Abstract
Regulatory T cells (Treg cells), whose differentiation and function are controlled by transcription factor Foxp3, express the closely related family member Foxp1. Here we explored Foxp1 function in Treg cells. We found that a large number of Foxp3-bound genomic sites in Treg cells were occupied by Foxp1 in both Treg cells and conventional T cells (Tconv cells). In Treg cells, Foxp1 markedly increased Foxp3 binding to these sites. Foxp1 deficiency in Treg cells resulted in their impaired function and competitive fitness, associated with markedly reduced CD25 expression and interleukin-2 (IL-2) responsiveness, diminished CTLA-4 expression and increased SATB1 expression. The characteristic expression patterns of CD25, Foxp3 and CTLA-4 in Treg cells were fully or partially rescued by strong IL-2 signaling. Our studies suggest that Foxp1 serves an essential non-redundant function in Treg cells by enforcing Foxp3-mediated regulation of gene expression and enabling efficient IL-2 signaling in these cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The datasets generated during the current study are included in this resource (and its supplementary information files) or are available in the Gene Expression Omnibus repository with accession code GSE121279. Datasets are relevant to Figs. 1, 2, 3, 7 and Supplementary Figs. 1, 3, 5. The ATAC-seq data relevant to Fig. 2 can be found in GEO (GSE83315) and the STAT5BCA dataset relevant to Fig. 7 can be found in GEO (GSE84553). Detailed information on experimental design and reagents may be found in the Life Sciences Reporting Summary.
References
Josefowicz, S. Z., Lu, L. & Rudensky, A. Y. Regulatory T cells: mechanisms of differentiation and function. Annu. Rev. Immunol. 30, 531–564 (2012).
Fontenot, J. D., Gavin, M. A. & Rudensky, A. Y. Foxp3 programs the development and function of CD4+CD25+ regulatory T cells. Nat. Immunol. 4, 330–336 (2003).
Hori, S., Nomura, T. & Sakaguchi, S. Control of regulatory T cell development by the transcription factor Foxp3. Science 299, 1057–1061 (2003).
Fontenot, J. D. et al. Regulatory T cell lineage specification by the forkhead transcription factor foxp3. Immunity 22, 329–341 (2005).
Lam, E. W., Brosens, J. J., Gomes, A. R. & Koo, C. Y. Forkhead box proteins: tuning forks for transcriptional harmony. Nat. Rev. Cancer 13, 482–495 (2013).
Zheng, Y. et al. Genome-wide analysis of Foxp3 target genes in developing and mature regulatory T cells. Nature 445, 936–940 (2007).
Marson, A. et al. Foxp3 occupancy and regulation of key target genes during T-cell stimulation. Nature 445, 931–935 (2007).
Song, X. et al. Structural and biological features of FOXP3 dimerization relevant to regulatory T cell function. Cell Rep. 1, 665–675 (2012).
Hu, H. et al. Foxp1 is an essential transcriptional regulator of B cell development. Nat. Immunol. 7, 819–826 (2006).
Wang, H. et al. The transcription factor Foxp1 is a critical negative regulator of the differentiation of follicular helper T cells. Nat. Immunol. 15, 667–675 (2014).
Feng, X. et al. Foxp1 is an essential transcriptional regulator for the generation of quiescent naïve T cells during thymocyte development. Blood 115, 510–518 (2010).
Feng, X. et al. Transcription factor Foxp1 exerts essential cell-intrinsic regulation of the quiescence of naïve T cells. Nat. Immunol. 12, 544–550 (2011).
Li, B. et al. FOXP3 is a homo-oligomer and a component of a supramolecular regulatory complex disabled in the human XLAAD/IPEX autoimmune disease. Int. Immunol. 19, 825–835 (2007).
Rudra, D. et al. Transcription factor Foxp3 and its protein partners form a complex regulatory network. Nat. Immunol. 13, 1010–1019 (2012).
Gerstein, M. B. et al. Architecture of the human regulatory network derived from ENCODE data. Nature 489, 91–100 (2012).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Samstein, R. M. et al. Foxp3 exploits a pre-existent enhancer landscape for regulatory T cell lineage specification. Cell 151, 153–166 (2012).
Koh, K. P., Sundrud, M. S. & Rao, A. Domain requirements and sequence specificity of DNA binding for the forkhead transcription factor FOXP3. PLoS ONE 4, e8109 (2009).
Chen, Y. et al. DNA binding by FOXP3 domain-swapped dimer suggests mechanisms of long-range chromosomal interactions. Nucleic Acids Res. 43, 1268–1282 (2015).
