Abstract
Progenitor-like CD8+ T cells mediate long-term immunity to chronic infection and cancer and respond potently to immune checkpoint blockade. These cells share transcriptional regulators with memory precursor cells, including T cell-specific transcription factor 1 (TCF1), but it is unclear whether they adopt distinct programs to adapt to the immunosuppressive environment. By comparing the single-cell transcriptomes and epigenetic profiles of CD8+ T cells responding to acute and chronic viral infections, we found that progenitor-like CD8+ T cells became distinct from memory precursor cells before the peak of the T cell response. We discovered a coexpression gene module containing Tox that exhibited higher transcriptional activity associated with more abundant active histone marks in progenitor-like cells than memory precursor cells. Moreover, thymocyte selection-associated high mobility group box protein TOX (TOX) promoted the persistence of antiviral CD8+ T cells and was required for the programming of progenitor-like CD8+ T cells. Thus, long-term CD8+ T cell immunity to chronic viral infection requires unique transcriptional and epigenetic programs associated with the transcription factor TOX.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All data generated during this study are available within the paper. The RNA-seq, ChIP-seq and scRNA-seq data have been deposited with the Gene Expression Omnibus (GEO, accession no. GSE119943). The TOX, NFAT1, IRF4, RUNX3, FOXO1, STAT3 and STAT5B ChIP-seq data are publicly available (GEO accession nos. GSE93953, GSE64407, GSE49930, GSE46943, GSE50128 and GSE102317).
References
Ahmed, R. & Gray, D. Immunological memory and protective immunity: understanding their relation. Science 272, 54–60 (1996).
Hashimoto, M. et al. CD8 T cell exhaustion in chronic infection and cancer: opportunities for interventions. Annu. Rev. Med. 69, 301–318 (2018).
Thommen, D. S. & Schumacher, T. N. T cell dysfunction in cancer. Cancer Cell 33, 547–562 (2018).
Kaech, S. M. & Wherry, E. J. Heterogeneity and cell-fate decisions in effector and memory CD8+ T cell differentiation during viral infection. Immunity 27, 393–405 (2007).
Blackburn, S. D., Shin, H., Freeman, G. J. & Wherry, E. J. Selective expansion of a subset of exhausted CD8 T cells by αPD-L1 blockade. Proc. Natl Acad. Sci. USA 105, 15016–15021 (2008).
Wu, T. et al. The TCF1-Bcl6 axis counteracts type I interferon to repress exhaustion and maintain T cell stemness. Sci. Immunol. 1, eaai8593 (2016).
Im, S. J. et al. Defining CD8+ T cells that provide the proliferative burst after PD-1 therapy. Nature 537, 417–421 (2016).
He, R. et al. Follicular CXCR5-expressing CD8+ T cells curtail chronic viral infection. Nature 537, 412–428 (2016).
Utzschneider, D. T. et al. T cell factor 1-expressing memory-like CD8+ T cells sustain the immune response to chronic viral infections. Immunity 45, 415–427 (2016).
Leong, Y. A. et al. CXCR5+ follicular cytotoxic T cells control viral infection in B cell follicles. Nat. Immunol. 17, 1187–1196 (2016).
Siddiqui, I. et al. Intratumoral Tcf1+PD-1+CD8+ T cells with stem-like properties promote tumor control in response to vaccination and checkpoint blockade immunotherapy. Immunity 50, 195–211.e10 (2019).
Man, K. et al. Transcription factor IRF4 promotes CD8+ T cell exhaustion and limits the development of memory-like T cells during chronic infection. Immunity 47, 1129–1141.e5 (2017).
Doering, T. A. et al. Network analysis reveals centrally connected genes and pathways involved in CD8+ T cell exhaustion versus memory. Immunity 37, 1130–1144 (2012).
Papalexi, E. & Satija, R. Single-cell RNA sequencing to explore immune cell heterogeneity. Nat. Rev. Immunol. 18, 35–45 (2018).
O’Flaherty, E. & Kaye, J. TOX defines a conserved subfamily of HMG-box proteins. BMC Genomics 4, 13 (2003).
Pircher, H., Bürki, K., Lang, R., Hengartner, H. & Zinkernagel, R. M. Tolerance induction in double specific T-cell receptor transgenic mice varies with antigen. Nature 342, 559–561 (1989).
