Abstract
Low exposure to microbial products, respiratory viral infections and air pollution are major risk factors for allergic asthma, yet the mechanistic links between such conditions and host susceptibility to type 2 allergic disorders remain unclear. Through the use of single-cell RNA sequencing, we characterized lung neutrophils in mice exposed to a pro-allergic low dose of lipopolysaccharide (LPS) or a protective high dose of LPS before exposure to house dust mites. Unlike exposure to a high dose of LPS, exposure to a low dose of LPS instructed recruited neutrophils to upregulate their expression of the chemokine receptor CXCR4 and to release neutrophil extracellular traps. Low-dose LPS–induced neutrophils and neutrophil extracellular traps potentiated the uptake of house dust mites by CD11b+Ly-6C+ dendritic cells and type 2 allergic airway inflammation in response to house dust mites. Neutrophil extracellular traps derived from CXCR4hi neutrophils were also needed to mediate allergic asthma triggered by infection with influenza virus or exposure to ozone. Our study indicates that apparently unrelated environmental risk factors can shape recruited lung neutrophils to promote the initiation of allergic asthma.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The scRNA-seq data provided in this manuscript have been deposited in the ArrayExpress database at EMBL-EBI (https://www.ebi.ac.uk/arrayexpress/ experiments/E-MTAB-6902) under accession number E-MTAB-6902. The figures that have associated scRNA-seq data are the following: Fig. 2, Supplementary Figs. 2 and 3. The source data underlying Fig. 1a–d,f, 2d, 3b,c,f,g,i,k, 4b–e,g,i–k,m, 5b–e,g, 6b,d,f, 7b–i,k–m and 8a–c,e,f–h,j and Supplementary Figs. 1b,d, 4c,d, 5b,c,e,g, 6b, 7b–d,f,g,i,j and 8a are provided as a Source Data file. Any data that support the findings of this study are available from the corresponding authors upon request.
References
Braman, S. S. The global burden of asthma. Chest 130, 4s–12s (2006).
Galli, S. J., Tsai, M. & Piliponsky, A. M. The development of allergic inflammation. Nature 454, 445–454 (2008).
Vercelli, D. Discovering susceptibility genes for asthma and allergy. Nat. Rev. Immunol. 8, 169–182 (2008).
Eder, W., Ege, M. J. & von Mutius, E. The asthma epidemic. N. Engl. J. Med. 355, 2226–2235 (2006).
Platts-Mills, T. A. The allergy epidemics: 1870–2010. J. Allergy Clin. Immunol. 136, 3–13 (2015).
Busse, W. W., Lemanske, R. F. Jr. & Gern, J. E. Role of viral respiratory infections in asthma and asthma exacerbations. Lancet 376, 826–834 (2010).
Peden, D. B. The epidemiology and genetics of asthma risk associated with air pollution. J. Allergy Clin. Immunol. 115, 213–219 (2005). quiz 220.
Braun-Fahrlander, C. et al. Environmental exposure to endotoxin and its relation to asthma in school-age children. N. Engl. J. Med. 347, 869–877 (2002).
Strachan, D. P. Family size, infection and atopy: the first decade of the ‘hygiene hypothesis’. Thorax 55, S2–S10 (2000).
Lloyd, C. M. & Snelgrove, R. J. Type 2 immunity: expanding our view. Sci. Immunol. 3, eaat1604 (2018).
Lambrecht, B. N. & Hammad, H. The immunology of asthma. Nat. Immunol. 16, 45–56 (2015).
Gregory, L. G. & Lloyd, C. M. Orchestrating house dust mite-associated allergy in the lung. Trends Immunol. 32, 402–411 (2011).
Locksley, R. M. Asthma and allergic inflammation. Cell 140, 777–783 (2010).
Ng, L. G., Ostuni, R. & Hidalgo, A. Heterogeneity of neutrophils. Nat. Rev. Immunol. 19, 255–265 (2019).
Papayannopoulos, V. Neutrophil extracellular traps in immunity and disease. Nat. Rev. Immunol. 18, 134–147 (2018).
