Abstract
Most adult B cell lymphomas originate from germinal center (GC) B cells, but it is unclear to what extent B cells in overt lymphoma retain the functional dynamics of GC B cells or are blocked at a particular stage of the GC reaction. Here we used integrative single-cell analysis of phenotype, gene expression and variable-region sequence of the immunoglobulin heavy-chain locus to track the characteristic human GC B cell program in follicular lymphoma B cells. By modeling the cyclic continuum of GC B cell transitional states, we identified characteristic patterns of synchronously expressed gene clusters. GC-specific gene-expression synchrony was lost in single lymphoma B cells. However, distinct follicular lymphoma–specific cell states co-existed within single patient biopsies. Our data show that lymphoma B cells are not blocked in a GC B cell state but might adopt new dynamic modes of functional diversity, which opens the possibility of novel definitions of lymphoma identity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Victora, G. D. & Nussenzweig, M. C. Germinal centers. Annu. Rev. Immunol. 30, 429–457 (2012).
Mesin, L., Ersching, J. & Victora, G. D. Germinal center B cell dynamics. Immunity 45, 471–482 (2016).
Allen, C. D., Okada, T. & Cyster, J. G. Germinal-center organization and cellular dynamics. Immunity 27, 190–202 (2007).
Victora, G. D. et al. Germinal center dynamics revealed by multiphoton microscopy with a photoactivatable fluorescent reporter. Cell 143, 592–605 (2010).
Victora, G. D. et al. Identification of human germinal center light and dark zone cells and their relationship to human B-cell lymphomas. Blood 120, 2240–2248 (2012).
Küppers, R. Mechanisms of B-cell lymphoma pathogenesis. Nat. Rev. Cancer 5, 251–262 (2005).
Kridel, R., Sehn, L. H. & Gascoyne, R. D. Pathogenesis of follicular lymphoma. J. Clin. Invest. 122, 3424–3431 (2012).
Roulland, S. et al. Early steps of follicular lymphoma pathogenesis. in Advances in Immunology Vol. 111 (ed. Alt, F. W.) Ch. 1, 1–46 (Academic Press, Amsterdam, the Netherlands, 2011).
Hardianti, M. S. et al. Activation-induced cytidine deaminase expression in follicular lymphoma: association between AID expression and ongoing mutation in FL. Leukemia 18, 826–831 (2004).
Alizadeh, A. A. et al. Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature 403, 503–511 (2000).
Sungalee, S. et al. Germinal center reentries of BCL2-overexpressing B cells drive follicular lymphoma progression. J. Clin. Invest. 124, 5337–5351 (2014).
Tellier, J. et al. Human t(14;18)positive germinal center B cells: a new step in follicular lymphoma pathogenesis? Blood 123, 3462–3465 (2014).
McHeyzer-Williams, L. J., Milpied, P. J., Okitsu, S. L. & McHeyzer-Williams, M. G. Class-switched memory B cells remodel BCRs within secondary germinal centers. Nat. Immunol. 16, 296–305 (2015).
Nutt, S. L., Hodgkin, P. D., Tarlinton, D. M. & Corcoran, L. M. The generation of antibody-secreting plasma cells. Nat. Rev. Immunol. 15, 160–171 (2015).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982, https://doi.org/10.1038/nmeth.4402 (2017).
Kowalczyk, M. S. et al. Single-cell RNA-seq reveals changes in cell cycle and differentiation programs upon aging of hematopoietic stem cells. Genome Res. 25, 1860–1872 (2015).
Green, M. R. et al. Mutations in early follicular lymphoma progenitors are associated with suppressed antigen presentation. Proc. Natl. Acad. Sci. USA 112, E1116–E1125 (2015).
Seifert, M. et al. Functional capacities of human IgM memory B cells in early inflammatory responses and secondary germinal center reactions. Proc. Natl. Acad. Sci. USA 112, E546–E555 (2015).
Rousseeuw, P. J. Silhouettes: A graphical aid to the interpretation and validation of cluster analysis. J. Comput. Appl. Math. 20, 53–65 (1987).
Nutt, S. L., Taubenheim, N., Hasbold, J., Corcoran, L. M. & Hodgkin, P. D. The genetic network controlling plasma cell differentiation. Semin. Immunol. 23, 341–349 (2011).
