtration (that is, doublet removal and other filtration steps that are described in Methods), within each individual. e, Bar plot highlighting the cell abundances across clusters (n = 13) for 10X run batches (upper panel) and IL7R+ and IL7R− groups (lower panel) after BBKNN batch effect correction. f, Bar plot highlighting the individual (n=12) cell abundances across clusters (n = 13) after BBKNN batch effect correction. Each color represents an individual. g, Number of cells in each cluster.
Extended Data Fig. 4 scRNA-seq comparison of re-clustered CD8+ T cells.
a, Violin plot showing the IL7R expression (as defined by scRNA-seq) across the 13 clusters. b, Dotplot showing the genes that are modulated in TN (top genes in red) and TEMRA (top genes in green) compartments. The scores (y axis) were defined using the Scanpy function (sc.tl.rank_genes_groups), based on Wilcoxon statistical test. FC = Fold change. TN (C0, C4 and C8) and TEMRA (C1, C2 and C6) compartments were extracted, a second round of clustering on the selected clusters (n = 39,634) was performed (as in Fig. 3) and UMAP plots highlighting c, IL7R groups (IL7R+ in green, IL7R− in purple, as defined by flow sorting) and d, of representative genes are shown.
Extended Data Fig. 5 Extended data on cytotoxicity and Sesn2 expression.
a, Titration curve of varying effector to target (E:T) ratios on cytotoxicity measured as specific lysis of K562 cells using a calcein-release of CD27−CD28−CD8+ (DN) T cells and NK cells isolated by FACS. Non-linear regression (5-parameter asymmetric) was performed (means and s.d., n = 3 donors). b, Calcein-release cytotoxicity assay of K562 cells by CD27+CD28+ (DP), CD27+CD28− (SP), DN CD8+ T cells, and NK cells at E:T 20:1. Cytotoxicity was assessed over a period of six hours (means and s.d., n = 3 donors). c, Representative dot plot of MICA/B expression in C1R and C1R–MICA*008 cells. d, Representative histogram of NKG2D express ion on CD28−CD8+ T cells after transfection with NK G2D siRNA (siNKG2D, black) or scrambled siRNA (siCtrl, gray), determined 36 hours after transfection. Numbers indicate MFI. e, Expression of DAP10 on human NK cells, and DP, SP, and DN CD8+ T cell subsets. Mean fluorescence intensity is shown (means and s.d., n = 4 for T cell, n = 3 NK cells). f, Sestrin 2 on CD8+ T cells from young (<35 years, n = 5) and old (>65 years, n = 4) donors. MFIs are shown (geometric means and geometric s.d. factor). Two-tailed, unpaired Welch’s t-test, ** p < 0.01.
Extended Data Fig. 7 Extended data on the murine delayed-type hypersensitivity model.
a, Spleen weight following mBSA-driven DTH response in young WT (Y WT, n = 4 mice), old WT (O WT, n = 8 mice), old Sesn1−/− (O Sesn1−/−, n = 5 mice), and old Sesn2−/− (O Sesn2−/−, n = 4 mice). Bars represent means and s.d.. b, Representative gating strategy to identify NK1.1+ NK cells (violet), TCRβ+CD1d tetramer reactive iNKT cells (purple), TCRβ+CD3+ CD4+ (blue) and CD8+ (red) T cells in mice. Similar results were obtained in all mice (n = 3 per group). c, Quantification of these cell types in the spleen (means and s.e.m., n = 3 mice per group). d, Dot plots showing relative frequencies of CD44−CD62L+ naive (gray), CD44+CD62L+ central (blue), and CD44+CD62L− effector (red) CD8+ T cells. e, Quantification of these cell types as a proportion of total splenic CD8+ T cells (means and s.e.m., n = 3 per group). f, Enumeration of NKG2D, NKG2A/C/E, KLRG1, Ly49, and NKp46 expression on CD8+ T cells from Y WT, O WT, O Sesn1−/−, and O Sesn2−/− mice (means and s.e.m., n = 3 per group). One-way ANOVA with Tukey’s multiple comparisons test. * p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001.
Supplementary information
Supplementary Table 1
Differentially expressed genes from microarray data.
Supplementary Tables 2–5
See tab “table legends” in file.
Source data
Source Data Fig. 1
Uncropped western blots.
Source Data Fig. 4
Uncropped western blots.
Source Data Fig. 5
Uncropped western blots.
Source Data Fig. 6
Uncropped western blots.
Rights and permissions
About this article
Cite this article
Pereira, B.I., De Maeyer, R.P.H., Covre, L.P. et al. Sestrins induce natural killer function in senescent-like CD8+ T cells. Nat Immunol 21, 684–694 (2020). https://doi.org/10.1038/s41590-020-0643-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-020-0643-3
This article is cited by
-
The senescence journey in cancer immunoediting
Molecular Cancer (2024)
-
Temporal trends, sex differences, and age-related disease influence in Neutrophil, Lymphocyte count and Neutrophil to Lymphocyte-ratio: results from InCHIANTI follow-up study
Immunity & Ageing (2023)
-
GoM DE: interpreting structure in sequence count data with differential expression analysis allowing for grades of membership
Genome Biology (2023)
-
Better understanding CD8+ T cells in cancer and viral infections
Nature Immunology (2023)
-
CISH impairs lysosomal function in activated T cells resulting in mitochondrial DNA release and inflammaging
Nature Aging (2023)