Krishnamoorthy, N. et al. Neutrophil cytoplasts induce TH17 differentiation and skew inflammation toward neutrophilia in severe asthma. Sci. Immunol. 3, eaao4747 (2018).
Radermecker, C., Louis, R., Bureau, F. & Marichal, T. Role of neutrophils in allergic asthma. Curr. Opin. Immunol. 54, 28–34 (2018).
Wang, Y. H. & Wills-Karp, M. The potential role of interleukin-17 in severe asthma. Curr. Allergy Asthma Rep. 11, 388–394 (2011).
Lachowicz-Scroggins, M. E. et al. Extracellular DNA, neutrophil extracellular traps, and inflammasome activation in severe asthma. Am. J. Respir. Crit. Care Med. 199, 1076–1085 (2019).
Toussaint, M. et al. Host DNA released by NETosis promotes rhinovirus-induced type-2 allergic asthma exacerbation. Nat. Med. 23, 681–691 (2017).
Eisenbarth, S. C. et al. Lipopolysaccharide-enhanced, toll-like receptor 4-dependent T helper cell type 2 responses to inhaled antigen. J. Exp. Med. 196, 1645–1651 (2002).
Schuijs, M. J. et al. Farm dust and endotoxin protect against allergy through A20 induction in lung epithelial cells. Science 349, 1106–1110 (2015).
Zheng, G. X. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Remijsen, Q. et al. Neutrophil extracellular trap cell death requires both autophagy and superoxide generation. Cell Res. 21, 290–304 (2011).
Adrover, J. M., Nicolas-Avila, J. A. & Hidalgo, A. Aging: a temporal dimension for neutrophils. Trends Immunol. 37, 334–345 (2016).
Casanova-Acebes, M. et al. Rhythmic modulation of the hematopoietic niche through neutrophil clearance. Cell 153, 1025–1035 (2013).
Dwyer, M. P. et al. Discovery of 2-hydroxy-N,N-dimethyl-3-{2-[[(R)-1-(5- methylfuran-2-yl)propyl]amino]-3,4-dioxocyclobut-1-enylamino}benzamide (SCH527123): a potent, orally bioavailable CXCR2/CXCR1 receptor antagonist. J. Med. Chem. 49, 7603–7606 (2006).
Reutershan, J. et al. Critical role of endothelial CXCR2 in LPS-induced neutrophil migration into the lung. J. Clin. Invest. 116, 695–702 (2006).
Cools-Lartigue, J. et al. Neutrophil extracellular traps sequester circulating tumor cells and promote metastasis. J. Clin. Invest. 123, 3446–3458 (2013).
Wang, Y. et al. Histone hypercitrullination mediates chromatin decondensation and neutrophil extracellular trap formation. J. Cell Biol. 184, 205–213 (2009).
Mesnil, C. et al. Resident CD11b+Ly6C– lung dendritic cells are responsible for allergic airway sensitization to house dust mite in mice. PloS ONE 7, e53242 (2012).
Plantinga, M. et al. Conventional and monocyte-derived CD11b+ dendritic cells initiate and maintain T helper 2 cell-mediated immunity to house dust mite allergen. Immunity 38, 322–335 (2013).
Marichal, T. et al. Interferon response factor 3 is essential for house dust mite-induced airway allergy. J. Allergy Clin. Immunol. 126, 836–844.e13 (2010).
Larché, M. et al. Costimulation through CD86 is involved in airway antigen-presenting cell and T cell responses to allergen in atopic asthmatics. J. Immunol. 161, 6375–6382 (1998).
Janss, T. et al. Interferon response factor-3 promotes the pro-Th2 activity of mouse lung CD11b+ conventional dendritic cells in response to house dust mite allergens. Eur. J. Immunol. 46, 2614–2628 (2016).
Al-Garawi, A. A. et al. Acute, but not resolved, influenza A infection enhances susceptibility to house dust mite-induced allergic disease. J. Immunol. 182, 3095–3104 (2009).
