Abstract
Within germinal centers (GCs), complex and highly orchestrated molecular programs must balance proliferation, somatic hypermutation and selection to both provide effective humoral immunity and to protect against genomic instability and neoplastic transformation. In contrast to this complexity, GC B cells are canonically divided into two principal populations, dark zone (DZ) and light zone (LZ) cells. We now demonstrate that, following selection in the LZ, B cells migrated to specialized sites within the canonical DZ that contained tingible body macrophages and were sites of ongoing cell division. Proliferating DZ (DZp) cells then transited into the larger DZ to become differentiating DZ (DZd) cells before re-entering the LZ. Multidimensional analysis revealed distinct molecular programs in each population commensurate with observed compartmentalization of noncompatible functions. These data provide a new three-cell population model that both orders critical GC functions and reveals essential molecular programs of humoral adaptive immunity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request. Bulk RNA-seq, scRNA-seq, ATAC–seq and ChIP–seq data have been deposited in the Gene Expression Omnibus database under accession code GSE133743. Proteome and phosphoproteome data have been uploaded to ProteomeXchange via the PRIDE database. The project name is ‘Proteome profiling of mouse GCBCs’, and the project accession code is PXD015524. Data from GSE100738 were also analyzed.
References
De Silva, N. S. & Klein, U. Dynamics of B cells in germinal centres. Nat. Rev. Immunol. 15, 137–148 (2015).
MacLennan, I. C. Germinal centers. Annu. Rev. Immunol. 12, 117–139 (1994).
Bannard, O. et al. Germinal center centroblasts transition to a centrocyte phenotype according to a timed program and depend on the dark zone for effective selection. Immunity 39, 912–924 (2013).
Victora, G. D. & Nussenzweig, M. C. Germinal centers. Ann. Rev. Immunol. 30, 429–457 (2012).
Victora, G. D. et al. Germinal center dynamics revealed by multiphoton microscopy with a photoactivatable fluorescent reporter. Cell 143, 592–605 (2010).
Shulman, Z. et al. Dynamic signaling by T follicular helper cells during germinal center B cell selection. Science 345, 1058–1062 (2014).
Khalil, A. M., Cambier, J. C. & Shlomchik, M. J. B cell receptor signal transduction in the GC is short-circuited by high phosphatase activity. Science 336, 1178–1181 (2012).
Calado, D. P. et al. The cell-cycle regulator c-Myc is essential for the formation and maintenance of germinal centers. Nat. Immunol. 13, 1092–1100 (2012).
Dominguez-Sola, D. et al. The proto-oncogene MYC is required for selection in the germinal center and cyclic reentry. Nat. Immunol. 13, 1083–1091 (2012).
Allen, C. D. et al. Germinal center dark and light zone organization is mediated by CXCR4 and CXCR5. Nat. Immunol. 5, 943–952 (2004).
Song, S. & Matthias, P. D. The transcriptional regulation of germinal center formation. Front. Immunol. 9, 2026 (2018).
Tunyaplin, C. et al. Direct repression of prdm1 by Bcl-6 inhibits plasmacytic differentiation. J. Immunol. 173, 1158–1165 (2004).
Ochiai, K. et al. Transcriptional regulation of germinal center B and plasma cell fates by dynamical control of IRF4. Immunity 38, 918–929 (2013).
Recaldin, T. & Fear, D. J. Transcription factors regulating B cell fate in the germinal centre. Clin. Exp. Immunol. 183, 65–75 (2016).
Dominguez-Sola, D. et al. The FOXO1 transcription factor instructs the germinal center dark zone program. Immunity 43, 1064–1074 (2015).
Trabucco, S. E., Gerstein, R. M. & Zhang, H. YY1 Regulates the germinal center reaction by inhibiting apoptosis. J. Immunol. 197, 1699–1707 (2016).
Perez-Garcia, A. et al. CTCF orchestrates the germinal centre transcriptional program and prevents premature plasma cell differentiation. Nat. Commun. 8, 16067 (2017).