Calado, D. P. et al. The cell-cycle regulator c-Myc is essential for the formation and maintenance of germinal centers. Nat. Immunol. 13, 1092–1100 (2012).
Dominguez-Sola, D. et al. The proto-oncogene MYC is required for selection in the germinal center and cyclic reentry. Nat. Immunol. 13, 1083–1091 (2012).
Dominguez-Sola, D. et al. The FOXO1 transcription factor instructs the germinal center dark zone program. Immunity 43, 1064–1074 (2015).
Sander, S. et al. PI3 kinase and FOXO1 transcription factor activity differentially control B cells in the germinal center light and dark zones. Immunity 43, 1075–1086 (2015).
Gitlin, A. D., Shulman, Z. & Nussenzweig, M. C. Clonal selection in the germinal centre by regulated proliferation and hypermutation. Nature 509, 637–640 (2014).
Morin, R. D. et al. Frequent mutation of histone-modifying genes in non-Hodgkin lymphoma. Nature 476, 298–303 (2011).
Okosun, J. et al. Integrated genomic analysis identifies recurrent mutations and evolution patterns driving the initiation and progression of follicular lymphoma. Nat. Genet. 46, 176–181 (2014).
Pasqualucci, L. et al. Genetics of follicular lymphoma transformation. Cell Reports 6, 130–140 (2014).
Kridel, R. et al. Histological transformation and progression in follicular lymphoma: a clonal evolution study. PLoS Med. 13, e1002197, https://doi.org/10.1371/journal.pmed.1002197 (2016).
Ortega-Molina, A. et al. The histone lysine methyltransferase KMT2D sustains a gene expression program that represses B cell lymphoma development. Nat. Med. 21, 1199–1208 (2015).
Zhang, J. et al. Disruption of KMT2D perturbs germinal center B cell development and promotes lymphomagenesis. Nat. Med. 21, 1190–1198 (2015).
García-Ramírez, I. et al. Crebbp loss cooperates with Bcl2 overexpression to promote lymphoma in mice. Blood 129, 2645–2656 (2017).
Hashwah, H. et al. Inactivation of CREBBP expands the germinal center B cell compartment, down-regulates MHCII expression and promotes DLBCL growth. Proc. Natl. Acad. Sci. USA 114, 9701–9706 (2017).
Jiang, Y. et al. CREBBP inactivation promotes the development of HDAC3-dependent lymphomas. Cancer Discov 7, 38–53 (2017).
Zhang, J. et al. The CREBBP acetyltransferase is a haploinsufficient tumor suppressor in B-cell lymphoma. Cancer Discov. 7, 322–337 (2017).
Béguelin, W. et al. EZH2 is required for germinal center formation and somatic EZH2 mutations promote lymphoid transformation. Cancer Cell 23, 677–692 (2013).
Koues, O. I. et al. Enhancer sequence variants and transcription-factor deregulation synergize to construct pathogenic regulatory circuits in B-cell lymphoma. Immunity 42, 186–198 (2015).
Jiang, Y., Dominguez, P. M. & Melnick, A. M. The many layers of epigenetic dysfunction in B-cell lymphomas. Curr. Opin. Hematol. 23, 377–384 (2016).
Pangault, C. et al. Follicular lymphoma cell niche: identification of a preeminent IL-4-dependent TFH-B cell axis. Leukemia 24, 2080–2089 (2010).
Amé-Thomas, P. et al. Characterization of intratumoral follicular helper T cells in follicular lymphoma: role in the survival of malignant B cells. Leukemia 26, 1053–1063 (2012).
Mourcin, F., Pangault, C., Amin-Ali, R., Amé-Thomas, P. & Tarte, K. Stromal cell contribution to human follicular lymphoma pathogenesis. Front. Immunol. 3, 280, https://doi.org/10.3389/fimmu.2012.00280 (2012)..
Pereira, J. P., Kelly, L. M. & Cyster, J. G. Finding the right niche: B-cell migration in the early phases of T-dependent antibody responses. Int. Immunol. 22, 413–419 (2010).
Gitlin, A. D. et al. Humoral immunity. T cell help controls the speed of the cell cycle in germinal center B cells. Science 349, 643–646 (2015).