Ahmed, R., Salmi, A., Butler, L. D., Chiller, J. M. & Oldstone, M. B. Selection of genetic variants of lymphocytic choriomeningitis virus in spleens of persistently infected mice. Role in suppression of cytotoxic T lymphocyte response and viral persistence. J. Exp. Med. 160, 521–540 (1984).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Singer, M. et al. A distinct gene module for dysfunction uncoupled from activation in tumor-infiltrating T cells. Cell 171, 1221–1223 (2017).
Aliahmad, P. & Kaye, J. Development of all CD4 T lineages requires nuclear factor TOX. J. Exp. Med. 205, 245–256 (2008).
Seehus, C. R. et al. The development of innate lymphoid cells requires TOX-dependent generation of a common innate lymphoid cell progenitor. Nat. Immunol. 16, 599–608 (2015).
Page, N. et al. Expression of the DNA-binding factor TOX promotes the encephalitogenic potential of microbe-induced autoreactive CD8+ T cells. Immunity 48, 937–950.e8 (2018).
Youngblood, B. et al. Effector CD8 T cells dedifferentiate into long-lived memory cells. Nature 552, 404–409 (2017).
Gray, S. M., Amezquita, R. A., Guan, T., Kleinstein, S. H. & Kaech, S. M. Polycomb repressive complex 2-mediated chromatin repression guides effector CD8+ T cell terminal differentiation and loss of multipotency. Immunity 46, 596–608 (2017).
He, B. et al. CD8+ T cells utilize highly dynamic enhancer repertoires and regulatory circuitry in response to infections. Immunity 45, 1341–1354 (2016).
Creyghton, M. P. et al. Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc. Natl Acad. Sci. USA 107, 21931–21936 (2010).
Wu, T. et al. TCF1 is required for the T follicular helper cell response to viral infection. Cell Rep. 12, 2099–2110 (2015).
Martinez, G. J. et al. The transcription factor NFAT promotes exhaustion of activated CD8+ T cells. Immunity 42, 265–278 (2015).
Shan, Q. et al. The transcription factor Runx3 guards cytotoxic CD8+ effector T cells against deviation towards follicular helper T cell lineage. Nat. Immunol. 18, 931–939 (2017).
Staron, M. M. et al. The transcription factor FoxO1 sustains expression of the inhibitory receptor PD-1 and survival of antiviral CD8+ T cells during chronic infection. Immunity 41, 802–814 (2014).
Cox, M. A., Kahan, S. M. & Zajac, A. J. Anti-viral CD8 T cells and the cytokines that they love. Virology 435, 157–169 (2013).
Xin, G. et al. A critical role of IL-21-induced BATF in sustaining CD8-T-cell-mediated chronic viral control. Cell Rep. 13, 1118–1124 (2015).
Doedens, A. L. et al. Hypoxia-inducible factors enhance the effector responses of CD8+ T cells to persistent antigen. Nat. Immunol. 14, 1173–1182 (2013).
Gattinoni, L. et al. Wnt signaling arrests effector T cell differentiation and generates CD8+ memory stem cells. Nat. Med. 15, 808–813 (2009).
Miller, B. C. et al. Subsets of exhausted CD8+ T cells differentially mediate tumor control and respond to checkpoint blockade. Nat. Immunol. 20, 326–336 (2019).
Sade-Feldman, M. et al. Defining T cell states associated with response to checkpoint immunotherapy in melanoma. Cell 175, 998–1013.e20 (2018).
Kurtulus, S. et al. Checkpoint blockade immunotherapy induces dynamic changes in PD-1−CD8+ tumor-infiltrating T cells. Immunity 50, 181–194.e6 (2019).
Snell, L. M. et al. CD8+ T cell priming in established chronic viral infection preferentially directs differentiation of memory-like cells for sustained immunity. Immunity 49, 678–694.e5 (2018).
Wherry, E. J. & Kurachi, M. Molecular and cellular insights into T cell exhaustion. Nat. Rev. Immunol. 15, 486–499 (2015).
Gattinoni, L., Speiser, D. E., Lichterfeld, M. & Bonini, C. T memory stem cells in health and disease. Nat. Med. 23, 18–27 (2017).
Gautam, S. et al. The transcription factor c-Myb regulates CD8+ T cell stemness and antitumor immunity. Nat. Immunol. 20, 337–349 (2019).
Rosenberg, S. A. & Restifo, N. P. Adoptive cell transfer as personalized immunotherapy for human cancer. Science 348, 62–68 (2015).