Caganova, M. et al. Germinal center dysregulation by histone methyltransferase EZH2 promotes lymphomagenesis. J. Clin. Invest. 123, 5009–5022 (2013).
Beguelin, W. et al. EZH2 is required for germinal center formation and somatic EZH2 mutations promote lymphoid transformation. Cancer Cell 23, 677–692 (2013).
Yoshida, H. et al. The cis-regulatory atlas of the mouse immune system. Cell 176, 897–912.e20 (2019).
Röhlich, K. Beitrag zur Cytologie der Keimzentren der Lymphknoten. Z. Mikrosk. Anat. Forsch 20, 287–297 (1930).
Endl, E. & Gerdes, J. Posttranslational modifications of the KI-67 protein coincide with two major checkpoints during mitosis. J. Cell. Physiol. 182, 371–380 (2000).
Chistiakov, D. A., Killingsworth, M. C., Myasoedova, V. A., Orekhov, A. N. & Bobryshev, Y. V. CD68/macrosialin: not just a histochemical marker. Lab. Invest. 97, 4–13 (2017).
Brink, R. & Phan, T. G. Self-reactive B cells in the germinal center reaction. Annu. Rev. Immunol. 36, 339–357 (2018).
Weber, T. S. Cell cycle-associated CXCR4 expression in germinal center B cells and its implications on affinity maturation. Front. Immunol. 9, 1313 (2018).
Allen, C. D., Okada, T., Tang, H. L. & Cyster, J. G. Imaging of germinal center selection events during affinity maturation. Science 315, 528–531 (2007).
Mesin, L., Ersching, J. & Victora, G. D. Germinal center B cell dynamics. Immunity 45, 471–482 (2016).
Gitlin, A. D., Shulman, Z. & Nussenzweig, M. C. Clonal selection in the germinal centre by regulated proliferation and hypermutation. Nature 509, 637–640 (2014).
Finkin, S., Hartweger, H., Oliveira, T. Y., Kara, E. E. & Nussenzweig, M. C. Protein amounts of the MYC transcription factor determine germinal center B cell division capacity. Immunity 51, 324–336.e5 (2019).
Vervoorts, J., Luscher-Firzlaff, J. & Luscher, B. The ins and outs of MYC regulation by posttranslational mechanisms. J. Biol. Chem. 281, 34725–34729 (2006).
Yao, S., Fan, L. Y. & Lam, E. W. The FOXO3-FOXM1 axis: a key cancer drug target and a modulator of cancer drug resistance. Semin. Cancer Biol. 50, 77–89 (2018).
Ersching, J. et al. Germinal center selection and affinity maturation require dynamic regulation of mTORC1 kinase. Immunity 46, 1045–1058.e6 (2017).
Gitlin, A. D. et al. T cell help controls the speed of the cell cycle in germinal center B cells. Science 349, 643–646 (2015).
Vitale, I., Galluzzi, L., Castedo, M. & Kroemer, G. Mitotic catastrophe: a mechanism for avoiding genomic instability. Nat. Rev. Mol. Cell Biol. 12, 385–392 (2011).
Kuppers, R. & Dalla-Favera, R. Mechanisms of chromosomal translocations in B cell lymphomas. Oncogene 20, 5580–5594 (2001).
Clark, M. R., Mandal, M., Ochiai, K. & Singh, H. Orchestrating B cell lymphopoiesis through interplay of IL-7 receptor and pre-B cell receptor signalling. Nat. Rev. Immunol. 14, 69–80 (2014).
Zhang, L., Reynolds, T. L., Shan, X. & Desiderio, S. Coupling of V(D)J recombination to the cell cycle suppresses genomic instability and lymphoid tumorigenesis. Immunity 34, 163–174 (2011).
Khair, L. et al. ATM increases activation-induced cytidine deaminase activity at downstream S regions during class-switch recombination. J. Immunol. 192, 4887–4896 (2014).
Schrader, C. E., Guikema, J. E., Linehan, E. K., Selsing, E. & Stavnezer, J. Activation-induced cytidine deaminase-dependent DNA breaks in class switch recombination occur during G1 phase of the cell cycle and depend upon mismatch repair. J. Immunol. 179, 6064–6071 (2007).