Zheng, Y. et al. Role of conserved non-coding DNA elements in the Foxp3 gene in regulatory T-cell fate. Nature 463, 808–812 (2010).
Ono, M. et al. Foxp3 controls regulatory T-cell function by interacting with AML1/Runx1. Nature 446, 685–689 (2007).
van der Veeken, J. et al. Memory of inflammation in regulatory T cells. Cell 166, 977–990 (2016).
Zhang, Y. et al. Foxp1 coordinates cardiomyocyte proliferation through both cell-autonomous and nonautonomous mechanisms. Genes Dev. 24, 1746–1757 (2010).
Hisaoka, T., Nakamura, Y., Senba, E. & Morikawa, Y. The forkhead transcription factors, Foxp1 and Foxp2, identify different subpopulations of projection neurons in the mouse cerebral cortex. Neuroscience 166, 551–563 (2010).
Cesario, J. M., Almaidhan, A. A. & Jeong, J. Expression of forkhead box transcription factor genes Foxp1 and Foxp2 during jaw development. Gene Expr. Patterns 20, 111–119 (2016).
Banham, A. H. et al. The FOXP1 winged helix transcription factor is a novel candidate tumor suppressor gene on chromosome 3p1. Cancer Res. 61, 8820–8829 (2001).
Gabut, M. et al. An alternative splicing switch regulates embryonic stem cell pluripotency and reprogramming. Cell 147, 132–146 (2011).
Hill, J. A. et al. Foxp3 transcription-factor-dependent and -independent regulation of the regulatory T cell transcriptional signature. Immunity 27, 786–800 (2007).
Kitagawa, Y. et al. Guidance of regulatory T cell development by Satb1-dependent super-enhancer establishment. Nat. Immunol. 18, 173–183 (2017).
Beyer, M. et al. Repression of the genome organizer SATB1 in regulatory T cells is required for suppressive function and inhibition of effector differentiation. Nat. Immunol. 12, 898–907 (2011).
Chinen, T. et al. An essential role for the IL-2 receptor in Treg cell function. Nat. Immunol. 17, 1322–1333 (2016).
Levine, A. G., Arvey, A., Jin, W. & Rudensky, A. Y. Continuous requirement for the TCR in regulatory T cell function. Nat. Immunol. 15, 1070–1078 (2014).
Sakaguchi, S., Sakaguchi, N., Asano, M., Itoh, M. & Toda, M. Immunologic self-tolerance maintained by activated T cells expressing IL-2 receptor alpha-chains (CD25). Breakdown of a single mechanism of self-tolerance causes various autoimmune diseases. J. Immunol. 155, 1151–1164 (1995).
Khattri, R., Cox, T., Yasayko, S. & Ramsdell, F. An essential role for Scurfin in CD4+CD25+ T regulatory cells. Nat. Immunol. 4, 337–342 (2003).
Lin, W. et al. Regulatory T cell development in the absence of functional Foxp3. Nat. Immunol. 8, 359–368 (2007).
Gavin, M. A. et al. Foxp3-dependent programme of regulatory T-cell differentiation. Nature 445, 771–775 (2007).
Song, X., Tang, Y. & Wang, Y. Genesis of the vertebrate FoxP subfamily member genes occurred during two ancestral whole genome duplication events. Gene 588, 156–162 (2016).
Shu, W. et al. Foxp2 and Foxp1 cooperatively regulate lung and esophagus development. Development 134, 1991–2000 (2007).
Shu, W., Yang, H., Zhang, L., Lu, M. M. & Morrisey, E. E. Characterization of a new subfamily of winged-helix/forkhead (Fox) genes that are expressed in the lung and act as transcriptional repressors. J. Biol. Chem. 276, 27488–27497 (2001).
Tamura, S., Morikawa, Y., Iwanishi, H., Hisaoka, T. & Senba, E. Foxp1 gene expression in projection neurons of the mouse striatum. Neuroscience 124, 261–267 (2004).
Wang, B. et al. Foxp1 regulates cardiac outflow tract, endocardial cushion morphogenesis and myocyte proliferation and maturation. Development 131, 4477–4487 (2004).
Banham, A. H. et al. Expression of the FOXP1 transcription factor Is strongly associated with inferior survival in patients with Diffuse Large B-Cell Lymphoma. Clin. Cancer Res. 11, 1065–1072 (2005).
Zabarovsky, E. R., Lerman, M. I. & Minna, J. D. Tumor suppressor genes on chromosome 3p involved in the pathogenesis of lung and other cancers. Oncogene 21, 6915–6935 (2002).