Hollingsworth, J. W. et al. Ozone activates pulmonary dendritic cells and promotes allergic sensitization through a Toll-like receptor 4-dependent mechanism. J. Allergy Clin. Immunol. 125, 1167–1170 (2010).
Stein, M. M. et al. Innate immunity and asthma risk in amish and hutterite farm children. N. Engl. J. Med. 375, 411–421 (2016).
Roan, F., Obata-Ninomiya, K. & Ziegler, S. F. Epithelial cell-derived cytokines: more than just signaling the alarm. J. Clin. Invest. 129, 1441–1451 (2019).
Marichal, T. et al. DNA released from dying host cells mediates aluminum adjuvant activity. Nat. Med. 17, 996–1002 (2011).
Bonnelykke, K., Vissing, N. H., Sevelsted, A., Johnston, S. L. & Bisgaard, H. Association between respiratory infections in early life and later asthma is independent of virus type. J. Allergy Clin. Immunol. 136, 81–86.e4 (2015).
Sabatel, C. et al. Exposure to bacterial CpG DNA protects from airway allergic inflammation by expanding regulatory lung interstitial macrophages. Immunity 46, 457–473 (2017).
Gentleman, R. C. et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 5, R80 (2004).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Acknowledgments
We thank F. Trottein (Pasteur Institute, Lille, France) for providing us with the Influenza A virus H1N1 PR8/34 strain; S. Ormenese, R. Stefan, J.-J. Goval and A. Hego from the GIGA Flow Cytometry and Cell Imaging Platform; P. Drion and all staff members from the GIGA Mouse facility and Transgenics Platform; C. Humblet and staff members from the GIGA Immunohistology Platform; B. Charloteaux, M. Deckers, L. Karim and members from the GIGA Genomics Platform; C. Desmet and D. Pirottin for insightful discussions; and C. François, R. Fares and I. Sbai for their excellent technical and administrative support. C.Radermecker and C.S. were research fellows of the F.R.S-FNRS; S.L.J. was supported by a Chair from Asthma UK (no. CH11SJ, to S.L.J.) and Medical Research Council Center grant (no. G1000758, to S.L.J.); F.B. is supported by the F.R.S.-FNRS for the FRFS-WELBIO (grant no. CR-2012S-01R to F.B.) and by an Excellence Of Science (EOS) grant. T.M. is a Research Associate of the F.R.S.-FNRS and is supported by an ‘Incentive Grant for Scientific Research’ of the F.R.S.-FNRS (grant no. F.4508.18 to T.M.), by the FRFS-WELBIO (grant no. CR-2017s-04 to T.M.), by the Acteria Foundation and by an ERC Starting Grant (grant no. ERC-StG-2018 IM-ID 801823 to T.M.).
Author information
Authors and Affiliations
Contributions
C.Radermecker, T.M. and F.B. conceived the project. T.M., F.B., M.T., C.Radermecker and C.S. were involved in experimental design. C.Radermecker and C.S. realized most experiments, compiled the data and contributed equally to this work. P.M., C.Ruscitti and J.S. were involved in (bone marrow-derived) dendritic cell-related experiments. M.T. and S.L.J. were involved in experiments aiming at detecting and inhibiting NETs. C.V., F.P., N.R. and D.C. contributed to experiments involving ozone exposure and invasive measurements of airway function. T.M. analyzed scRNA-seq data with the help of the GIGA Genomics Platform. C.Radermecker and T.M. prepared the figures, and T.M. wrote the manuscript. All authors provided feedback on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Ioana Visan was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 BALB/c wild-type mice administered i.n. with different LPS doses exhibit distinct profiles of innate neutrophil recruitment and HDM-induced airway eosinophilia.