Dave, S. S. et al. Prediction of survival in follicular lymphoma based on molecular features of tumor-infiltrating immune cells. N. Engl. J. Med. 351, 2159–2169 (2004).
Carlotti, E. et al. High throughput sequencing analysis of the immunoglobulin heavy chain gene from flow-sorted B cell sub-populations define the dynamics of follicular lymphoma clonal evolution. PLoS One 10, e0134833 (2015).
Carlotti, E. et al. Transformation of follicular lymphoma to diffuse large B-cell lymphoma may occur by divergent evolution from a common progenitor cell or by direct evolution from the follicular lymphoma clone. Blood 113, 3553–3557 (2009).
Finak, G., Perez, J.-M., Weng, A. & Gottardo, R. Optimizing transformations for automated, high throughput analysis of flow cytometry data. BMC Bioinformatics 11, 546 (2010).
Smith, K. et al. Rapid generation of fully human monoclonal antibodies specific to a vaccinating antigen. Nat. Protoc. 4, 372–384 (2009).
Tiller, T. et al. Efficient generation of monoclonal antibodies from single human B cells by single cell RT-PCR and expression vector cloning. J. Immunol. Methods 329, 112–124 (2008).
van Dongen, J. J. M. et al. Design and standardization of PCR primers and protocols for detection of clonal immunoglobulin and T-cell receptor gene recombinations in suspect lymphoproliferations: report of the BIOMED-2 Concerted Action BMH4-CT98-3936. Leukemia 17, 2257–2317 (2003).
van der Maaten, L. & Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
McDavid, A. et al. Data exploration, quality control and testing in single-cell qPCR-based gene expression experiments. Bioinformatics 29, 461–467 (2013).
Golub, T. R. et al. Molecular classification of cancer: class discovery and class prediction by gene expression monitoring. Science 286, 531–537 (1999).
Wu, Z., Irizarry, R. A., Gentleman, R., Martinez-Murillo, F. & Spencer, F. A model-based background adjustment for oligonucleotide expression arrays. J. Am. Stat. Assoc. 99, 909–917 (2004).
Li, Q., Birkbak, N. J., Gyorffy, B., Szallasi, Z. & Eklund, A. C. Jetset: selecting the optimal microarray probe set to represent a gene. BMC Bioinformatics 12, 474 (2011).
Giudicelli, V., Chaume, D. & Lefranc, M.-P. IMGT/V-QUEST, an integrated software program for immunoglobulin and T cell receptor V-J and V-D-J rearrangement analysis. Nucleic Acids Res 32, W435–W440 (2004).
Huson, D. H. et al. Dendroscope: an interactive viewer for large phylogenetic trees. BMC Bioinformatics 8, 460 (2007).
Acknowledgements
We thank all members of the Nadel laboratory, especially S. Roulland, for discussions and comments; the bioinformatics platform of Centre d’Immunologie de Marseille-Luminy; J. Hardwigsen (Assistance Publique – Hôpitaux de Marseille) for normal human spleen samples; Mi-Mabs and Cancéropôle Provence-Alpes-Côte d’Azur for support in the single-cell qPCR analysis platform; and HalioDX for providing access to the 10x Genomics Chromium system. Supported by Fondation ARC (fellowship to P.M.; PGA 120150202381 to B.N.; and grants to P.M.), Cancéropôle Provence-Alpes-Côte d’Azur (I.C.-M.; and grants to P.M.), Medimmune (11799A10 for M.-L.M.), Fondation pour la Recherche Médicale (G.B.) and GEFLUC Marseille (grants to P.M.).
Author information
Authors and Affiliations
Contributions
P.M. designed the study, performed the experiments, supervised data analysis and wrote the manuscript; I.C.-M. analyzed the data and wrote the manuscript; M.-L.M. performed the experiments; B.T. analyzed the public microarray data; G.B., A.T.-G. and G.S. provided human lymphoma samples and critical insight in follicular lymphoma pathology; L.S. supervised data analysis, analyzed the data and wrote the manuscript; B.N. provided direction in the study design and wrote the manuscript; and all authors reviewed and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Integrative single-cell analysis for human B cell subsets segregation.