Bi, E. et al. Foxo1 and Foxp1 play opposing roles in regulating the differentiation and antitumor activity of TH9 cells programmed by IL-7. Sci. Signal. 10, eaak974 (2017).
Rudra, D. et al. Runx-CBFbeta complexes control expression of the transcription factor Foxp3 in regulatory T cells. Nat. Immunol. 10, 1170–1177 (2009).
Liu, W. et al. CD127 expression inversely correlates with FoxP3 and suppressive function of human CD4+ T reg cells. J. Exp. Med. 203, 1701–1711 (2006).
Feuerer, M. et al. Genomic definition of multiple ex vivo regulatory T cell subphenotypes. Proc. Natl. Acad.Sci. USA 107, 5919–5924 (2010).
Wing, K. et al. CTLA-4 control over Foxp3+ regulatory T cell function. Science 322, 271–276 (2008).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Liao, Y., Smyth, G. K. & Shi, W. The Subread aligner: fast, accurate and scalable read mapping by seed-and-vote. Nucleic Acids Res. 41, e108 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Grant, C. E., Bailey, T. L. & Noble, W. S. FIMO: scanning for occurrences of a given motif. Bioinformatics 27, 1017–1018 (2011).
Acknowledgements
We thank the Memorial Sloan Kettering Cancer Center (MSKCC) integrated genomics operation for sequencing RNA-seq and ChIP-seq libraries and preprocessing data. We thank A.C. Scott and D. Tran for help with tissue lymphocyte isolation and experiment optimization and all members of the A.Y. Rudensky laboratory for helpful discussions and technical assistance. This study was supported by NIH grants R37 AI034206 (A.Y.R.), P30 CA008748 (A.Y.R.) and U01 HG007893 (A.Y.R. and C.S.L.), as well as the Hilton–Ludwig Cancer Prevention Initiative funded by the Conrad N. Hilton Foundation and Ludwig Cancer Research (A.Y.R.). A.Y.R. is an investigator with the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
C.K. and A.Y.R. conceived of and designed the experiments. C.K., Y.P., C.S.L. and A.Y.R. interpreted the results. C.K. and A.Y.R. wrote the manuscript. C.K. performed the experiments. Y.R. generated Foxp1fl mice. Y.P. and C.S.L. designed computational analyses. Y.P. performed computational analyses.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Integrated supplementary information
Supplementary Figure 1 ChIP signal analysis.
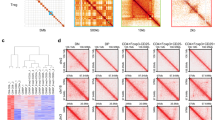
(a) The 5’ ends of strand-separated tags from a sample of peaks, aligned by the center of the peaks, for Foxp3 ChIP-seq samples (left) and Foxp1 ChIP-seq samples (right). Plots were generated by MACS2 (see Methods). (b) Foxp3 ChIP signal at sites also bound by Foxp1 (n = 1980) or sites not bound by Foxp1 (n = 1101, top) and Foxp1 ChIP signal at sites also bound by Foxp3 (n = 2011) or sites not bound by Foxp3 (n = 54, bottom) Box shows interquartile range (IQR) of a sample centered at median, notches show a confidence interval around the median estimated as median ± 1.57 x IQR/(√sample size), whiskers extend IQR endpoints by 1.5 x IQR in each direction, outliers beyond interval between whiskers are omitted from the plot, two-sided Mann-Whitney U test). (c) Foxp3 ChIP-seq read count analysis across Foxp1+ Treg and Foxp1– Treg samples in flanking regions of peaks identified library size scaling factors for each sample (left). These scaling factors were then used for the same read count analysis in actual Foxp3 peaks (right). (d, e) HOMER analysis of Foxp3 (d) and Foxp1 (e) bound sites to identify enriched transcription factor motifs. Only the top 10 most significantly enriched transcription factor motifs are shown; see Methods.
Supplementary Figure 2 Generation of the conditional Foxp1 allele.
(a) Diagram of the targeting vector containing Foxp1 exon 11 and of the targeting strategy. (b) PCR analysis of homozygous Foxp1fl/fl, heterozygous Foxp1fl/+ and homozygous Foxp1+/+ from tail DNA showing the co–integration of the 3′ downstream loxP site (376 bp) and wild-type allele product (257 bp). (c) No detectable off-target Foxp3Cre expression causes Foxp1 recombination within non-Treg cells (representative of 2 independent experiments). (d) Representative flow cytometric plots gated on splenic CD4+ T cells showing specific Foxp1 deletion in Foxp3+ Treg cells when Foxp1fl/fl was crossed to Foxp3YFP-Cre (representative of several independent experiments). (e) Representative histogram of Foxp1 protein level in naïve Foxp1+ and Foxp1– Treg cells and in naïve Tconv cells (representative of several independent experiments). (f) Summary plot of Foxp1 MFI in Foxp1+ and Foxp1– Treg cells from 8–10-week old male mice (n = 5 independent mice per group, representative of 3 independent experiments, error bars represent SEM centered on mean, ****P < 0.0001, unpaired two-tailed Student’s t test).