Fooksman, D. R., Nussenzweig, M. C. & Dustin, M. L. Myeloid cells limit production of antibody-secreting cells after immunization in the lymph node. J. Immunol. 192, 1004–1012 (2014).
Araki, K. et al. Pathogenic virus-specific T cells cause disease during treatment with the calcineurin inhibitor FK506: implications for transplantation. J. Exp. Med. 207, 2355–2367 (2010).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9, 559 (2008).
Schmidl, C., Rendeiro, A. F., Sheffield, N. C. & Bock, C. ChIPmentation: fast, robust, low-input ChIP-seq for histones and transcription factors. Nat. Methods 12, 963–965 (2015).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Trapnell, C. & Salzberg, S. L. How to map billions of short reads onto genomes. Nat. Biotechnol. 27, 455–457 (2009).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Acknowledgements
We thank L. Garrett (NHGRI), E. Escobar (NHGRI), C. Rivas (NHGRI), I. Ginty (NHGRI), W. Pridgen (NHGRI), G. Gutierrez-Cruz (National Institute of Arthritis and Musculoskeletal and Skin Diseases (NIAMS)) and S. Dell’Orso (NIAMS) for their excellent technical support and J. Kaye for the ToxloxP/loxP mice. This work was supported in part by the intramural programs of the NHGRI, National Institute of Allergy and Infectious Diseases, NIAMS and National Institute of Neurological Disorders and Stroke, and the National Cancer Institute, National Institutes of Health (NIH) and DDIR Innovation Awards to P.L.S., J.J.O. and L.G., and NIH grant (no. AG056524 to T.W). E.J.W. is a member of the Parker Institute for Cancer Immunotherapy, which supported the University of Pennsylvania cancer immunotherapy program.
Author information
Authors and Affiliations
Contributions
T.W. conceived the study. T.W., C.Y., P.L.S. and J.J.O. designed the experiments. T.W., C.Y., Y.J., E.A.M., H.-Y.S., N.E.L., M.K., S.A., J.C., J.F., J.H., R.H., J.R. and S.G. performed the experiments. O.K. and E.J.W. provided the P14 ToxloxP/loxP CD4-Cre mice. T.W., C.Y., H.-W.S., P.L.S., J.J.O., D.B.M., L.G. and E.F.H. analyzed and interpreted the results. T.W., C.Y. and P.L.S. drafted the manuscript.
Corresponding authors
Ethics declarations
Competing interests
E.J.W. has consulting agreements with and/or is on the scientific advisory board for Merck, Roche, Pieris, Elstar and Surface Oncology. E.J.W. has a patent licensing agreement on the PD-1 pathway with Roche/Genentech. Y.J. has stock in Cellular Biomedicine Group.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 scRNA-seq analysis of virus-specific CD8+ T cells from chronic LCMV infection.
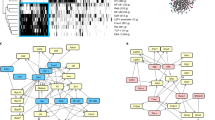
Experimental setup has been described in Fig. 1. (a) Gating strategy for sorting P14 CD8+ T cells (CD45.1+ CD8+) on day 7 post-infection. (b) Percentages of cells that belong to each cluster, as defined in Fig. 1a. (c) Violin plots showing the expression of Tcf7, Id3, Tox, Ikzf2, and Hif1a in cells from cluster 0 through cluster 3, as defined in Fig. 1a. The violin represents the probability density at each value; each dot represents one cell (n = 2,597 cells). (d) Violin plots showing the expression of Gzmb, Id2, Birc5, Top2a, and Ezh2 in cells from cluster 0 through cluster 3. The violin represents the probability density at each value; each dot represents one cell (n = 2,597 cells). (e) Left panel: Histogram of the Area Under the Curve (AUC) score, determined by AUCell. The red vertical line separates the cells with progenitor-like gene set “on” from ones with the gene set “off”. Right panel: t-SNE plots indicate whether progenitor-like gene set is “on” (red) or “off” (grey) in each cell. Sample size n = 2,597 cells. (f) X-axis represents values of component 2, determined by Monocle 2. Y-axis represents the normalized expression of Tcf7 in each cell (n = 2,597 cells). Dark blue circles represent the spline. (g) The cell-cycle phase (G1, S, or G2/M) of each cell (n = 2,597 cells) was determined by Seurat v.2. The distribution of the values of component 1, determined by Monocle 2, in cells from each cell-cycle phase is plotted as histogram (G1: red; S: blue; G2/M: green). (h) Left panel: Cell-cycle phase (G1: red; S: blue; G2/M: green) of each cell (n = 2,597 cells) is shown in t-SNE plots. Right panel: Percentages of cells from each cluster, defined in Fig. 1a, in each cell-cycle phase (G1: red; S: blue; G2/M: green).