(a) Integrative single-cell analysis strategy used on normal human B cell subsets. (b) Flow cytometry gating strategy for single-cell sorting of GC B cells from human spleen or tonsil samples. Index sorting data for CXCR4 and CD83 expression allowed us to assign sorted GC B cells to the LZ (CXCR4loCD83hi) or DZ (CXCR4hiCD83lo) subsets a posteriori. In some instances, to promote balanced representation of LZ and DZ subsets in sorted single cells, equal numbers of cells were sorted from 4 separate gates spanning the spectrum of CXCR4 and CD83 expression levels. (c) Flow cytometry gating strategy for single-cell sorting of plasmablasts / plasma cells and memory B cells from human spleen or tonsil samples. Index sorting data for CD19 and CD20 expression on CD38hiSSChiCD3-CD27+ single-cells allowed us to assign sorted cells to the early plasmablast (CD19+CD20+), late plasmablasts (CD19+CD20-), or plasma cells (CD19−CD20−) subsets a posteriori. Four different subsets of isotype-defined CD27+ memory B cells were sorted based on IgD, IgM and IgG expression. (d) Gene expression correlation dot plots of actual gene expression levels measured in 10-cell samples of B cells from the indicated sample (y-axis), compared to their extrapolated values from single-cell measured values (x-axis). The first diagonal (red line), the linear regression line (blue line), and the Pearson correlation coefficient (R2) are indicated. N = 364 gene expression measures per sample (pooled from 4 replicates). (e) Average gene expression in 1-cell, 10-cell and 30-cell samples of GC B cells for 71 genes for which 100% of 30-cell GC B cell samples were positive. Number of cells per sample is plotted on a log2 scale. Each gene’s average values are linked by a connecting line. (f) Hierarchical clustering of single-cell gene expression values in normal human B cell subsets (cells: Spearman correlation distance, genes: euclidean distance, average linking). Number of cells = 767. (g) Projection of single human B cells (n = 767) on the first 2 principal components computed by PCA on the 91-gene expression matrix (PC1: 19% of total variability, PC2: 8% of total variability). Cells are colored based on their phenotype. (h) Visualization of PCA gene loadings on PC1 and PC2 for top contributing genes (accounting for 60% of total information for each PC).
Supplementary Figure 2 Modeling of human GC B cell gene expression changes based on single-cell analysis.
(a) Volcano plot of 91 genes showing the difference of the mean expression between DZ cells and LZ cells vs. -log10 of the LRT P-value. The grey line shows the 0.05 level of significance on LRT test. The red lines show Z-score values of -1 and + 1. Significant differentially expressed genes are highlighted in red and labelled. (b) Projection of single human GC B cells (n = 503) on the first 2 principal components computed by PCA on the 91-gene expression matrix (PC1: 12% of total variability, PC2: 9% of total variability). Cells are colored based on their sample of origin. (c) Distribution of euclidean distances of single GC B cells from the (0,0) origin in the PC1 x PC2 projection. The mean (red line) and 95% CI (blue lines) are indicated. Note that only 4.57% of cells are at a distance < 5 from the (0,0) origin. (d) k-means clustering of the single-cell 91-gene expression matrix was performed with the indicated values of k (from 3 to 7). For each k, the PC1 x PC2 projection of GC B cells colored by cluster identity (top) and the distribution of θGC values of cells in each cluster (bottom) are shown. Note that the radial repartition of clusters in the PC1 x PC2 space is conserved for all values of k. (e) Sample origin repartition of GC B cells in each k-means cluster (k = 5). (f) Index sorting defined phenotype repartition of GC B cells in each k-means cluster (k = 5). (DZ, CXCR4hiCD83lo; LZ, CXCR4loCD83hi; other, CXCR4loCD83lo). (g) CCNB1 gene expression in single human GC B cells laid out on the circular model. (h) TriggerPulseWidth parameter (y-axis, a proxy for cell size) in single human GC B cells ordered along the θGC pseudotime (x-axis). Black profile line indicates average TriggerPulseWidth levels evolution along θGC. Cells are colored based on CCNB1 expression (n.d.: not detected).
Supplementary Figure 3 High throughput single-cell RNAseq analysis of human GC B cells and FL B cells.