Petersen, S. et al. AID is required to initiate Nbs1/γ-H2AX focus formation and mutations at sites of class switching. Nature 414, 660–665 (2001).
Sharbeen, G., Yee, C. W., Smith, A. L. & Jolly, C. J. Ectopic restriction of DNA repair reveals that UNG2 excises AID-induced uracils predominantly or exclusively during G1 phase. J. Exp. Med. 209, 965–974 (2012).
Wang, Q. et al. The cell cycle restricts activation-induced cytidine deaminase activity to early G1. J. Exp. Med. 214, 49–58 (2017).
Storb, U. Why does somatic hypermutation by AID require transcription of its target genes? Adv. Immunol. 122, 253–277 (2014).
Zan, H. & Casali, P. Regulation of Aicda expression and AID activity. Autoimmunity 46, 83–101 (2013).
Stewart, I., Radtke, D., Phillips, B., McGowan, S. J. & Bannard, O. Germinal center B cells replace their antigen receptors in dark zones and fail light zone entry when immunoglobulin gene mutations are damaging. Immunity 49, 477–489.e7 (2018).
Hodson, D. J. et al. Regulation of normal B-cell differentiation and malignant B-cell survival by OCT2. Proc. Natl Acad. Sci. USA 113, E2039–E2046 (2016).
Vilagos, B. et al. Essential role of EBF1 in the generation and function of distinct mature B cell types. J. Exp. Med. 209, 775–792 (2012).
Kwon, K. et al. Instructive role of the transcription factor E2A in early B lymphopoiesis and germinal center B cell development. Immunity 28, 751–762 (2008).
Milpied, P. et al. Human germinal center transcriptional programs are de-synchronized in B cell lymphoma. Nat. Immunol. 19, 1013–1024 (2018).
Veselits, M. et al. Igβ ubiquitination activates PI3K signals required for endosomal sorting. J. Exp. Med. 214, 3775–3790 (2017).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Mandal, M. et al. CXCR4 signaling directs Igk recombination and the molecular mechanisms of late B lymphopoiesis. Nat. Immunol. 20, 1393–1403 (2019).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at ArXiv https://arxiv.org/abs/1303.3997 (2013).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Li, Q., Brown, J., Huang, H. & Bickel, P. Measuring reproducibility of high-throughput experiments. Ann. Appl. Stat. 5, 1752–1779 (2011).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Tan, H. et al. Integrative proteomics and phosphoproteomics profiling reveals dynamic signaling networks and bioenergetics pathways underlying T cell activation. Immunity 46, 488–503 (2017).
Tan, H. et al. Refined phosphopeptide enrichment by phosphate additive and the analysis of human brain phosphoproteome. Proteomics 15, 500–507 (2015).
Wang, H. et al. Systematic optimization of long gradient chromatography mass spectrometry for deep analysis of brain proteome. J. Proteome Res. 14, 829–838 (2015).
Niu, M. et al. Extensive peptide fractionation and y 1 ion-based interference detection method for enabling accurate quantification by isobaric labeling and mass spectrometry. Anal. Chem. 89, 2956–2963 (2017).
Stewart, E. et al. Identification of therapeutic targets in rhabdomyosarcoma through integrated genomic, epigenomic, and proteomic analyses. Cancer Cell 34, 411–426 e419 (2018).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinf. 9, 559 (2008).
Macosko, E. Z. et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161, 1202–1214 (2015).
Qiu, X. et al. Single-cell mRNA quantification and differential analysis with Census. Nat. Methods 14, 309–315 (2017).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Acknowledgements
We thank D. Leclerc and the UChicago flow cytometry core for providing the latest flow cytometry technologies. We also thank Y. Wang for assistance in proteomics experiments. This work is supported by the US National Institutes of Health grant nos. R01AI143778 and R21AI128785 (to M.R.C.), T32 HL007605 and F32AI143120 (to D.E.K.), T32HD007009 (to M.K.O.), R01AG047928 (to J.P.) and the Cancer Center Support grant (no. P30CA014599) for the UChicago flow cytometry core.