Supplementary Figure 2 scRNA-seq analysis of virus-specific CD8+ T cells from acute and chronic LCMV infections.
Experimental setup has been described in Fig. 2. (a) t-SNE plots of scRNA-seq data merged from two independent batches of D4.5 Arm, D4.5 Cl13, D7 Arm and D7 Cl13 P14 CD8+ T cells. Batch#1: n = 19,497 cells; and batch#2: n = 16,042 cells. (b) Correlation of gene expression (27,998 genes) between scRNA-seq replicates (upper panel) and between scRNA-seq data and bulk RNA-seq data (lower panel) with Pearson’s R indicated. Numbers on the axes are in TPM+1. (c) Violin plots showing the numbers of unique mRNA molecules (left panel) and detected genes (right panel) in cells from each sample (D4.5 Arm: 4,651 cells; D4.5 Cl13: 4,116 cells; D7 Arm: 4,678 cells; D7 Cl13: 2,597 cells). The violin represents the probability density at each value; each dot represents one cell. (d) Enriched pathways of top expressed genes in P14 cells (n = 2,597 cells) from day 7 after LCMV clone 13 infection, determined by Metascape. (e) Enriched pathways of top expressed genes in P14 cells (n = 4,678 cells) from day 7 after LCMV Armstrong infection, determined by Metascape. (f) Violin plots showing the expression of genes in cells (n = 16,042 cells) from cluster 0 through cluster 10, as defined in Fig. 2c. The violin represents the probability density at each value; each dot represents one cell.
Supplementary Figure 3 Comparison of single-cell transcriptomes between progenitor-like CD8+ T cells and memory precursor cells.
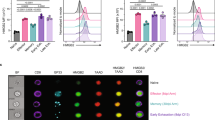
(a) Enrichment (log2 P-values) of progenitor-like gene signature in each cell (n = 16,042 cells) from the clusters defined in Fig. 2c was determined by one-sided Fisher’s exact test, illustrated in violin plots. The violin represents the probability density at each value; each dot represents one cell. (b) Enrichment (log2 P-values) of memory precursor gene signature in each cell (n = 16,042 cells) was determined by one-sided Fisher’s exact test. Left panel: t-SNE plots with P-values color-coded. Middle panel: Enrichment (log2 P-values) in each cell from the clusters defined in Fig. 2c, illustrated in violin plots. The violin represents the probability density at each value; each dot represents one cell. Right panel: Percentages of cells in each cluster with P-values above (red) or below (turquoise) 0.05. (c) Relationships among gene modules defined in Fig. 3e. The heatmap depicts adjacencies among module eigengenes. At the top of the heatmap is a hierarchical clustering of module eigengenes. Color-codes on the left side and the bottom of the heatmap represent the modules defined in Fig. 3e. Asterisks indicate modules 29 (saddlebrown) and 12 (tan). (d) Enrichment (log2 P-values) of genes in module 29 (saddlebrown) in each cell (n = 16,042 cells) from the clusters defined in Fig. 2c was determined by one-sided Fisher’s exact test, illustrated in violin plots. The violin represents the probability density at each value; each dot represents one cell. (e) Enrichment (log2 P-values) of genes in module 12 (tan) in each cell (n = 16,042 cells) from the clusters defined in Fig. 2c was determined by one-sided Fisher’s exact test, illustrated in violin plots. The violin represents the probability density at each value; each dot represents one cell. (f) Gating strategy used to sort progenitor-like (Tim3loBlimp-1lo) and terminally exhausted (Tim3hiBlimp-1hi) P14 cells on day 7 after LCMV clone 13 infection. (g) Gating strategy used to sort memory precursor (KLRG1lo) and short-lived effector (KLRG1hi) P14 cells on day 7 after LCMV Armstrong infection. (h) RNA-seq analysis of Tox mRNA in D4.5 Arm (n = 3 biological replicates), D4.5 Cl13 (n = 3 biological replicates), D7 Arm memory precursor (n = 3 biological replicates), D7 Arm short-lived effector (n = 3 biological replicates), D7 Cl13 progenitor-like (n = 2 biological replicates), and D7 Cl13 terminally exhausted (n = 2 biological replicates) P14 cells. (i) Left panel: Representative FACS plots of TOX and TCF1 expression in P14 cells from mice seven days after LCMV clone 13 infection. Right panel: FACS analysis of TOX expression in progenitor-like (TCF1hi, red), terminally exhausted (TCF1lo, green), memory precursor (KLRG1lo, orange), and short-lived effector (KLRG1hi, blue) P14 cells on day 7 post-infection. (j) TOX protein levels in D4.5 Arm (n = 5 mice), D4.5 Cl13 (n = 5 mice), D7 Arm memory precursor (n = 4 mice), D7 Arm short-lived effector (n = 4 mice), D7 Cl13 progenitor-like (n = 5 mice), and D7 Cl13 terminally exhausted (n = 5 mice) P14 cells, as determined by TOX mean fluorescent intensity (MFI) divided by isotype control MFI. In h and j, statistical significance was determined by two-sided Student’s t-test; centers and error bars represent the mean and SD. Data in f, g, i are representative of two independent experiments. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Supplementary Figure 4 Distinct H3K27ac profile in progenitor-like CD8+ T cells.