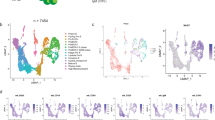
(a) Flow cytometry gating strategy for cell sorting of IgDneg B cells and GC B cells from human spleen for high throughput single-cell RNAseq analysis. (b) Experimental strategy for high throughput single-cell RNAseq of human B cells enriched in GC B cells. (c) Projection of single human B cells (n = 859) on the first 2 principal components computed by PCA on the 1146 variable genes (PC1: 23% of total variability, PC2: 5% of total variability). Cells are colored based on their expression of the GC marker BCL6. GC B cells were discriminated from IgDneg non-GC B cells based on their PC1 projection, as shown. (d) Experimental strategy for high throughput single-cell RNAseq of human FL cells. (e) Projection of single human FL cells (n = 1848) on the first 2 principal components computed by PCA on the 1025 variable genes (PC1: 15% of total variability, PC2: 2% of total variability). Cells are colored based on their expression of the T cell marker CD3D (left) and the B cell marker MS4A1 (right). Malignant FL B cells were discriminated from T cell microenvironment based on their PC1 projection, as shown. (f) Single-cell gene expression heatmaps for our 91-gene panel, as measured by qPCR (left) and high throughput single-cell RNAseq (right). Single-cells (columns) are grouped based on their sample of origin (color-coded in the first row). Genes (rows) are ordered by descending average expression (top to bottom). (g) Percentage of cells with detected expression of each gene of our 91-gene panel within the indicated cell type (left, GC B cells; right, FL B cells) computed from qPCR and RNAseq methods. Grey lines connect identical genes. Red line indicates mean. **** two-tailed P-value < 0.0001 in Wilcoxon matched pairs signed rank test (n = 91). (h) Assignment of cell cycle phase to human GC B cells (n = 358) after computing expression scores of lists of genes characteristic of S phase (x-axis) or G2/M phase (y-axis). Cells in the red gate were assigned to the G1 phase; cells in the blue gate were assigned to the S phase; cells in the green gate were assigned to the G2/M phase. (i) Projection of single human GC B cells (n = 358) on the PC2 and PC3 components computed by PCA on the 1450 variable genes (PC2: 3% of total variability, PC3: 2% of total variability). Cells are colored based on their assigned cell cycle phase. (j) Single cells scatter plot of the expression score of lists of genes characteristic of LZ B cells (x-axis) or DZ B cells (y-axis). Cells are colored based on their assigned cell cycle phase. (k) Single-cell gene expression heatmap of human GC B cells (n = 358) for 397 genes previously published as being differentially expressed between human DZ and LZ GC B cells. Single cells (columns) are ordered based on their DZ-LZ score (Fig. 3c). Assigned cell cycle phase is color-coded in the first row. Genes (rows) are ordered by descending (genes up in DZ) or ascending (genes up in LZ) average expression (top to bottom).
Supplementary Figure 4 Gene expression evolution in GC B cells along θGC pseudotime.
Evolution of gene expression in human GC B cells along the θGC pseudotime for the indicated genes grouped by functional categories.
Supplementary Figure 5 The 91-gene panel is relevant to discriminate gene expression profiles of normal B cell subsets and FL B cells in bulk gene expression datasets.
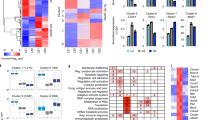
(a) Published gene expression microarray datasets for FACS-purified FL B cells (Green et al.), human LZ and DZ GC B cells (Victora et al.), and human circulating naïve and memory B cells (Seifert et al.), were downloaded and reanalyzed together. The top 500 variable genes on the combined dataset were used to generate a hierarchical clustering dendrogram (Euclidean distance, Ward agglomerative method) of all samples. (b) Only the probesets corresponding to the 91-gene panel analyzed by single-cell qPCR were retained for heatmap representation and hierarchical clustering (genes: Pearson correlation; samples: Euclidean distance; Ward agglomerative method). Of note, 21 out of the 91 genes were present in the top 500 variable genes.
Supplementary Figure 6 Single-cell analysis of FL B cells heterogeneity.