Author information
Authors and Affiliations
Contributions
D.E.K. and M.R.C. conceived and designed experiments. D.E.K. performed and analyzed most of the experiments. D.E.K. and M.M.-C. analyzed the high throughput sequencing data. M.K.O assisted in scRNA-seq experiments. J.A. assisted with imaging experiments. M.V. assisted in influenza experiments. Y.D., H.W., J.P. and H.C. assisted with proteomics experiments and analyses. M.M. and K.C.M. assisted with some experiments. M.M. provided valuable insights into the study design. D.E.K. and M.R.C. oversaw the entire project and edited the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information L.A. Dempsey was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
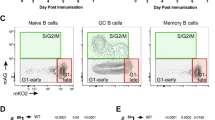
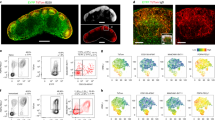
Extended Data Fig. 1 Transcriptional analysis of GCBC subsets in support of Fig. 1.
a, Schematic of immunization strategy. Mice were immunized intraperitoneally with sheep red blood cells (SRBCs), boosted on day 5, followed by isolation of GCBC subsets from spleens. b, RNA-Seq volcano plot displaying the -Log10 FDR vs Log2 fold-change of genes differentially expressed between DZ and LZ cells isolated by the Classical GC strategy. c, RNA-Seq heatmap and volcano plot displaying head-to-head differentially expressed genes between DZ vs LZ B cells isolated using the New Strategy in Fig. 1c. d and e, RNA-Seq heatmaps and volcano plots displaying head-to-head differentially expressed genes between GZ vs LZ (d) and DZ vs GZ (e) B cells isolated using the New Strategy in Fig. 1c. For RNA-Seq, n = 2 per cell type. Each n represents cells pooled from 20 mice. q values were generated with edgeR (see Methods).
Extended Data Fig. 2 Epigenetic differences between GCBC subsets in support of Fig. 2.
a, Genome accessibility and enhancer tracks aligned at the Otub2 locus. b, mRNA expression of Otub2. c, mRNA expression of the indicated TF in LZ, GZ and DZ B cells. d, TF motifs enriched in accessible regions for the indicated genome accessibility cluster and associated GCBC subset. p values were generated using HOMER (see Methods). For each cluster, n= the number of accessibility peaks indicated in Fig. 2f. e and f, Genome accessibility and enhancer tracks aligned at the Foxn2 (e) and Ccnb1 (f) locus for GCBCs isolated by the Classic GC method. For ATAC-Seq data, n = 2 per cell type. Each n represents cells pooled from 20 mice. (b,c) Each dot corresponds to an independent biological sample.
Extended Data Fig. 3 Phosphoproteomic analysis of GCBC subsets in support of Fig. 3.
a, Schematic of phosphoproteome differential expression analysis on GC populations isolated with the New gating strategy. b, Phosphoproteome cell distance plot of the indicated cell populations. Scale represents Euclidian distance. c, Box plots of phosphoproteomic clustering analysis (left). Boxes represent interquartile ranges (IQRs; Q1–Q3 percentile) and black vertical lines represent median values. Maximum and minimum values (ends of whiskers) are defined as Q3 + 1.5× the IQR and Q1 − 1.5× the IQR, respectively. Pathway analysis associated with the indicated cluster (right). Numbers correspond to -log10 p value. For each cluster, n= the number of phosphoproteins indicated above boxplot. P values were generated by Metascape using an established hypergeometric test coupled with Benjamini-Hochberg p-value correction algorithm. Light blue, gray, and dark blue (left side of heatmap) correspond to the LZ, GZ, or DZ subsets respectively. d, Total protein expression of Ki67. e, Relative levels of the indicated Ki67 phosphopeptides from cluster 1. f, List of GZ upregulated phosphopeptides for Ki67 from cluster 1. For phosphoproteome data, n = 2 per cell type. Cells were isolated from a total of 120 mice. Each n was generated from 5-6 million purified B cell subsets. See also Supplementary Data 5.