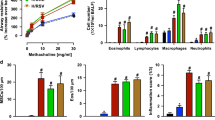
a, Experimental outline for panel b. b, Eosinophil cell counts in the broncho-alveolar lavage fluid (BALF), assessed 3 d after the second HDM administration in mice exposed to different doses of LPS, ranging from 0.1 ng to 10 µg and instilled i.n. with 40 and 10 µg HDM, 1 and 8 d later, respectively. Data shown are mean +/- s.e.m and are pooled from 4 independent experiments (PBS; LPS 0.1 ng; 1 ng; 10 ng; 100 ng; 1 µg; 10 µg: n = 9;16;22;20;23;24;25 mice per group, respectively). c, Experimental outline for panel d. d, Absolute numbers of lung CD45+CD11bhiLy-6Ghi neutrophils assessed over time after i.n. vehicle, low-dose LPS (LPSlo) or high-dose LPS (LPShi) exposure. Data show mean +/- s.e.m. and are pooled from 2 independent experiments (n = 3 [PBS, LPSlo] or 4 [LPShi] mice per group). P values were calculated using (b) a one-way or (d) a two-way ANOVA with Tukey’s post hoc test. In d, the symbol * compares the indicated group vs. PBS; the symbol ° compares LPSlo vs. LPShi groups. °P<0.05; **/°°P<0. 01; ***P<0.001. i.n., intranasal.
Supplementary Figure 2 Lung neutrophils from vehicle, LPSlo and LPShi mice were isolated and subjected to droplet-based scRNA-seq and quality-control filtering and analysis.
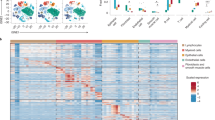
a, Simplified experimental outline for scRNA-seq analysis. b, Representative FACS sorting strategy of CD45+ cells after MACS neutrophil enrichment of lung single-cell suspensions (n = 3 pooled mice/group). One PBS-injected mouse is shown. c, Post-sort purity of FACS-sorted CD45+ cells from vehicle, LPSlo and LPShi mice 24 h after treatment. Representative post-sort flow cytometry dot plots are shown. Insets indicate the percentage of cells within the indicated gate. d,e, Gene numbers (left), Unique Molecular Identifiers (UMI) numbers (middle) and percentage of mitochondrial genes (right) detected in vehicle, LPSlo and LPShi neutrophils before (d) and after (e) selection and filtering, presented as violin plots (height: gene numbers [left], UMI numbers [middle] and percentage mitochondrial genes [right]; width: abundance of cells) and individual dots representing individual cells. The red frames indicate cells that were selected for downstream analyses. f, tSNE plots depicting the transcriptional identity of neutrophils merged from vehicle, LPSlo and LPShi mice 24 h after treatment, analyzed by scRNA-seq. g, Dot plots showing average expression of the indicated genes and percentage of cells expressing the indicated genes within each cluster. High expression of S100a8, S100a9 and Csf3r identified the 6 clusters as neutrophils, which was further supported by the virtual absence of expression of markers of other cell types.
Supplementary Figure 3 Clusters identified by scRNA-seq were analyzed for differential gene expression.
a, Simplified experimental outline for scRNA-seq analysis. b, tSNE plots depicting the transcriptional identity of neutrophils merged from vehicle, LPSlo and LPShi mice 24 h after treatment, analyzed by scRNA-seq (n = 3 pooled mice/condition). c, PANTHER GO statistical enrichment tests on the genes significantly upregulated in cluster 0 as compared to the other clusters. d, Dot plots showing average expression of the indicated genes and percentage of cells expressing the indicated genes within neutrophil clusters. Depicted genes belong to the lists of genes characterizing the biological responses shown in c. e, PANTHER GO statistical enrichment tests on the common LPS signature. f, Dot plots, as in d. Depicted genes belong to the lists of genes characterizing the biological responses shown in e. (c,e) Number of genes (histograms) and -log P value (gray squares) of the enriched biological processes are plotted. P values were calculated using a two-tailed Mann-Whitney U test with Benjamini-Hochberg False Discovery Rate (FDR) correction.