Supplementary Figure 3 Gene expression changes associated with Foxp1 deficiency in Foxp3- and Foxp1-bound genes.
(a) Gene ontology pathway enrichment analysis performed on clusters of genes identified in Fig. 2g. The size of each cluster (and list of all genes in each cluster) is shown in Supplementary Table 4. The size of the background set of expressed genes is 13499 genes. The enrichment tests used were one-sided hypergeometric tests (see Methods). (b) Distribution of gene expression changes associated with Foxp1 or Foxp3-bound genes. P-values calculated using two-sided KS test comparing genes associated with a binding site to all genes.
Supplementary Figure 4 Foxp1 deficiency causes Tconv cell activation and effector cytokine production.
(a) CD4+ T cell numbers in lymphoid organs of 10-week old CD4Cre Foxp1fl/fl mice (n = 3 biologically independent mice per group, representative of 2 independent experiments, error bars represent SEM centered on the mean, *P (left to right)=0.0008, 0.035, unpaired two-tailed Student’s t test). (b) Frequency of cytokine-producing T cells among all splenic CD4+ Tconv cells or CD8+ T cells from 10-week old CD4Cre Foxp1fl/fl mice after 3h in vitro stimulation with PMA and ionomycin and with addition of brefeldin A (n = 3 biologically independent mice per group, representative of 2 independent experiments, error bars represent SEM centered on the mean, *P < 0.05; **P < 0.01, unpaired two-tailed Student’s t test). (c) CD44hi cell frequency among splenic or lymph node CD4+ Tconv cell or CD8+ T cells from 10-week old CD4Cre Foxp1fl/fl mice (n = 2 and n = 3 biologically independent mice in control group and experimental group respectively, representative of 2 independent experiments, error bars represent SEM centered on the mean). (d) Ki-67+ T cell frequency among CD4+ Tconv cells or CD8+ T cells (n = 3 biologically independent mice per group, representative of 2 independent experiments, error bars represent SEM centered on the mean, *P < 0.05; **P < 0.01, unpaired two-tailed Student’s t test).
Supplementary Figure 5 Discordant Foxp1-dependent regulation of IL7R, CD25 and SATB1 expression in Treg cells versus Tconv cells.
Foxp3 repression of Il7ra is Foxp1 independent: (a) Representative histograms of IL7R protein level in naïve Foxp1+ and Foxp1– Treg cells and in naïve Tconv cells (representative of several independent experiments). (b) Summary plots of IL7R MFI in Foxp1+ and Foxp1– naïve Treg cells from 8–10-week old male mice (n = 4 biologically independent mice per group, representative of 3 independent experiments, error bars represent SEM centered on the mean, *P (left to right)=0.023, 0.0066, unpaired two-tailed Student’s t test). (c) Proliferation of sorted Foxp1+ and Foxp1– Treg cells or Foxp1– Tconv cells after 7d in vitro culture with IL-7. Foxp3– ex-Treg cells and Treg cells that maintained Foxp3 expression are both shown (Representative plots, n = 3 biologically independent mice per group with 3 technical replicates each, representative of 2 independent experiments). (d) Foxp3 and Foxp1 ChIP-seq tracks at the Il7ra locus (tracks show mean of 3 independent biological samples, Foxp1+ Treg track shows mean of 2 independent biological samples). (e) Induction of CD25 expression on activation of conventional CD4 T cells is unimpeded in the absence of Foxp1. Summary plots of CD25 MFI in Foxp1+ and Foxp1– Tconv cells after 3h in vitro culture in the absence or presence of soluble CD3 and CD28 antibodies (n = 2 and n = 3 biologically independent mice in the control and experimental group respectively, representative of 3 independent experiments, error bars represent SEM centered on the mean). (f) Summary plots of SATB1 MFI in Foxp1+ and Foxp1– Tconv cells (n = 2 and n = 3 biologically independent mice in the control and experimental group respectively, representative of 3 independent experiments, error bars represent SEM centered on the mean).
Supplementary Figure 6 In vivo suppression assay analysis.