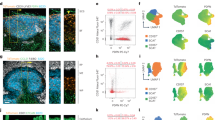
Extended Data Fig. 4 Histological analysis of the GZ in support of Fig. 4.
a and b, Immunofluorescence microscopy of GCs 14 days post SRBC immunization. Single panels and merged image displaying GL7, CD35, and Cyclin B1 in two distinct GCs. (n > 7 mice) c, Expression of Lars2 (mitochondrial leucyl-tRNA synthetase) in LZ, GZ and DZ. Each dot corresponds to an independent biological sample. d, Immunofluorescence microscopy of GC 14 days post SRBC immunization examining colocalization of Cyclin B1 and LARS2. (a) is related to Fig. 4 panel a.
Extended Data Fig. 5 Analysis of single GCBC transcription in support of Fig. 6.
scRNA-Seq UMAP plots generated with Monocle3 displaying the enrichment for gene expression signatures derived from bulk RNA-Seq clusters 1-8. Bulk RNA-Seq gene expression trend is displayed to the side of each scRNA-Seq UMAP plot. Boxes represent interquartile ranges (IQRs; Q1–Q3 percentile) and black vertical lines represent median values. Maximum and minimum values (ends of whiskers) are defined as Q3 + 1.5× the IQR and Q1 − 1.5× the IQR, respectively.
Extended Data Fig. 6 Integration of transcriptional and proteomic analysis in support of Fig. 7.
a-c, Heatmap of gene expressions (left) and protein expressions (right) for the indicated genes. Full lists related to Fig. 7a, c, e. d-i, Heatmaps of head-to-head GCBC subset comparisons for IPA upstream regulator analysis. For factors indicated. d-f, Analysis of total proteome dataset. g-i, Analysis of phosphoproteome dataset. j, Summary of relative activation status for panels (d-i), for the indicated activated factors and cell types. For head-to-head comparisons, increased activation for the indicated factors corresponds to increased color intensity GZ (green), DZ (magenta), and LZ (blue).
Extended Data Fig. 7 Visualization of GZ proliferative clusters in support of Fig. 8.
a, Immunofluorescence microscopy 3D z stacks displaying whole GCs and focusing on GZ clusters within the GC. GL7 (green), Cyclin B1 (red), DAPI (blue). (n = 4) Right panel is related to GZ displayed in Fig. 7m. b, Schematic of EdU and BrdU injection experimental design. c and d, Immunofluorescence microscopy of whole GCs for the indicated markers 5.5 hr post BrdU injection (n≥3 mice per timepoint indicated in panel b). e and f, Immunofluorescent microscopy of GCs for the indicated markers (e) and quantification of EdU intensity within the GZ (f) 5.5 hr post BrdU injection. Each dot represents a cell. g, Schematic representing dilution of EdU during cell division (5.5 hr). h, Immunofluorescent microscopy of GCs for the indicated markers visualizing cells in multiple stages of active cell division within a GZ cluster 5.5 hr post BrdU injection (n > 3 mice).
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2.
Supplementary Data
Supplementary Data 1–5.
Rights and permissions
About this article
Cite this article
Kennedy, D.E., Okoreeh, M.K., Maienschein-Cline, M. et al. Novel specialized cell state and spatial compartments within the germinal center. Nat Immunol 21, 660–670 (2020). https://doi.org/10.1038/s41590-020-0660-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41590-020-0660-2
This article is cited by
-
B cells orchestrate tolerance to the neuromyelitis optica autoantigen AQP4
Nature (2024)
-
Defective mitochondria remodelling in B cells leads to an aged immune response
Nature Communications (2024)
-
The endogenous repertoire harbors self-reactive CD4+ T cell clones that adopt a follicular helper T cell-like phenotype at steady state
Nature Immunology (2023)
-
Rapid and sensitive single-cell RNA sequencing with SHERRY2
BMC Biology (2022)
-
Cellular metabolic adaptations in rheumatoid arthritis and their therapeutic implications
Nature Reviews Rheumatology (2022)