Supplementary Figure 4 LPSlo neutrophils, unlike steady-state or LPShi neutrophils, are dedicated to form NETs ex vivo.
a, Experimental outline. b, Representative photographs of FACS-sorted CXCR4loCD49dlo, CXCR4loCD49dlo and CXCR4hiCD49dhi lung neutrophils from vehicle, LPSlo and LPShi mice 24 h after treatment. c, Quantification of the size of neutrophils from mice as in b. d, Quantification of the nucleus/cytoplasm ratio of neutrophils from mice as in b. e-f, Representative high-resolution confocal microscopy pictures of Cit-H3+MPO+DAPI+ NETs released from ex vivo-cultured neutrophils as in b. e, Representative pictures of the individual and merged stainings, magnification 20x. f, Additional representative pictures of the merged stainings, magnification 10x. g, Quantification of NET volume in ex vivo-cultured neutrophils as in b. Data show mean + s.e.m., each symbol representing independent sorting experiments in which cells from 2-3 mice were pooled by group (c-d,g: n = 6,4 independent experiments, respectively). P values were calculated using a one-way ANOVA with Tukey’s post hoc test. **P<0.01; ***P<0.001. i.n., intranasal. ns, not significant. Scale bar = (b) 10 µm; (e,f) 50 µm.
Supplementary Figure 5 NETs are not detected in LPSlo mice treated with α-Ly-6G neutrophil-depleting antibodies.
a, Experimental outline. b, Neutrophil cell counts in the broncho-alveolar lavage fluid (BALF) of vehicle and LPSlo mice treated or not i.p. with α-Ly-6G antibodies one day before, at the time of and one day after LPSlo exposure. c, Levels of extracellular dsDNA in the BALF of mice as in b. d, Representative blots of Cit-H3 and HSP90α (loading control) assessed by Western blot of lung protein extracts from mice as in b. e, Quantification of normalized Cit-H3 levels in lung protein extracts of mice as in b. f, Confocal microscopy stainings of Cit-H3+MPO+ NETs on lung sections of mice as in b. Pictures shown are representative of one of >3 mice analyzed, each of them giving similar results. g, Quantification of NET volume in lung sections of mice as in b. (b,c,e,g) Data show mean + s.e.m., as well as individual mice, and (b,c,e) are representative of one of 3 independent experiments yielding similar results (b,c,e: n = 3 mice per group; g: n = 4 mice per group). P values were calculated using a one-way ANOVA with Tukey’s post hoc test. ***P<0.001. BALF, bronchoalveolar lavage fluid; i.p., intraperitoneal; ns, not significant. Scale bar = 50 µm.
Supplementary Figure 6 NETs appear in the lung 18 h after LPSlo exposure and are impaired in lungs of LPSlo mice treated with DNAse, Cl-amidine or NEi.
a, Confocal microscopy stainings of Cit-H3+MPO+ NETs on lung sections of mice 0, 6, 12, 18, 24 and 48 h after LPSlo treatment. Pictures are representative of one of 4 mice analyzed, each of them giving similar results. b, Quantification of NET volume on lung sections of mice as in a. c, Confocal microscopy stainings of Cit-H3+MPO+ NETs on lung sections, assessed 24 h after LPSlo in mice treated i.p. with DNAse one day before and at the time of LPSlo treatment, or treated 4 times i.p. with NEi or Cl-amidine, every 12 h starting one day before LPSlo. Pictures are representative of one of 5 mice analyzed, each of them giving similar results. Quantification of NET volume is shown in Fig. 5b. (b) Data show mean + s.e.m., as well as individual mice (n = 4 per time point). P values were calculated using a one-way ANOVA with Tukey’s post hoc tests and compare the indicated time points with the baseline (hour 0). ***P<0.001. NEi, neutrophil elastase inhibitor. Scale bar = 50 µm.