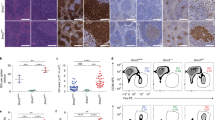
(a) Integrative single-cell analysis strategy used on FL B cells. (b) Flow cytometry gating strategy for single-cell sorting of FL B cells from human lymph nodes harvested at diagnosis. The example of patient FL 1 is shown here. The Ig light chain restriction gate was specifically tailored to each patient’s case depending on the known isotype restriction. Although not used in the gating strategy, surface CXCR4 and CD83 expression in sorted cells was recorded through index sorting. (c) Hierarchical clustering of single-cell gene expression values in FL B cells sorted from the indicated patients (top row) (cells: euclidean distance, genes: euclidean distance, average linking). (d) Projection of single human FL B cells (n = 714) on the PC1 x PC2 (top left) and PC2 x PC3 (bottom left) components computed by PCA on the 91-gene expression matrix (PC1: 10% of total variability, PC2: 5% of total variability, PC3: 4% of total variability). Cells are colored based on their sample of origin. Corresponding PCA gene loadings on PC1 x PC2 (top right) and PC2 x PC3 (bottom right) for top contributing genes (accounting for 60% of total information for each PC). (e) Projection of single human FL B cells (n = 714) on the PC1 x PC2 (left) components computed by PCA on the 5-protein surface expression matrix from index sorting (PC1: 46% of total variability, PC2: 24% of total variability). Cells are colored based on their sample of origin. Corresponding PCA protein surface expression loadings on PC1 x PC2 (right).
Supplementary Figure 7 Somatic hypermutation analysis of single-cell IGH sequencing of FL B cells.
(a-e) For each FL sample (FL 1 to 5, a to e panels, respectively), single-cell IGH sequences were analyzed to infer the VH, D and JH genes used (top, inside parentheses), and the position and number of somatic mutations. Somatic mutations along the IGH gene, broken in framework (FR1, FR2, FR3, FR4) and complementarity determining regions (CDR1, CDR2, CDR3) as indicated, are indicated by red lines (mutation inducing an amino acid change) or blue lines (silent mutations). (1) shows mutations inferred to have occurred during evolution from the unmutated common ancestor (UCA) to the nearest common ancestor (NCA) of all analyzed FL B cells. (2) shows the mutations in all observed single FL B cells inferred to have occurred during evolution from the NCA. Mutation bar height represents the number of unique FL B cell IGH sequences with a mutation in that position. AID hotspots (RGYW sequence) are indicated by a purple bar below the graph. (3) shows the density of UCA to NCA IGH somatic mutations in FR and CDR coding regions, shown as the percentage of nucleotide sites with inferred somatic mutations. (4) shows the density of NCA to FL B cell IGH somatic mutations in FR and CDR coding regions, shown as the percentage of nucleotide sites with observed somatic mutations.
Supplementary Information
Supplementary Text and Figures
Supplementary Figures 1-7 and Supplementary Tables 1-5
Single-cell qPCR and IGHseq data
Raw annotated single-cell qPCR gene expression data and IGHseq data, when available, for all cells analyzed in the study
Single-cell qPCR data summary
Summary of the performance of each of the 91 qPCR gene expression assays on all B cell samples analyzed in the study
Gene lists used in single-cell RNAseq analyses
Annotated list of genes used to compute the DZ and LZ scores for Fig. 3c and Supplementary Fig. 3j (sheet 1), and annotated lists of genes in clusters 1, 3 and 5 from Fig. 4c-d (sheets 2-4)
Rights and permissions
About this article
Cite this article
Milpied, P., Cervera-Marzal, I., Mollichella, ML. et al. Human germinal center transcriptional programs are de-synchronized in B cell lymphoma. Nat Immunol 19, 1013–1024 (2018). https://doi.org/10.1038/s41590-018-0181-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-018-0181-4
This article is cited by
-
Recent advances in understanding the biology of follicular lymphoma
International Journal of Hematology (2024)
-
Single-cell spatial metabolomics with cell-type specific protein profiling for tissue systems biology
Nature Communications (2023)
-
Germinal center-dependent and -independent immune responses of tumor-infiltrating B cells in human cancers
Cellular & Molecular Immunology (2023)
-
Transfer learning enables predictions in network biology
Nature (2023)
-
Understanding repertoire sequencing data through a multiscale computational model of the germinal center
npj Systems Biology and Applications (2023)