(a) Frequency of CD45.1xCD45.2 transferred Treg cells of all CD4+ T cells in lymphoid organs on d22 post-transfer (n = 4 and n = 3 biologically independent mice for Foxp1+ group and Foxp1– group respectively, representative of 2 independent experiments, error bars represent SEM centered on the mean; *P (left to right)=0.044, 0.020, 0.0063, unpaired two-tailed Student’s t test). (b) Percentage of CD45.1xCD45.2 transferred Treg cells on d 22 post-transfer that maintained expression of Foxp3 (n = 4 and n = 3 biologically independent mice for Foxp1+ group and Foxp1– group respectively, error bars represent SEM centered on the mean; *P (left to right)=0.44, 0.44, 0.45, unpaired two-tailed Student’s t test). (c) Frequency of cytokine-producing T cells among all splenic CD45.1 CD4+ Tconv cells, CD45.1 CD8+ T cells and CD45.1xCD45.2 Treg cells after 3h in vitro stimulation with PMA and ionomycin and with addition of brefeldin A (n = 3 biologically independent mice per group, error bars represent SEM centered on the mean; *P (left to right)=0.032, 0.053, 0.026, 0.017, 0.023, 0.018, 0.11, 0.023, 0.0025, unpaired two-tailed Student’s t test).
Supplementary Figure 7 Rescue of CD25, CTLA-4, and Foxp3 expression by IL-2 exposure in Foxp1-deficient Treg cells.
(a) Representative histograms of CD25, CTLA-4, Foxp3, and SATB1 protein level in naïve Foxp1+ and Foxp1– splenic Treg cells after 24h in vitro culture in presence or absence of 500 U/mL IL-2 (representative of 2 experiments). (b) Summary plots of CD25, CTLA-4, Foxp3, and SATB1 MFI in naïve Foxp1+ and Foxp1– splenic Treg cells after 24h in vitro culture in presence or absence of 500 U/mL IL-2. In all panels n = 3 biologically independent mice per group, representative of 2 independent experiments, error bars represent SEM centered on the mean; CD25 MFI *P (left to right)=0.11, 0.003, 0.02, 0.25, CTLA4 MFI *P (left to right)=0.00085, 0.012, 0.028, 0.14, Foxp3 MFI *P (left to right)=0.05, 0.0025, 0.018, 0.19, SATB1 MFI *P (left to right)=0.05, 0.012, 0.05, 0.0014, unpaired two-tailed Student’s t test).
Supplementary information
Supplementary Figures 1-7
Supplementary Tables 1, 6 and 7
Supplementary Table 2: Foxp3 and Foxp1 ChIP-seq putative peaks
Table including sets of significant Foxp3 and Foxp1 ChIP-seq peaks in Treg cells and Tconv cells. Differential ChIP signal analysis was done with a two-sided Wald test
Supplementary Table 3: Differential gene expression analysis of Foxp1-sufficient and Foxp1-deficient Treg cells
Differential gene expression and normalized RNA-seq counts in all genes comparing naïve CD62Lhi and activated CD44hi YFP+ Treg cells isolated from Foxp3YFP-Cre/+ Foxp1fl/fl female mice. Differential expression was estimated with a two-sided Wald test using DESeq2 (see Methods)
Supplementary Table 4: GO pathway analysis of Foxp1-bound gene clusters
The size of the background set of genes (all expressed genes) is 13499 genes. Enrichment tests were run using GOstats and GO.db and are based on one-sided hyper-geometric tests (see Methods
Supplementary Table 5: STAT5BCA up and down gene signatures and Treg cell up and down genes signatures
Signatures were obtained and analyzed as described in Methods
Rights and permissions
About this article
Cite this article
Konopacki, C., Pritykin, Y., Rubtsov, Y. et al. Transcription factor Foxp1 regulates Foxp3 chromatin binding and coordinates regulatory T cell function. Nat Immunol 20, 232–242 (2019). https://doi.org/10.1038/s41590-018-0291-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-018-0291-z
This article is cited by
-
The regulation and differentiation of regulatory T cells and their dysfunction in autoimmune diseases
Nature Reviews Immunology (2024)
-
BRD9-mediated chromatin remodeling suppresses osteoclastogenesis through negative feedback mechanism
Nature Communications (2023)
-
Stepwise differentiation of follicular helper T cells reveals distinct developmental and functional states
Nature Communications (2023)
-
EBV miRNAs BART11 and BART17-3p promote immune escape through the enhancer-mediated transcription of PD-L1
Nature Communications (2022)
-
The emerging role of transcription factor FOXP3 in thyroid cancer
Reviews in Endocrine and Metabolic Disorders (2022)