Supplementary Figure 7 NET-releasing neutrophils are recruited to the lungs on influenza virus PR8 infection and ozone exposure.
a, Experimental outline for panels b–d and g-k. b, Body weight expressed as the percentage of body weight before influenza virus (H1N1 strain PR8) infection, assessed over time in PBS-injected and PR8-infected mice. c, Lung relative RNA expression of viral non-structural protein 1 (NS1) assessed by RT-qPCR over time in PBS-injected or PR8-infected mice. d, Absolute numbers of lung CD45+CD11bhiLy-6Ghi neutrophils assessed over time in PBS-injected or PR8-infected mice. e, Experimental outline for f. f, Absolute numbers of lung CD45+CD11bhiLy-6Ghi neutrophils assessed in air-exposed and ozone-exposed mice 1 d after the last ozone exposure. g, Levels of extracellular dsDNA in the BALF of mice following PR8 infection. h, Representative blots of Cit-H3 and HSP90α (loading control) assessed by Western blot of lung protein extracts of mice following PR8 infection. i, Quantification of normalized Cit-H3 levels in lung protein extracts of mice following PR8 infection. j, Quantification of NET volume in lung sections of mice following PR8 infection. k, Confocal microscopy stainings of Cit-H3+MPO+ NETs on lung sections of mice following PR8 infection. Pictures shown are representative of one of 4 mice analyzed, each of them giving similar results. (b,c) Data show mean +/- s.e.m. and are pooled from 4 independent experiments, each symbol representing individual mice, ([b] PBS: n = 20, 20, 20, 18, 19, 20 at days 0, 1, 3, 5, 7, 10, respectively; PR8: n=20,20,20,18,20,10 at days 0,1,3,5,7,10, respectively; [c] PBS: n=8 at each time point; PR8: n=8,8,13,13,12,8 at days 0,1,3,5,7,10 respectively). (d,f) Data show mean (d) +/- or (f) + s.e.m. and are pooled from 2 independent experiments, each symbol representing individual mice (n = 3-4 mice per group). (g,i,j) Data show mean +/- s.e.m. and are pooled from 3-5 independent experiments, each symbol representing (g,j) individual mice (g: n = 9,7,16,17,16,7 at days 0,1,3,5,7,10, respectively; j: n=4 mice per time point), or (i) independent experiments in which lungs from 3-5 mice were pooled by group (n = 4 independent experiments). P values were calculated using (b,c,d) a two-way ANOVA with Tukey’s post hoc test and compare PBS vs. PR8 at the indicated time points, (f) a two-tailed unpaired Student’s t test or (g,i,j) a one-way ANOVA with Tukey’s post hoc test and compare the indicated time points with the baseline. *P < 0.05; **P < 0.01; ***P < 0.001. BALF, bronchoalveolar lavage fluid; ns, not significant i.n., intranasal; PFU, plaque forming unit. Scale bar = 50 µm.
Supplementary Figure 8 Ozone-potentiated HDM-induced perivascular and peribronchial leukocyte infiltration is decreased upon treatment with NEi, Cl-Amidine or DNAse.
a, Inflammatory score estimating perivascular and peribronchial inflammation, quantified from H&E-stained lung sections 3 d after the second HDM administration in vehicle-HDM and ozone-HDM mice treated daily for 4 d with DNAse i.p., starting the first day of ozone exposure, or treated every 12 h with NEi or Cl-amidine i.p., for the same duration as for DNAse. b, Representative H&E staining of lung sections of mice as in a. Pictures are representative of one of 6 mice analyzed, each of them giving comparable results. (a) Data show mean + s.e.m. and are pooled from 2 independent experiments (n = 6 mice per group). P values were calculated using a one-way ANOVA with Tukey’s post hoc test. **P < 0.01; ***P<0.001. NEi, neutrophil elastase inhibitor. Scale bar = 32 µm.
Supplementary information
Supplementary Information
Supplementary Figs. 1–8.
Source data
Rights and permissions
About this article
Cite this article
Radermecker, C., Sabatel, C., Vanwinge, C. et al. Locally instructed CXCR4hi neutrophils trigger environment-driven allergic asthma through the release of neutrophil extracellular traps. Nat Immunol 20, 1444–1455 (2019). https://doi.org/10.1038/s41590-019-0496-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-019-0496-9