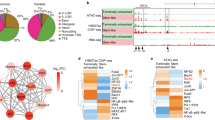
(a, b) Venn diagrams illustrating pairwise comparisons of H3K27ac peaks between progenitor-like CD8+ T cells (red) and terminally exhausted CD8+ T cells (green) (a), and between progenitor-like CD8+ T cells (red) and memory precursors (orange) (b). (c) Pie charts demonstrating the distribution of all H3K27ac peaks across the genome (three prime untranslated region [3′ UTR], five prime untranslated region [5′ UTR], exon, intergenic, intron, non-coding, promoter-transcription start sites [TSS], and transcription termination site [TTS]). (d) Hierarchical clustering of D7 Cl13 progenitor-like (cluster 10), D7 Cl13 terminally exhausted (cluster 0), D7 Arm memory precursor (cluster 7), and D7 Arm short-lived effector (clusters 2, 6, and 8) cells using scRNA-seq data from Fig. 2. (e, f) Deposition of H3K27ac centered on TSS of progenitor-like (e) or terminally exhausted (f) signature genes in progenitor-like (red) and terminally exhausted (green) CD8+ T cells. (g) Normalized H3K27ac profile at Id2 locus in progenitor-like (red), terminally exhausted (green), memory precursor (orange), and short-lived effector (blue) CD8+ T cells. Data is representative of two independent experiments. (h) Distribution of NFAT1, IRF4, RUNX3, FOXO1, STAT3, and STAT5B ChIP-seq peaks at Tox locus. (i) 5,000 naïve P14 CD8+ T cells were transferred to C57BL/6 mice that were subsequently infected with LCMV clone 13. Mice were subcutaneously injected with vehicle (n = 4 mice) or FK506 (n = 5 mice) daily starting from day 4 post-infection. TOX expression in P14 cells was determined by flow cytometry on day 7 post-infection. Two-sided Student’s t-test was used to determine statistical significance. Centers and error bars represent the mean and SD. ***P < 0.001.
Supplementary Figure 5 The effects of TOX overexpression on virus-specific CD8+ T cells during chronic LCMV infection.
Experimental setup has been described in Fig. 5. (a, b) Fold increase of Tox mRNA in progenitor-like (a) or terminally exhausted (b) TOX-overexpressing P14 cells relative to control cells (pMIG), as determined by RNA-seq. N = 3 biological replicates per group. (c) Representative FACS plots of TCF1 and Tim3 levels in TOX-overexpressing and control (pMIG) P14 cells on day 7 and 14 post-infection (gated on CD45.1+ CD8+ cells). (d) Gating strategy for sorting Tim3hi terminally exhausted P14 CD8+ T cells on day 7 after LCMV clone 13 infection. (e) Representative FACS plots and percentages of TCF1loTim3hi cells in TOX-overexpressing (n = 5 mice) and control (pMIG, n = 5 mice) P14 cells on day 5 post-transfer (gated on CD45.1+ CD8+ cells), as described in Fig. 5d. (f) Gating strategy for sorting Tim3loLy108hi (progenitor-like) and Tim3hiLy108lo (terminally exhausted) P14 CD8+ T cells on day 7 after LCMV clone 13 infection. (g-i) FACS analyses of PD1 (g, n = 5 mice per group), TIGIT (h, n = 5 mice per group), and LILRB4 (i, n = 4 mice per group) expression in control (pMIG) and TOX-overexpressing P14 CD8+ T cells on day 7 after LCMV clone 13 infection. (j–m) FACS analyses of IFNγ (j, k), TNF (j, l), and IL-2 (m) expression by control (pMIG, n = 5 mice) and TOX-overexpressing (n = 5 mice) P14 CD8+ T cells after 5-hr stimulation with GP33 peptide on day 7 after LCMV clone 13 infection. (n) in vitro killing of GP33-peptide pulsed EL4 target cells by control (pMIG, n = 3 wells) and TOX-overexpressing (n = 3 wells) P14 CD8+ T cells after 4-hr co-culture. (o–s) GSEA illustrating the enrichment of genes in hypoxia (o), oxidative phosphorylation (p), mTOR signaling (q), IFN-α responses (r), and DNA repair pathways (s), in TOX-overexpressing versus control (pMIG) progenitor-like (Tim3loLy108hi, left panels) or terminally exhausted (Tim3hiLy108lo, right panels) P14 CD8+ T cells. N = 3 biological replicates per group. Data in c, d, e are representative of two independent experiments. Data in f is representative of three independent experiments. In a, b, e, g–i, k–n, two-sided Student’s t-test was used to determine statistical significance; centers and error bars represent the mean and SD. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Supplementary Figure 6 Tox deficiency led to loss of progenitor-like CD8+ T cells.
(a) Experimental setup has been described in Fig. 6. Left panel: Representative FACS plots of H-2Db GP33 tetramer staining on WT and Tox-/- CD8+ T cells from bone marrow chimeras on day 7 post-infection. Right panel: Frequencies of H-2Db GP33 tetramer+ cells within CD45.2 WT (white) and CD45.1 WT or Tox-/- (filled) splenic CD8+ T cells. Five mice were included in each group. Data are representative of at least two independent experiments. Statistical significance was determined by two-sided paired t-test. (b, c) Naïve Toxf/f; CD4-Cre (Tox cKO) or Toxf/f (WT) P14 CD8+ T cells were transferred into WT CD45.1 recipients that were subsequently infected with LCMV clone 13. On day 7 post-infection, the numbers of P14 cells (b) and frequencies of TCF1hiTim3lo cells within P14 cells (c) were determined. Five mice were included in each group. Statistical significance was determined by two-sided Student’s t-test. Centers and error bars in a-c represent the mean and SD. **P < 0.01.
Supplementary information
Supplementary Information
Supplementary Figs. 1–6
Supplementary Table 1
Marker genes of different LCMV-specific CD8 subsets identified by scRNA-seq
Supplementary Table 2
Progenitor-like signature genes
Supplementary Table 3
Memory precursor signature genes
Supplementary Table 4
Genes in module 29 (saddlebrown)
Supplementary Table 5
Genes in module 12 (tan)
Supplementary Table 6
Terminally exhausted CD8 signature genes
Supplementary Table 7
Differentially expressed genes in TOX overexpressing Tim3loLy108hi P14 cells
Supplementary Table 8
Differentially expressed genes in TOX overexpressing Tim3hiLy108lo P14 cells
Rights and permissions
About this article
Cite this article
Yao, C., Sun, HW., Lacey, N.E. et al. Single-cell RNA-seq reveals TOX as a key regulator of CD8+ T cell persistence in chronic infection. Nat Immunol 20, 890–901 (2019). https://doi.org/10.1038/s41590-019-0403-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-019-0403-4
This article is cited by
-
Nonlinear dimensionality reduction based visualization of single-cell RNA sequencing data
Journal of Analytical Science and Technology (2024)
-
Modeling T cell temporal response to cancer immunotherapy rationalizes development of combinatorial treatment protocols
Nature Cancer (2024)
-
A type 1 immunity-restricted promoter of the IL−33 receptor gene directs antiviral T-cell responses
Nature Immunology (2024)
-
FOXP1 and KLF2 reciprocally regulate checkpoints of stem-like to effector transition in CAR T cells
Nature Immunology (2024)
-
Single-cell RNA-sequencing of virus-specific cellular immune responses in chronic hepatitis B patients
Scientific Data (2024)